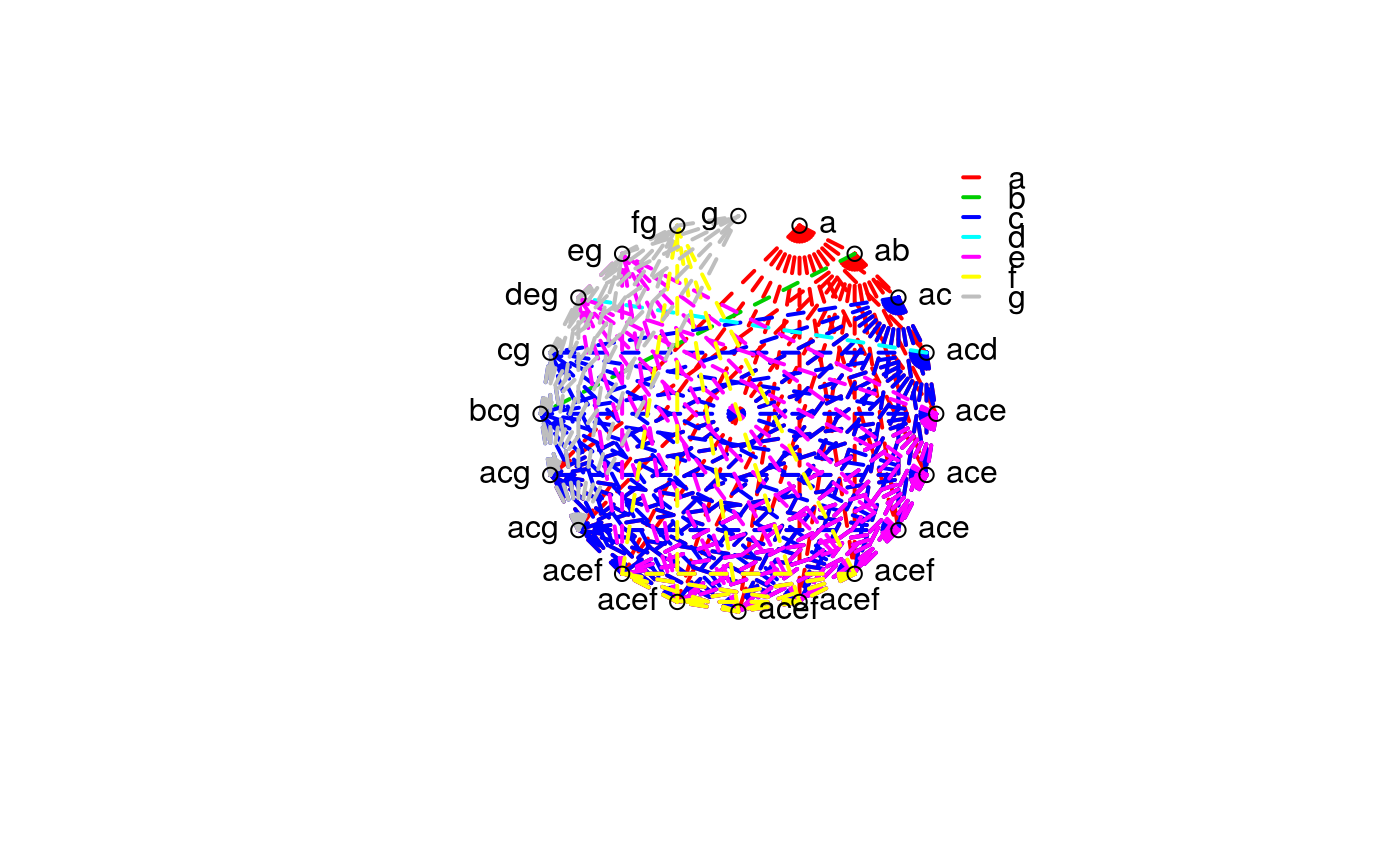
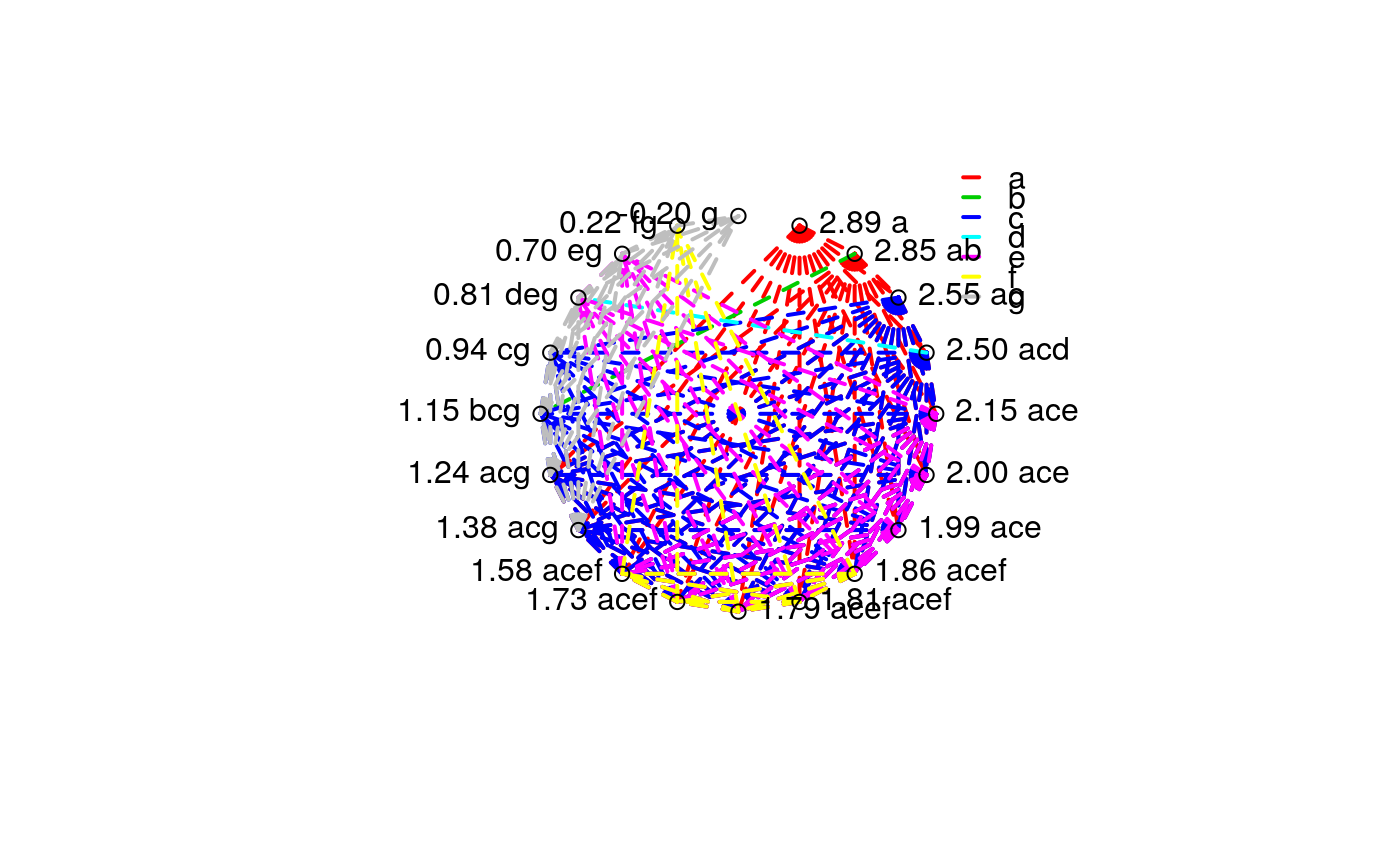
This function does a radial plot based on the vector of letters resulted from pairwise comparisons.
radial_cld(cld, labels = cld, col = NULL, means = NULL, perim = FALSE, legend = TRUE)
Arguments
| cld | Character vector with strings of letters that indicates which pair of treatment cells are not different. |
|---|---|
| labels | Vector of text to be annotated next each point. |
| col | Vector of colors to be used in the segments that joint points. |
| means | Numeric vector with the estimated means of treatment cells. It is used to place points at distances proportional to the differences on means. |
| perim | Logical value (default is |
| legend | Logical value (default is |
Value
None is returned, only the plot is done.
See also
cld2().
Examples
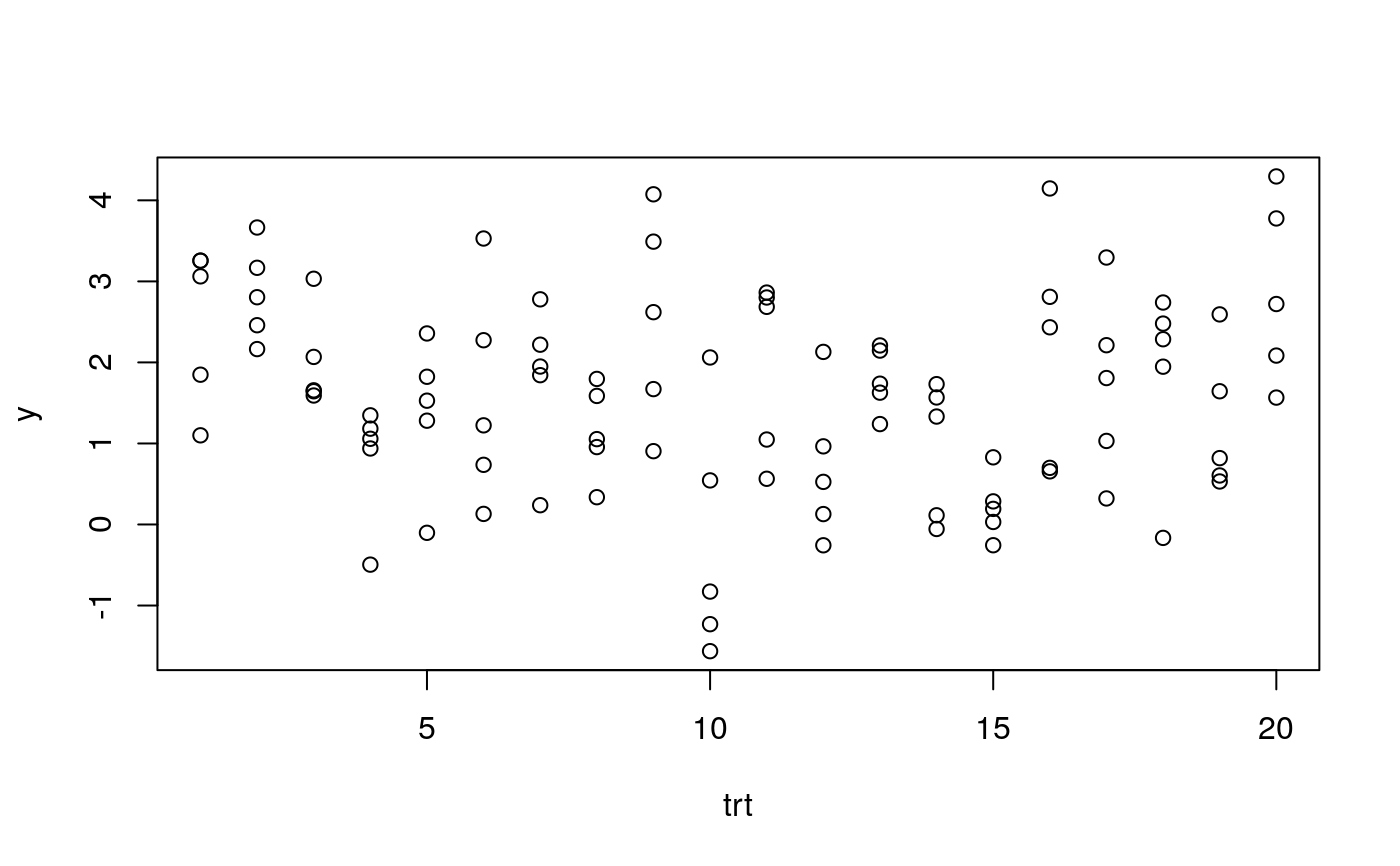
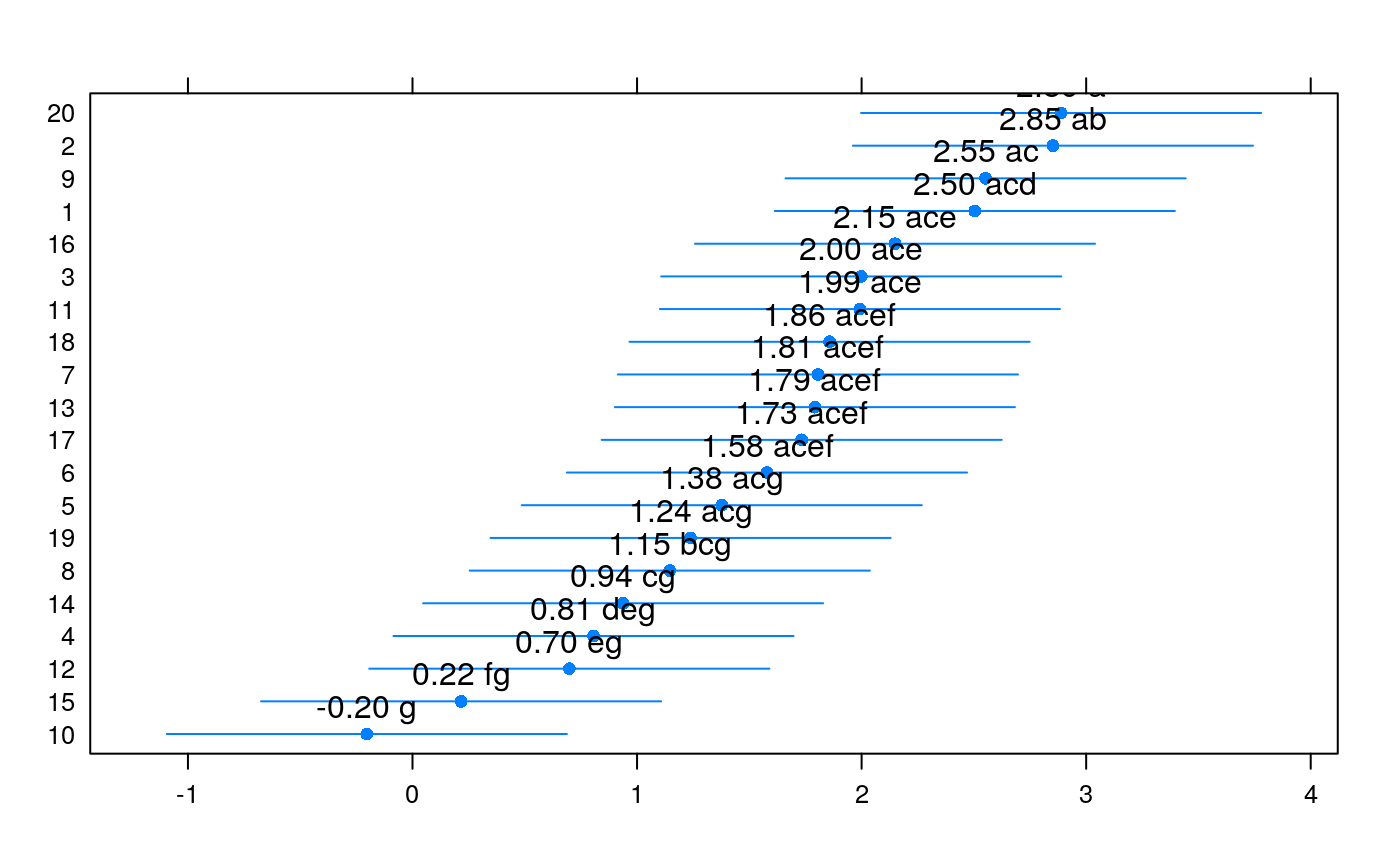
set.seed(4321) td <- data.frame(trt = rep(sample(1:20), each = 5)) td$y <- rnorm(nrow(td), mean = 0.15 * sort(td$trt), sd = 1) plot(y ~ trt, data = td)#> Analysis of Variance Table #> #> Response: y #> Df Sum Sq Mean Sq F value Pr(>F) #> trt 19 65.815 3.4640 3.4576 5.194e-05 *** #> Residuals 80 80.146 1.0018 #> --- #> Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1summary(m0)#> #> Call: #> lm(formula = y ~ trt, data = td) #> #> Residuals: #> Min 1Q Median 3Q Max #> -2.0226 -0.6637 0.0693 0.6348 2.2642 #> #> Coefficients: #> Estimate Std. Error t value Pr(>|t|) #> (Intercept) 2.50426 0.44762 5.595 2.99e-07 *** #> trt2 0.34782 0.63303 0.549 0.584225 #> trt3 -0.50600 0.63303 -0.799 0.426472 #> trt4 -1.69826 0.63303 -2.683 0.008870 ** #> trt5 -1.12692 0.63303 -1.780 0.078841 . #> trt6 -0.92557 0.63303 -1.462 0.147624 #> trt7 -0.69882 0.63303 -1.104 0.272939 #> trt8 -1.35856 0.63303 -2.146 0.034894 * #> trt9 0.04789 0.63303 0.076 0.939889 #> trt10 -2.70744 0.63303 -4.277 5.20e-05 *** #> trt11 -0.51198 0.63303 -0.809 0.421043 #> trt12 -1.80581 0.63303 -2.853 0.005517 ** #> trt13 -0.71239 0.63303 -1.125 0.263802 #> trt14 -1.56631 0.63303 -2.474 0.015465 * #> trt15 -2.28781 0.63303 -3.614 0.000525 *** #> trt16 -0.35552 0.63303 -0.562 0.575952 #> trt17 -0.77093 0.63303 -1.218 0.226867 #> trt18 -0.64693 0.63303 -1.022 0.309886 #> trt19 -1.26594 0.63303 -2.000 0.048916 * #> trt20 0.38425 0.63303 0.607 0.545568 #> --- #> Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1 #> #> Residual standard error: 1.001 on 80 degrees of freedom #> Multiple R-squared: 0.4509, Adjusted R-squared: 0.3205 #> F-statistic: 3.458 on 19 and 80 DF, p-value: 5.194e-05 #>library(multcomp) library(doBy) X <- LE_matrix(m0, effect = "trt") rownames(X) <- levels(td$trt) ci <- apmc(X, m0, focus = "trt", test = "fdr") ci$cld <- with(ci, ordered_cld(cld, fit)) ci <- ci[order(ci$fit, decreasing = TRUE), ] library(latticeExtra) segplot(reorder(trt, fit) ~ lwr + upr, centers = fit, data = ci, draw = FALSE, cld = ci$cld) + layer(panel.text(x = centers, y = z, labels = sprintf("%0.2f %s", centers, cld), pos = 3))radial_cld(cld = ci$cld)#> Warning: Length of vector `col` is different of the number of unique letters. Colors will be recycled.radial_cld(cld = ci$cld, means = ci$fit, perim = TRUE)#> Warning: Length of vector `col` is different of the number of unique letters. Colors will be recycled.radial_cld(cld = ci$cld, col = 1:3)#> Warning: Length of vector `col` is different of the number of unique letters. Colors will be recycled.radial_cld(cld = ci$cld, col = 1:3)#> Warning: Length of vector `col` is different of the number of unique letters. Colors will be recycled.#> Warning: Length of vector `col` is different of the number of unique letters. Colors will be recycled.