Desenvolvimento de Mudas de Eucalipto em Resposta à Doses de P e Biocarvão
Walmes M. Zeviani & Milson E. Serafim
11 de julho de 2019
Source:vignettes/biocar_eucal.Rmd
biocar_eucal.RmdDescrição e Análise Exploratória
# Estrutura dos dados.
str(biocar_eucal)## 'data.frame': 90 obs. of 22 variables:
## $ biocar: Factor w/ 3 levels "S_BP","SP","SP_B": 1 1 1 1 1 1 1 1 1 1 ...
## $ dosep : num 0 0 0 1.5 1.5 1.5 3 3 3 6 ...
## $ bloc : Factor w/ 3 levels "I","II","III": 1 2 3 1 2 3 1 2 3 1 ...
## $ altini: num 17.6 17.6 17.6 17.6 17.6 17.6 17.6 17.6 17.6 17.6 ...
## $ diaini: num 2.2 2.2 2.2 2.2 2.2 2.2 2.2 2.2 2.2 2.2 ...
## $ altfin: num 83 57 58 64 64 83 59 67 60 66 ...
## $ diafin: num 6.2 5.1 5.6 6.4 5.5 5.8 6.6 6.3 6.4 6.1 ...
## $ mfpa : num 51.1 44 41.3 58.1 35 61.4 68.4 56.1 52.7 57.6 ...
## $ mfr : num 21.4 7.3 12.1 15.4 7.6 31 18.5 18.8 5.3 17.9 ...
## $ mspa : num 12.7 9.8 8.7 14.5 6.8 15.2 19.4 13 12.4 16.8 ...
## $ msr : num 10.1 6 6.5 7.5 4.1 11.7 9.7 8.4 4.2 9.4 ...
## $ ph : num 7.6 7.2 7.3 7.1 6.1 6.7 7.3 7.6 5.6 6.7 ...
## $ phkcl : num 7.2 6.5 6.6 6.7 5.3 6.1 7.1 7.3 5.1 5.8 ...
## $ pcz : num 6.8 5.8 5.9 6.3 4.5 5.5 6.9 7 4.6 4.9 ...
## $ p : num 42.3 35.2 35.8 64.1 67.6 57.2 73.6 73 33.6 43.3 ...
## $ k : num 21.1 16.5 13.4 12.5 15 15.8 13.5 16.6 13.6 16.3 ...
## $ ca : num 5.9 4.3 5.2 4.8 4.1 4.6 6.1 5.3 4.3 4.7 ...
## $ mg : num 0.9 2.6 1.2 1.5 1.2 1.7 1.3 2.6 1.8 2.5 ...
## $ hal : num 1 1.2 1 1.1 1.7 1.2 1.1 1 2.1 1.7 ...
## $ are : num 69.4 71.3 70.3 70.3 70.6 70.9 70.9 69.8 70.8 70.2 ...
## $ sil : num 15.1 13.6 14.5 14.5 14 13.8 13.9 15 13.8 14.1 ...
## $ arg : num 15.5 15.1 15.2 15.2 15.4 15.4 15.2 15.2 15.4 15.7 ...# Usa objeto de nome curto mais fácil de manipular.
bio <- biocar_eucal
# Tabela de frequência do desenho experimental.
xtabs(~dosep + biocar, data = bio)## biocar
## dosep S_BP SP SP_B
## 0 3 3 3
## 1.5 3 3 3
## 3 3 3 3
## 6 3 3 3
## 12 3 3 3
## 24 3 3 3
## 48 3 3 3
## 96 3 3 3
## 192 3 3 3
## 384 3 3 3# Verifica se alguma variável resposta tem valor perdido.
sapply(bio[, -(1:3)], FUN = function(x) any(is.na(x)))## altini diaini altfin diafin mfpa mfr mspa msr ph
## FALSE FALSE FALSE FALSE FALSE FALSE FALSE FALSE FALSE
## phkcl pcz p k ca mg hal are sil
## FALSE FALSE TRUE FALSE FALSE FALSE FALSE FALSE FALSE
## arg
## FALSE# Empilha a variável resposta para fazer gráficos.
bioe <- melt(bio,
id.vars = 1:3,
measure.vars = 6:ncol(bio),
variable_name = "resp")
xyplot(value ~ dosep^0.3 | resp,
groups = biocar,
data = bioe,
as.table = TRUE,
type = c("p", "smooth"),
auto.key = list(title = "Biocarvão",
cex.title = 1.1,
columns = 3),
scales = list(y = list(relation = "free")))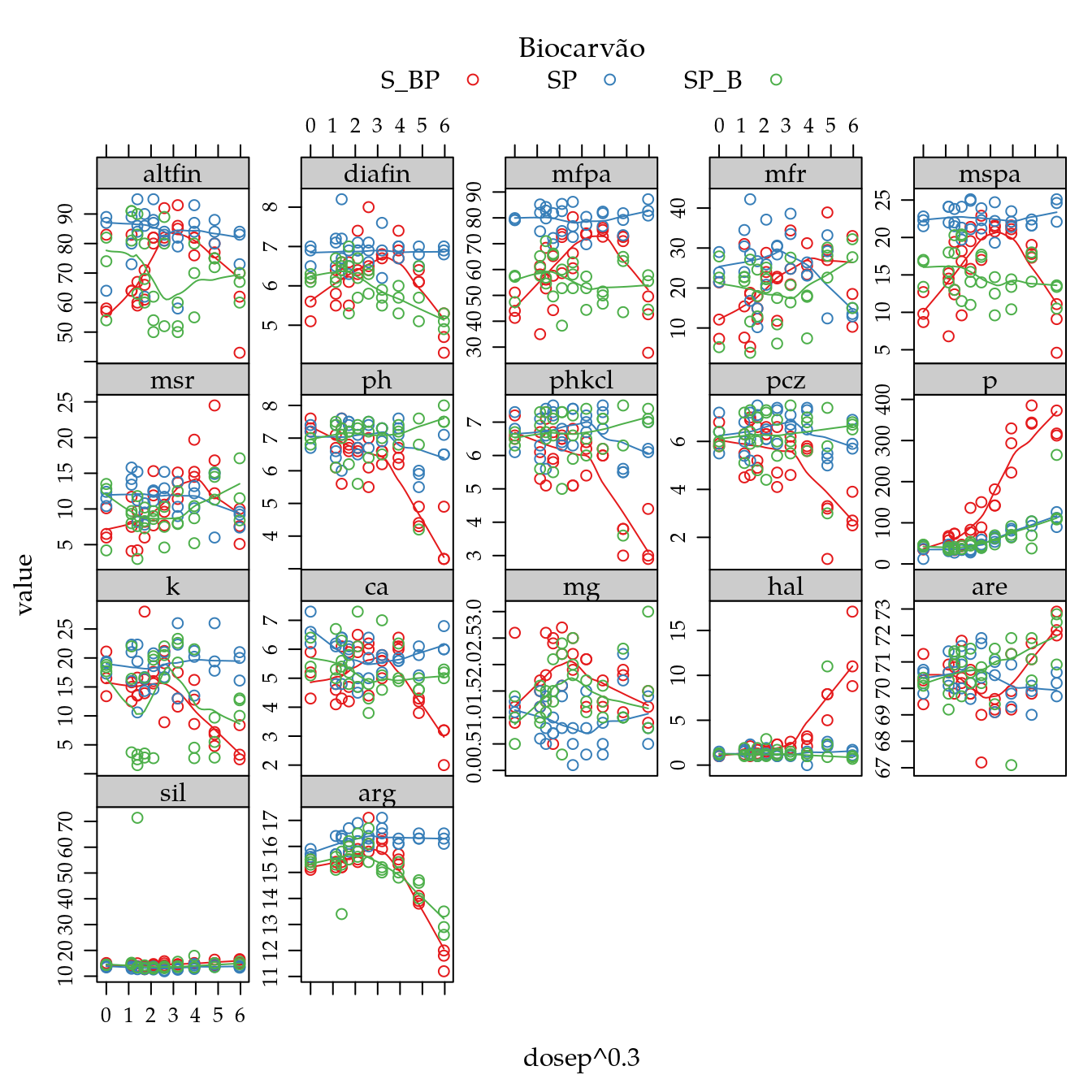
Transformação potência para equalizar distância entre níveis
## [1] 0.0 1.5 3.0 6.0 12.0 24.0 48.0 96.0 192.0 384.0# Variância das distância entre níveis em escala unitária.
esp <- function(p) {
u <- x^p
u <- (u - min(u))
u <- u/max(u)
var(diff(u))
}
# Otimiza para obter o valor de potência para maior uniformidade.
op <- optimize(f = esp, interval = c(0, 1))
op$minimum## [1] 0.3064936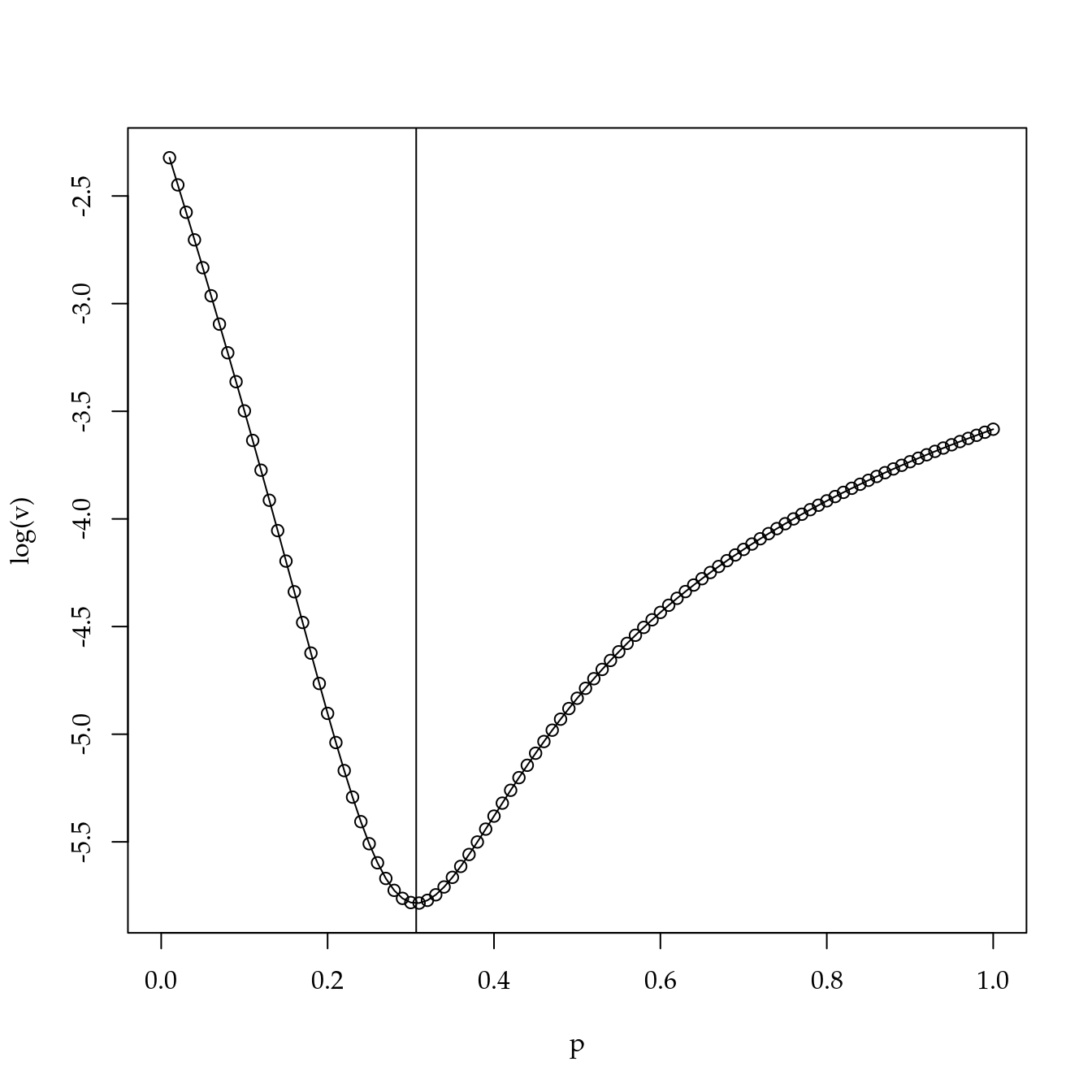
# Cria potência da dose.
bio$dose <- bio$dosep^0.3
bio$P <- factor(bio$dosep)Altura final
#-----------------------------------------------------------------------
# Modelo saturado (considera dose como qualitativo).
m0 <- lm(altfin ~ bloc + biocar * P, data = bio)
par(mfrow = c(2, 2))
plot(m0)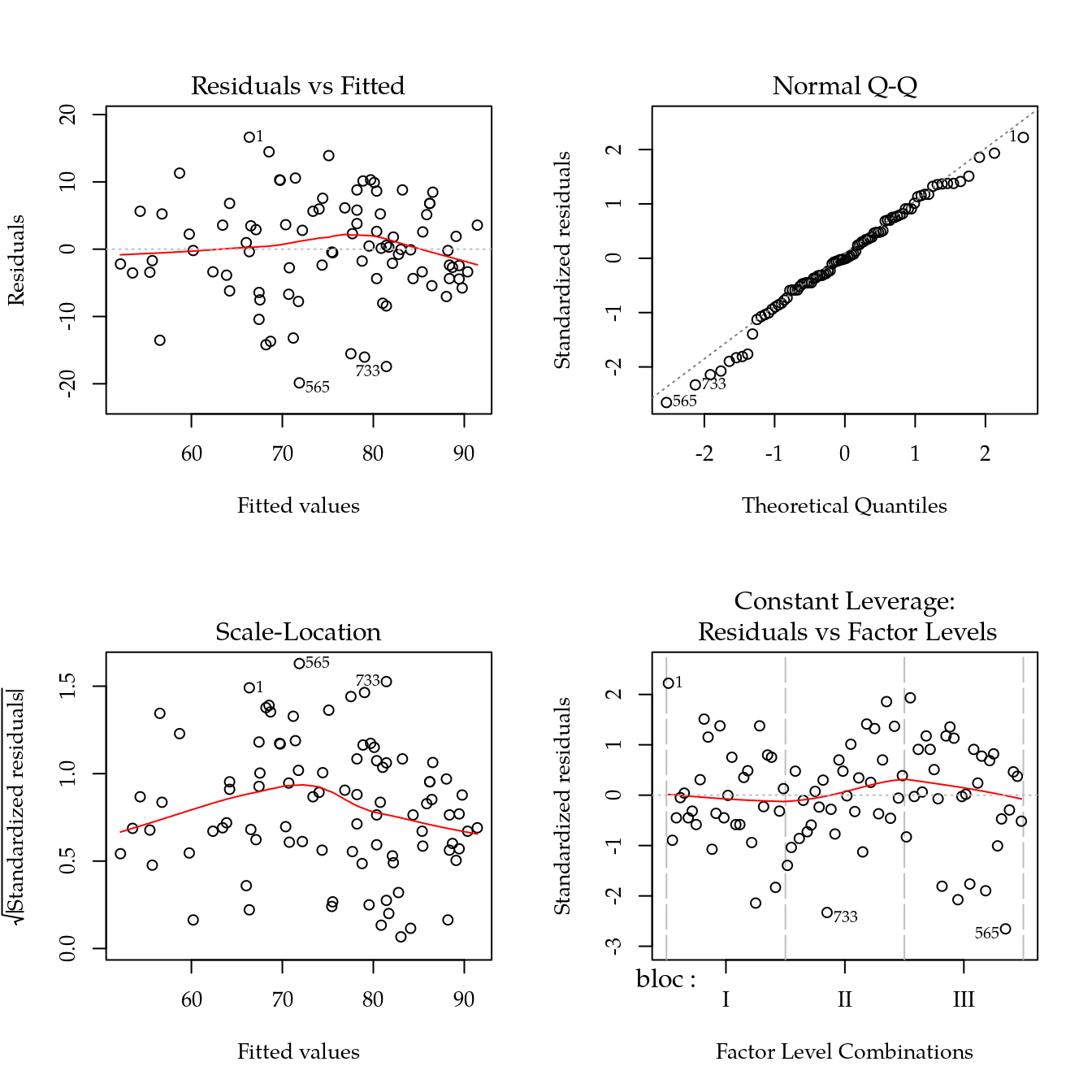
## Analysis of Variance Table
##
## Response: altfin
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 162.9 81.43 0.9373 0.3975388
## biocar 2 2654.9 1327.43 15.2786 4.676e-06 ***
## P 9 1431.4 159.05 1.8306 0.0818713 .
## biocar:P 18 4847.8 269.32 3.0999 0.0005685 ***
## Residuals 58 5039.1 86.88
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1L <- LE_matrix(m0, effect = c("biocar", "P"))
pred <- equallevels(attr(L, "grid"), bio)
ci <- confint(glht(m0, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
str(pred)## 'data.frame': 30 obs. of 5 variables:
## $ biocar : Factor w/ 3 levels "S_BP","SP","SP_B": 1 2 3 1 2 3 1 2 3 1 ...
## $ P : Factor w/ 10 levels "0","1.5","3",..: 1 1 1 2 2 2 3 3 3 4 ...
## $ Estimate: num 66 80 70 70.3 80.7 ...
## $ lwr : num 55.2 69.2 59.2 59.6 69.9 ...
## $ upr : num 76.8 90.8 80.8 81.1 91.4 ...pch <- c(1, 4, 5)
key <- list(title = "Biocarvão",
cex.title = 1.1,
text = list(levels(bio$biocar)),
lines = list(pch = pch, lty = 1),
columns = 3,
type = "o",
divide = 1)
segplot(P ~ lwr + upr,
centers = Estimate,
data = pred,
draw = FALSE,
groups = biocar,
xlab = "Dose de P",
ylab = "Altura final (cm)",
key = key,
horizontal = FALSE,
pch = pch,
gap = 0.1,
panel = panel.groups.segplot)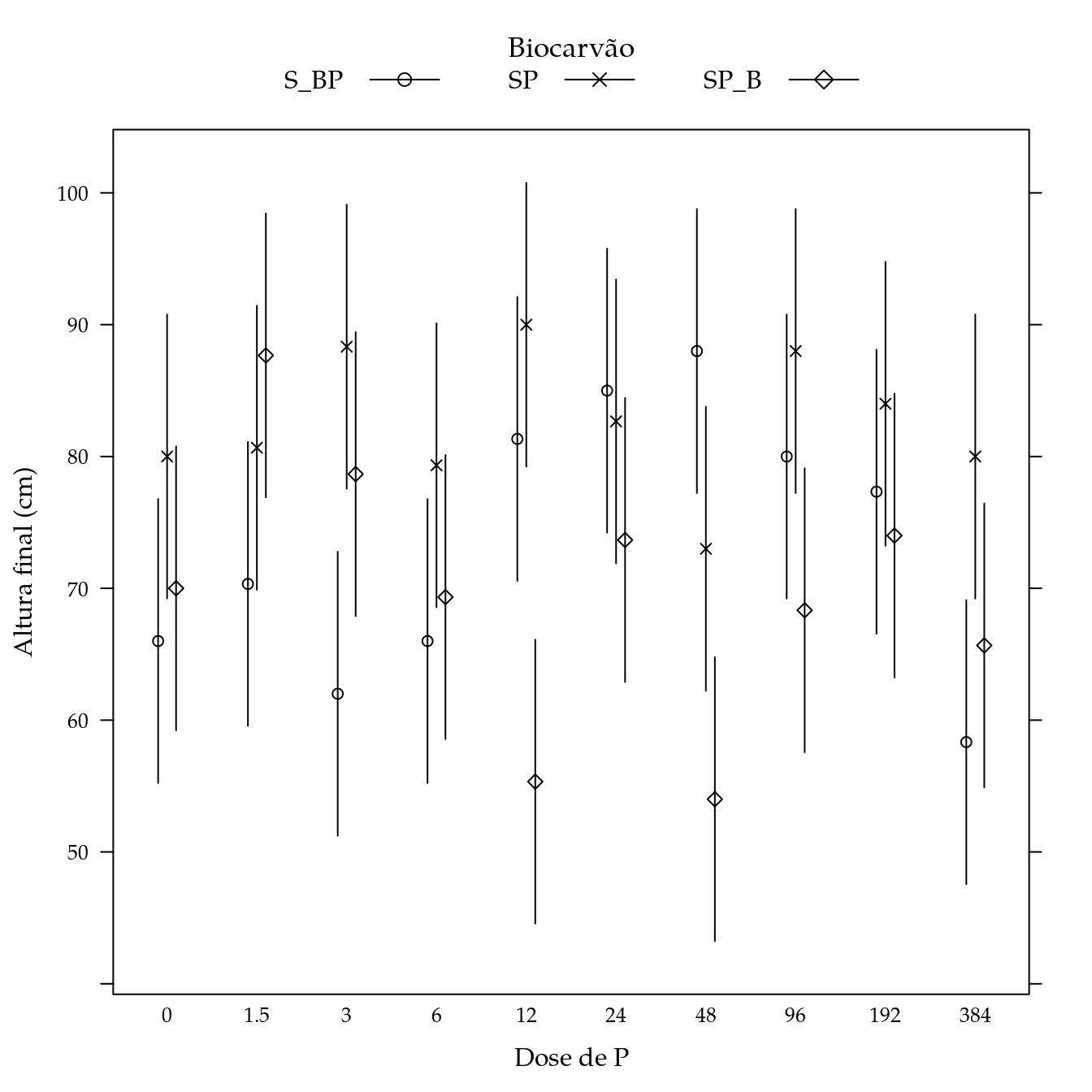
#-----------------------------------------------------------------------
# Modelo reduzido baseado em splines.
m1 <- update(m0, ~ bloc + biocar * bs(dose, df = 4))
par(mfrow = c(2, 2))
plot(m1)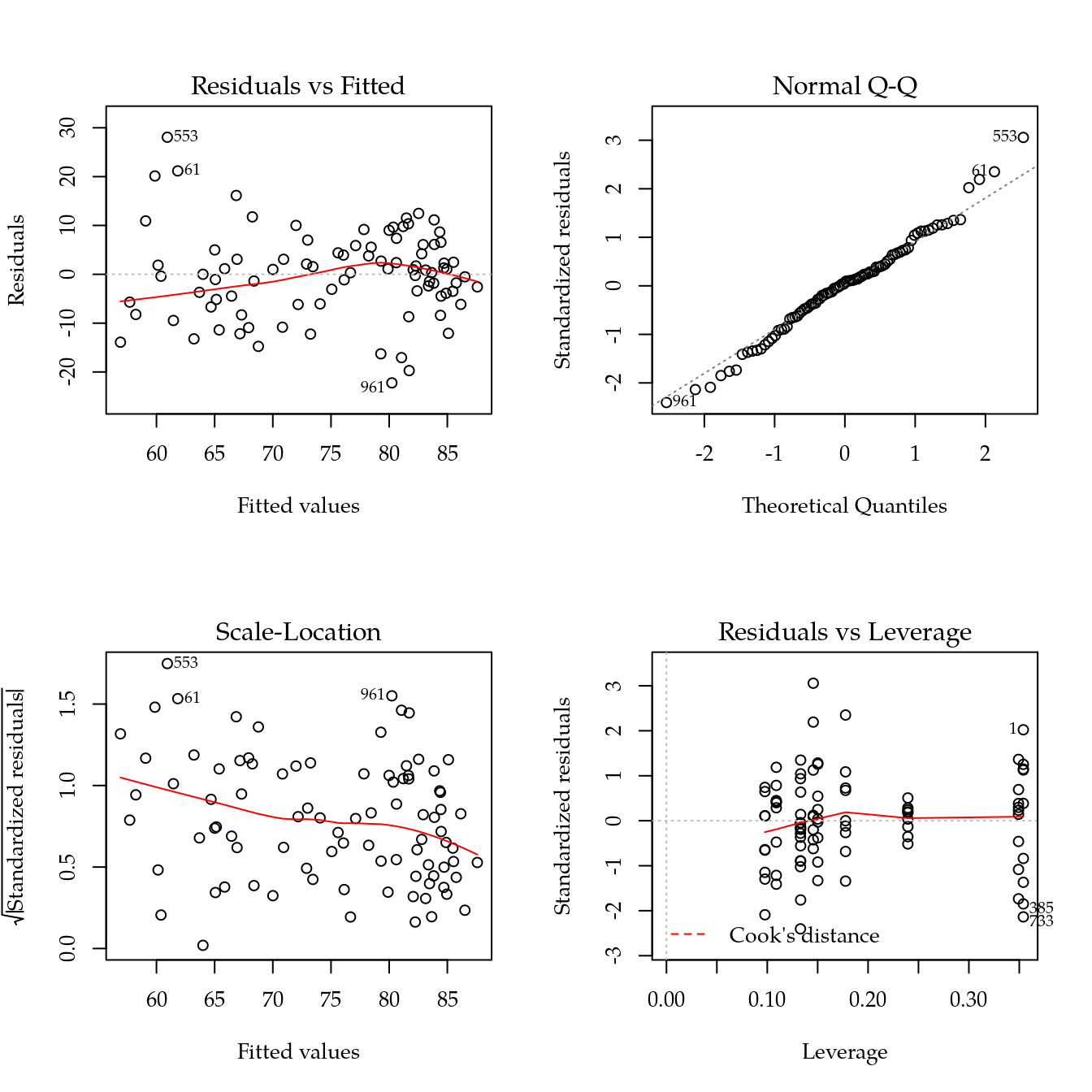
## Analysis of Variance Table
##
## Response: altfin
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 162.9 81.43 0.8254 0.442094
## biocar 2 2654.9 1327.43 13.4552 1.06e-05 ***
## bs(dose, df = 4) 4 738.8 184.69 1.8721 0.124470
## biocar:bs(dose, df = 4) 8 3377.7 422.21 4.2797 0.000295 ***
## Residuals 73 7201.9 98.66
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Teste para a falta de ajuste.
anova(m1, m0)## Analysis of Variance Table
##
## Model 1: altfin ~ bloc + biocar + bs(dose, df = 4) + biocar:bs(dose, df = 4)
## Model 2: altfin ~ bloc + biocar * P
## Res.Df RSS Df Sum of Sq F Pr(>F)
## 1 73 7201.9
## 2 58 5039.1 15 2162.8 1.6595 0.086 .
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Novos valores em grid para fazer o gráfico de bandas.
pred <- with(bio,
expand.grid(bloc = factor(levels(bloc)[1],
levels = levels(bloc)),
biocar = levels(biocar),
dose = seq(min(dose),
max(dose),
length.out = 60)))
# Extrai a especificação do tempo base splines.
form <- attr(terms(formula(m1)), "term.labels")[3]
B <- eval(parse(text = form), envir = bio)
str(B)## 'bs' num [1:90, 1:4] 0 0 0 0.648 0.648 ...
## - attr(*, "dimnames")=List of 2
## ..$ : NULL
## ..$ : chr [1:4] "1" "2" "3" "4"
## - attr(*, "degree")= int 3
## - attr(*, "knots")= Named num 2.35
## ..- attr(*, "names")= chr "50%"
## - attr(*, "Boundary.knots")= num [1:2] 0 5.96
## - attr(*, "intercept")= logi FALSE# Gera a matriz para os valores de predição.
X <- predict(B, pred$dos)
# Cria a matriz do modelo para fazer a predição.
L <- model.matrix(~bloc + biocar * X, data = pred)
# Dá pesos para os efeitos de blocos correspondente à LSmeans.
i <- attr(L, "assign") == 1
L[, i] <- 1/(sum(i) + 1)
# Obtém os valores preditos com IC.
ci <- confint(glht(m1, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
xyplot(altfin ~ dose,
groups = biocar,
data = bio,
auto.key = list(points = FALSE,
lines = TRUE,
divide = 1,
type = "o")) +
as.layer(xyplot(Estimate ~ dose,
groups = biocar,
data = pred,
type = "l",
ly = pred$lwr,
uy = pred$upr,
cty = "bands",
alpha = 0.3,
prepanel = prepanel.cbH,
panel.groups = panel.cbH,
panel = panel.superpose))
Diâmetro final
#-----------------------------------------------------------------------
# Modelo saturado (considera dose como qualitativo).
m0 <- lm(diafin ~ bloc + biocar * P, data = bio)
par(mfrow = c(2, 2))
plot(m0)
## Analysis of Variance Table
##
## Response: diafin
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 1.1527 0.5763 3.1239 0.0514683 .
## biocar 2 13.1307 6.5653 35.5856 8.234e-11 ***
## P 9 8.9588 0.9954 5.3954 2.357e-05 ***
## biocar:P 18 9.8982 0.5499 2.9806 0.0008518 ***
## Residuals 58 10.7007 0.1845
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1L <- LE_matrix(m0, effect = c("biocar", "P"))
pred <- equallevels(attr(L, "grid"), bio)
ci <- confint(glht(m0, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
str(pred)## 'data.frame': 30 obs. of 5 variables:
## $ biocar : Factor w/ 3 levels "S_BP","SP","SP_B": 1 2 3 1 2 3 1 2 3 1 ...
## $ P : Factor w/ 10 levels "0","1.5","3",..: 1 1 1 2 2 2 3 3 3 4 ...
## $ Estimate: num 5.63 6.8 6.2 5.9 6.93 ...
## $ lwr : num 5.14 6.3 5.7 5.4 6.44 ...
## $ upr : num 6.13 7.3 6.7 6.4 7.43 ...pch <- c(1, 4, 5)
key <- list(title = "Biocarvão",
cex.title = 1.1,
text = list(levels(bio$biocar)),
lines = list(pch = pch, lty = 1),
columns = 3,
type = "o",
divide = 1)
segplot(P ~ lwr + upr,
centers = Estimate,
data = pred,
draw = FALSE,
groups = biocar,
xlab = "Dose de P",
ylab = "Altura final (cm)",
key = key,
horizontal = FALSE,
pch = pch,
gap = 0.1,
panel = panel.groups.segplot)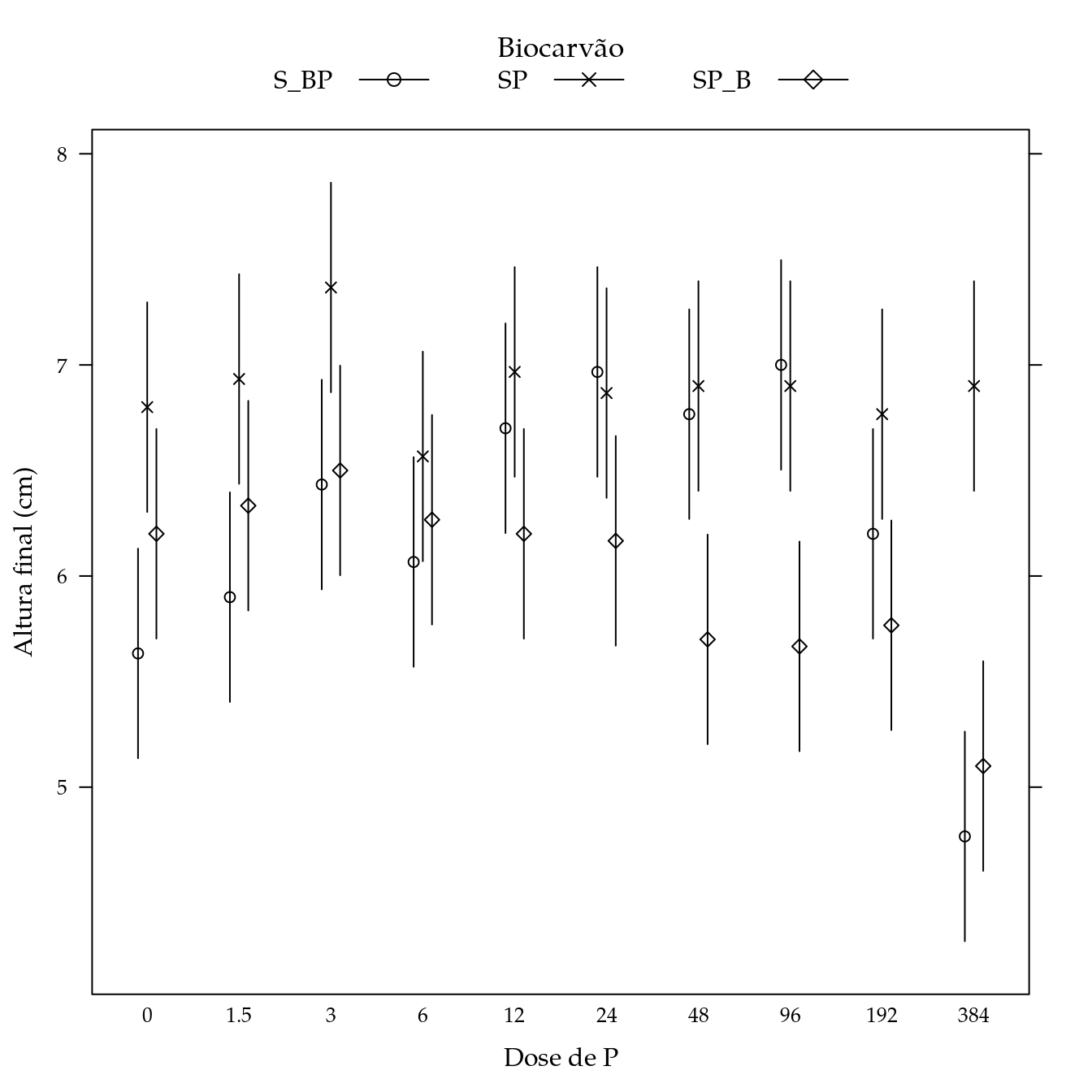
#-----------------------------------------------------------------------
# Modelo reduzido baseado em splines.
m1 <- update(m0, ~ bloc + biocar * bs(dose, df = 3))
par(mfrow = c(2, 2))
plot(m1)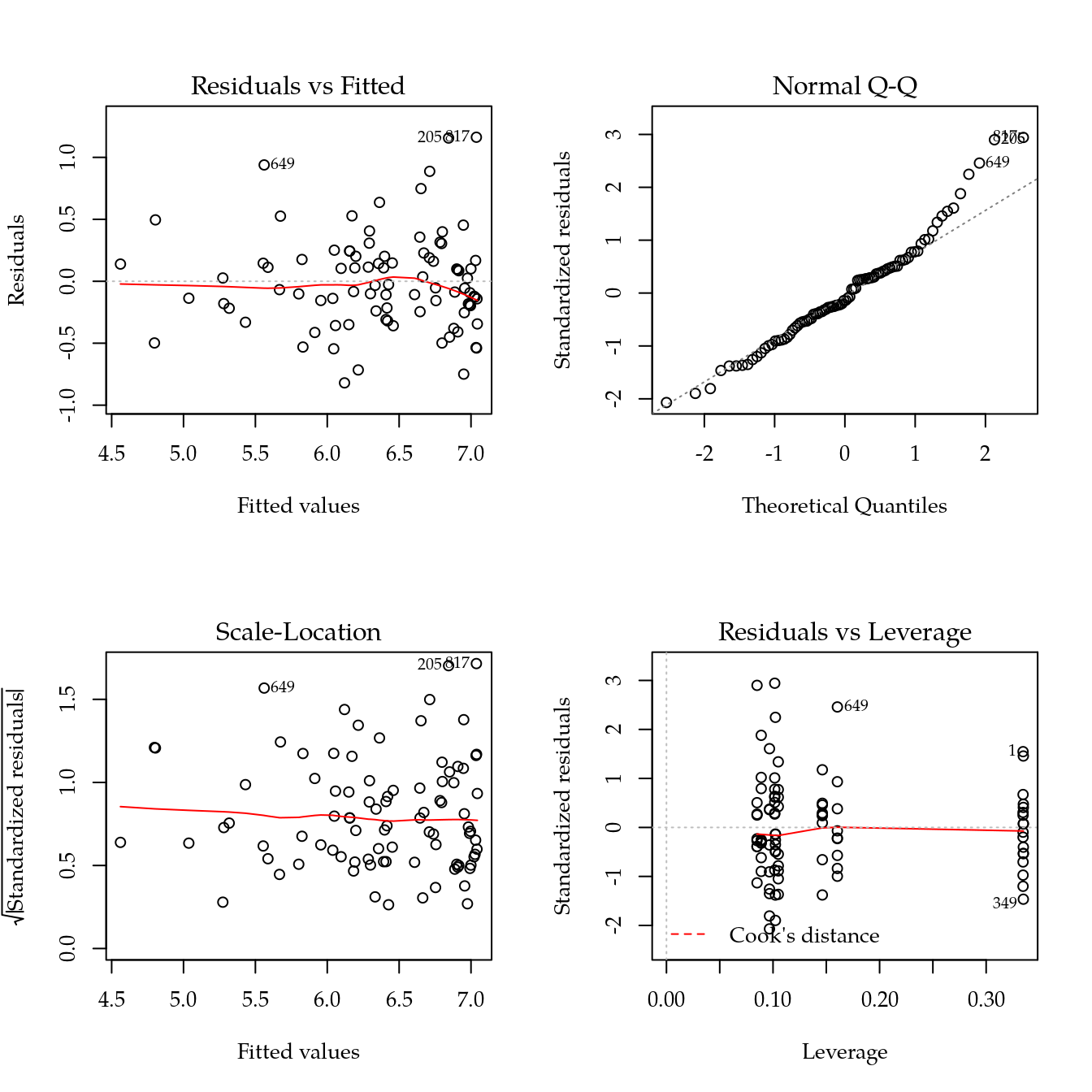
## Analysis of Variance Table
##
## Response: diafin
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 1.1527 0.5763 3.3175 0.04156 *
## biocar 2 13.1307 6.5653 37.7911 4.039e-12 ***
## bs(dose, df = 3) 3 7.5109 2.5036 14.4113 1.598e-07 ***
## biocar:bs(dose, df = 3) 6 8.8435 1.4739 8.4841 4.684e-07 ***
## Residuals 76 13.2033 0.1737
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Teste para a falta de ajuste.
anova(m1, m0)## Analysis of Variance Table
##
## Model 1: diafin ~ bloc + biocar + bs(dose, df = 3) + biocar:bs(dose, df = 3)
## Model 2: diafin ~ bloc + biocar * P
## Res.Df RSS Df Sum of Sq F Pr(>F)
## 1 76 13.203
## 2 58 10.701 18 2.5026 0.7536 0.7421# Novos valores em grid para fazer o gráfico de bandas.
pred <- with(bio,
expand.grid(bloc = factor(levels(bloc)[1],
levels = levels(bloc)),
biocar = levels(biocar),
dose = seq(min(dose),
max(dose),
length.out = 60)))
# Extrai a especificação do tempo base splines.
form <- attr(terms(formula(m1)), "term.labels")[3]
B <- eval(parse(text = form), envir = bio)
str(B)## 'bs' num [1:90, 1:3] 0 0 0 0.373 0.373 ...
## - attr(*, "dimnames")=List of 2
## ..$ : NULL
## ..$ : chr [1:3] "1" "2" "3"
## - attr(*, "degree")= int 3
## - attr(*, "knots")= num(0)
## - attr(*, "Boundary.knots")= num [1:2] 0 5.96
## - attr(*, "intercept")= logi FALSE# Gera a matriz para os valores de predição.
X <- predict(B, pred$dos)
# Cria a matriz do modelo para fazer a predição.
L <- model.matrix(~bloc + biocar * X, data = pred)
# Dá pesos para os efeitos de blocos correspondente à LSmeans.
i <- attr(L, "assign") == 1
L[, i] <- 1/(sum(i) + 1)
# Obtém os valores preditos com IC.
ci <- confint(glht(m1, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
xyplot(diafin ~ dose,
groups = biocar,
data = bio,
auto.key = list(points = FALSE,
lines = TRUE,
divide = 1,
type = "o")) +
as.layer(xyplot(Estimate ~ dose,
groups = biocar,
data = pred,
type = "l",
ly = pred$lwr,
uy = pred$upr,
cty = "bands",
alpha = 0.3,
prepanel = prepanel.cbH,
panel.groups = panel.cbH,
panel = panel.superpose))
Massa fresca de parte aérea (mfpa)
#-----------------------------------------------------------------------
# Modelo saturado (considera dose como qualitativo).
m0 <- lm(mfpa ~ bloc + biocar * P, data = bio)
par(mfrow = c(2, 2))
plot(m0)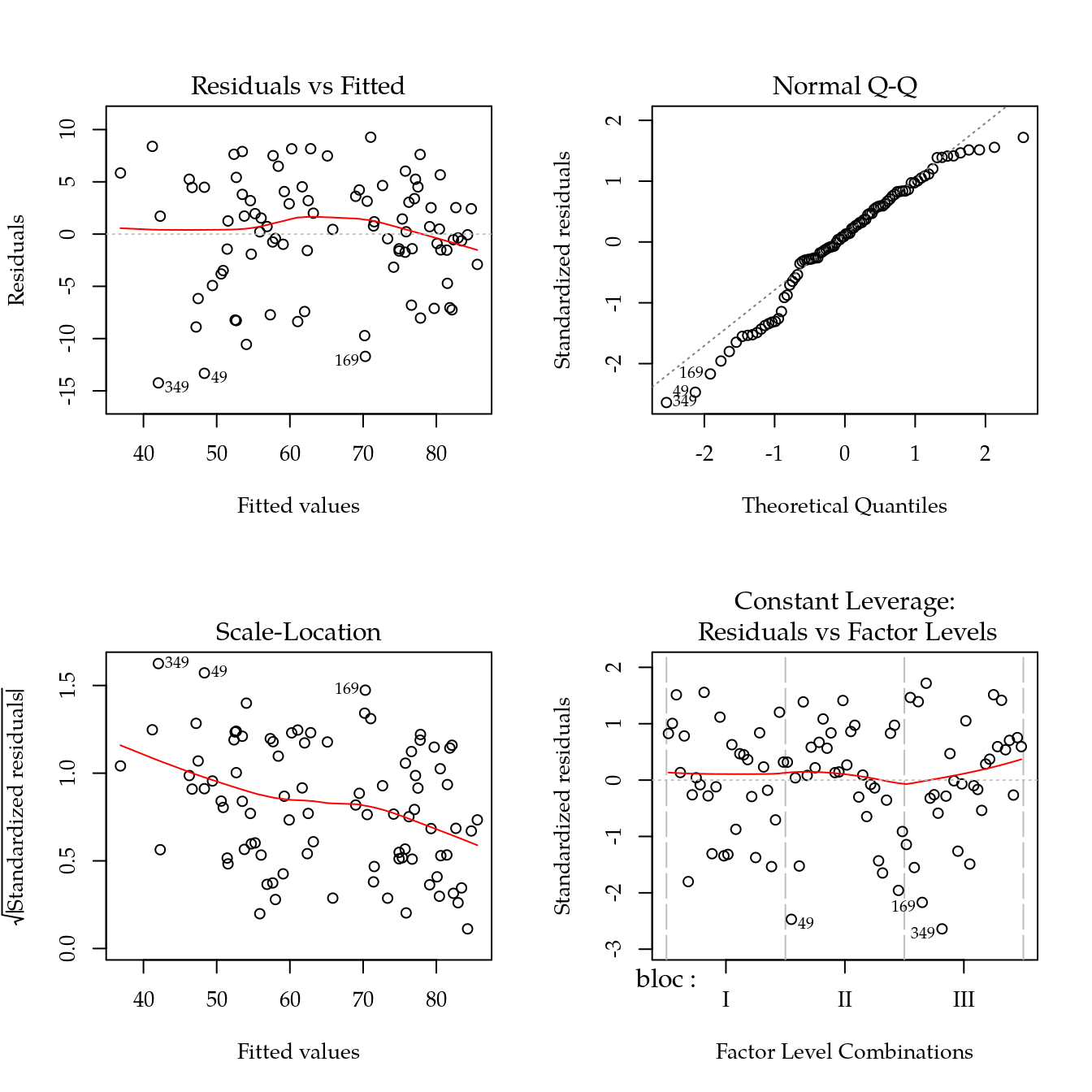
## Analysis of Variance Table
##
## Response: mfpa
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 464.9 232.5 5.1533 0.00871 **
## biocar 2 9399.0 4699.5 104.1834 < 2.2e-16 ***
## P 9 1085.3 120.6 2.6733 0.01143 *
## biocar:P 18 3860.4 214.5 4.7545 2.858e-06 ***
## Residuals 58 2616.3 45.1
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1L <- LE_matrix(m0, effect = c("biocar", "P"))
pred <- equallevels(attr(L, "grid"), bio)
ci <- confint(glht(m0, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
str(pred)## 'data.frame': 30 obs. of 5 variables:
## $ biocar : Factor w/ 3 levels "S_BP","SP","SP_B": 1 2 3 1 2 3 1 2 3 1 ...
## $ P : Factor w/ 10 levels "0","1.5","3",..: 1 1 1 2 2 2 3 3 3 4 ...
## $ Estimate: num 45.5 79.4 54.1 51.5 80.7 ...
## $ lwr : num 37.7 71.7 46.3 43.7 72.9 ...
## $ upr : num 53.2 87.2 61.8 59.3 88.4 ...pch <- c(1, 4, 5)
key <- list(title = "Biocarvão",
cex.title = 1.1,
text = list(levels(bio$biocar)),
lines = list(pch = pch, lty = 1),
columns = 3,
type = "o",
divide = 1)
segplot(P ~ lwr + upr,
centers = Estimate,
data = pred,
draw = FALSE,
groups = biocar,
xlab = "Dose de P",
ylab = "Altura final (cm)",
key = key,
horizontal = FALSE,
pch = pch,
gap = 0.1,
panel = panel.groups.segplot)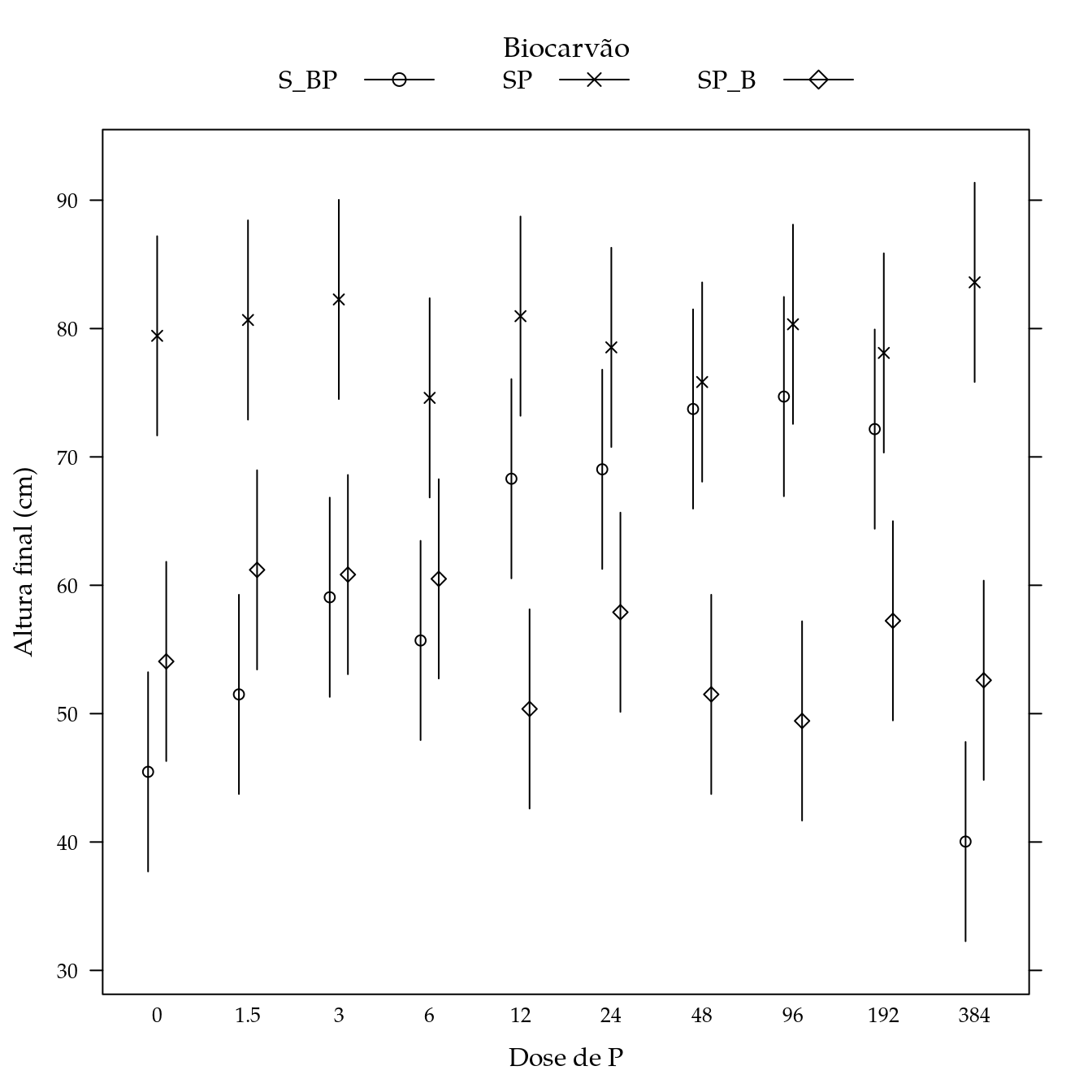
#-----------------------------------------------------------------------
# Modelo reduzido baseado em splines.
m1 <- update(m0, ~ bloc + biocar * bs(dose, df = 3))
par(mfrow = c(2, 2))
plot(m1)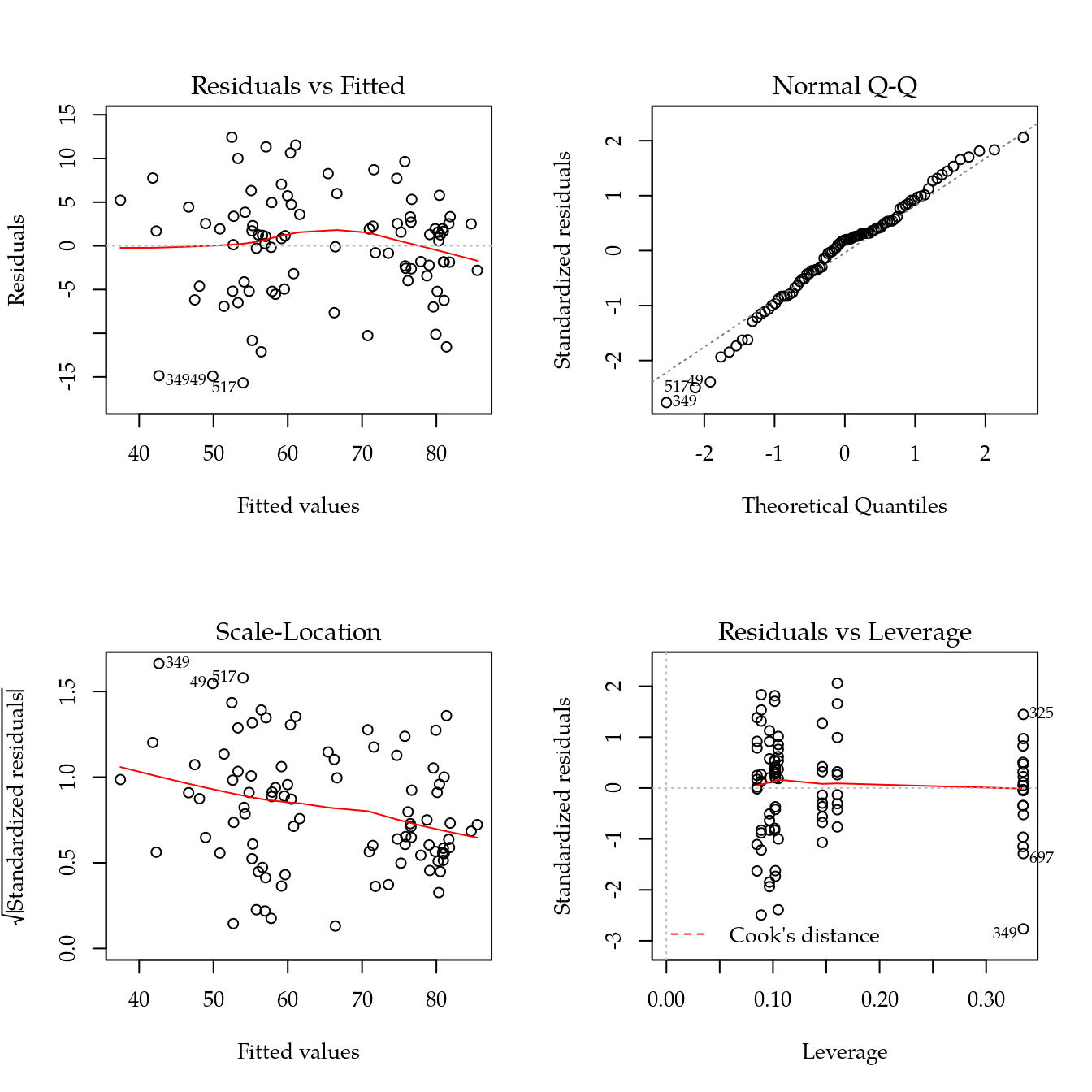
## Analysis of Variance Table
##
## Response: mfpa
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 464.9 232.5 5.3557 0.0066685 **
## biocar 2 9399.0 4699.5 108.2758 < 2.2e-16 ***
## bs(dose, df = 3) 3 872.5 290.8 6.7008 0.0004504 ***
## biocar:bs(dose, df = 3) 6 3390.8 565.1 13.0205 4.524e-10 ***
## Residuals 76 3298.6 43.4
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Teste para a falta de ajuste.
anova(m1, m0)## Analysis of Variance Table
##
## Model 1: mfpa ~ bloc + biocar + bs(dose, df = 3) + biocar:bs(dose, df = 3)
## Model 2: mfpa ~ bloc + biocar * P
## Res.Df RSS Df Sum of Sq F Pr(>F)
## 1 76 3298.6
## 2 58 2616.3 18 682.37 0.8404 0.6473# Novos valores em grid para fazer o gráfico de bandas.
pred <- with(bio,
expand.grid(bloc = factor(levels(bloc)[1],
levels = levels(bloc)),
biocar = levels(biocar),
dose = seq(min(dose),
max(dose),
length.out = 60)))
# Extrai a especificação do tempo base splines.
form <- attr(terms(formula(m1)), "term.labels")[3]
B <- eval(parse(text = form), envir = bio)
str(B)## 'bs' num [1:90, 1:3] 0 0 0 0.373 0.373 ...
## - attr(*, "dimnames")=List of 2
## ..$ : NULL
## ..$ : chr [1:3] "1" "2" "3"
## - attr(*, "degree")= int 3
## - attr(*, "knots")= num(0)
## - attr(*, "Boundary.knots")= num [1:2] 0 5.96
## - attr(*, "intercept")= logi FALSE# Gera a matriz para os valores de predição.
X <- predict(B, pred$dos)
# Cria a matriz do modelo para fazer a predição.
L <- model.matrix(~bloc + biocar * X, data = pred)
# Dá pesos para os efeitos de blocos correspondente à LSmeans.
i <- attr(L, "assign") == 1
L[, i] <- 1/(sum(i) + 1)
# Obtém os valores preditos com IC.
ci <- confint(glht(m1, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
xyplot(mfpa ~ dose,
groups = biocar,
data = bio,
auto.key = list(points = FALSE,
lines = TRUE,
divide = 1,
type = "o")) +
as.layer(xyplot(Estimate ~ dose,
groups = biocar,
data = pred,
type = "l",
ly = pred$lwr,
uy = pred$upr,
cty = "bands",
alpha = 0.3,
prepanel = prepanel.cbH,
panel.groups = panel.cbH,
panel = panel.superpose))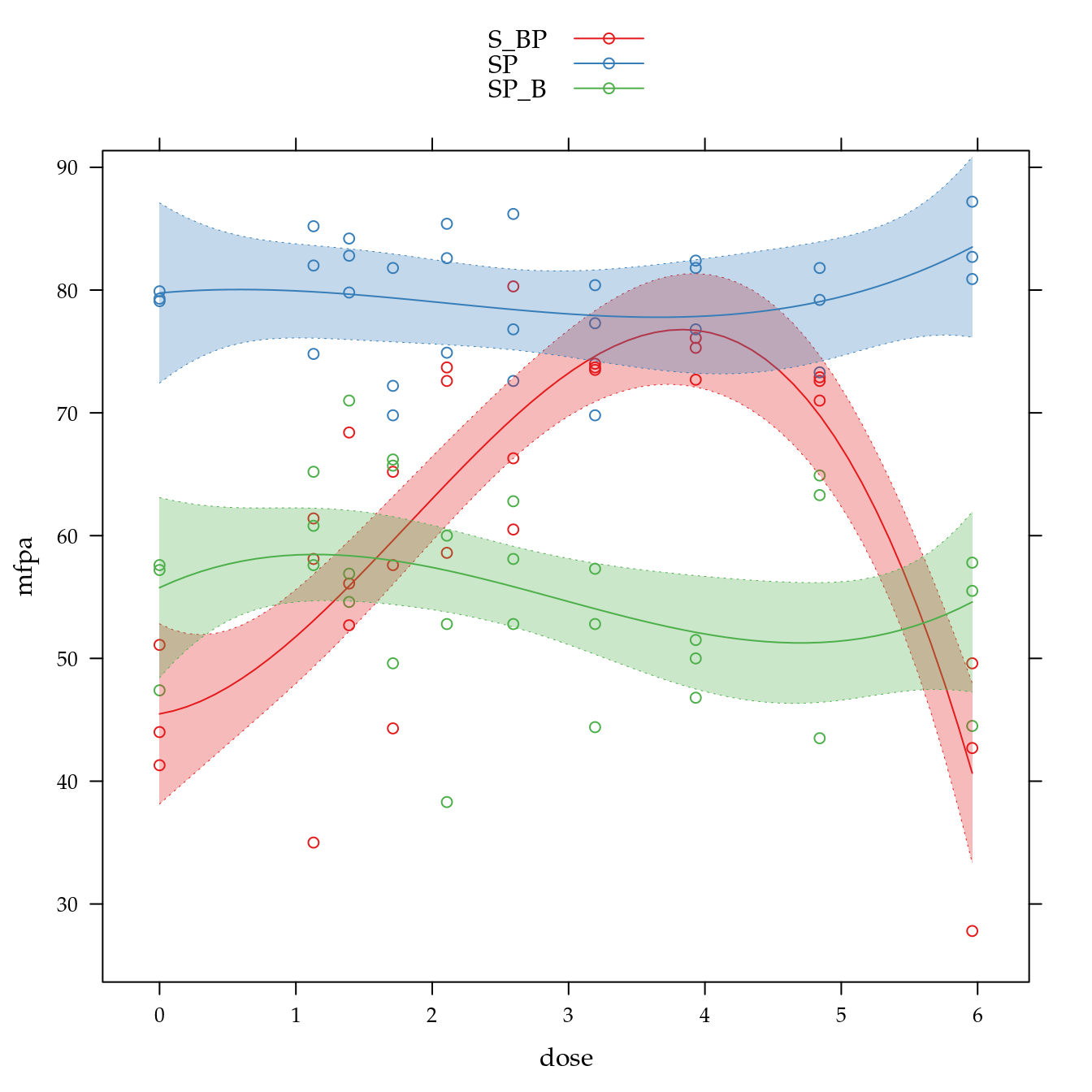
Massa fresca de raízes (mfr)
#-----------------------------------------------------------------------
# Modelo saturado (considera dose como qualitativo).
m0 <- lm(mfr ~ bloc + biocar * P, data = bio)
par(mfrow = c(2, 2))
plot(m0)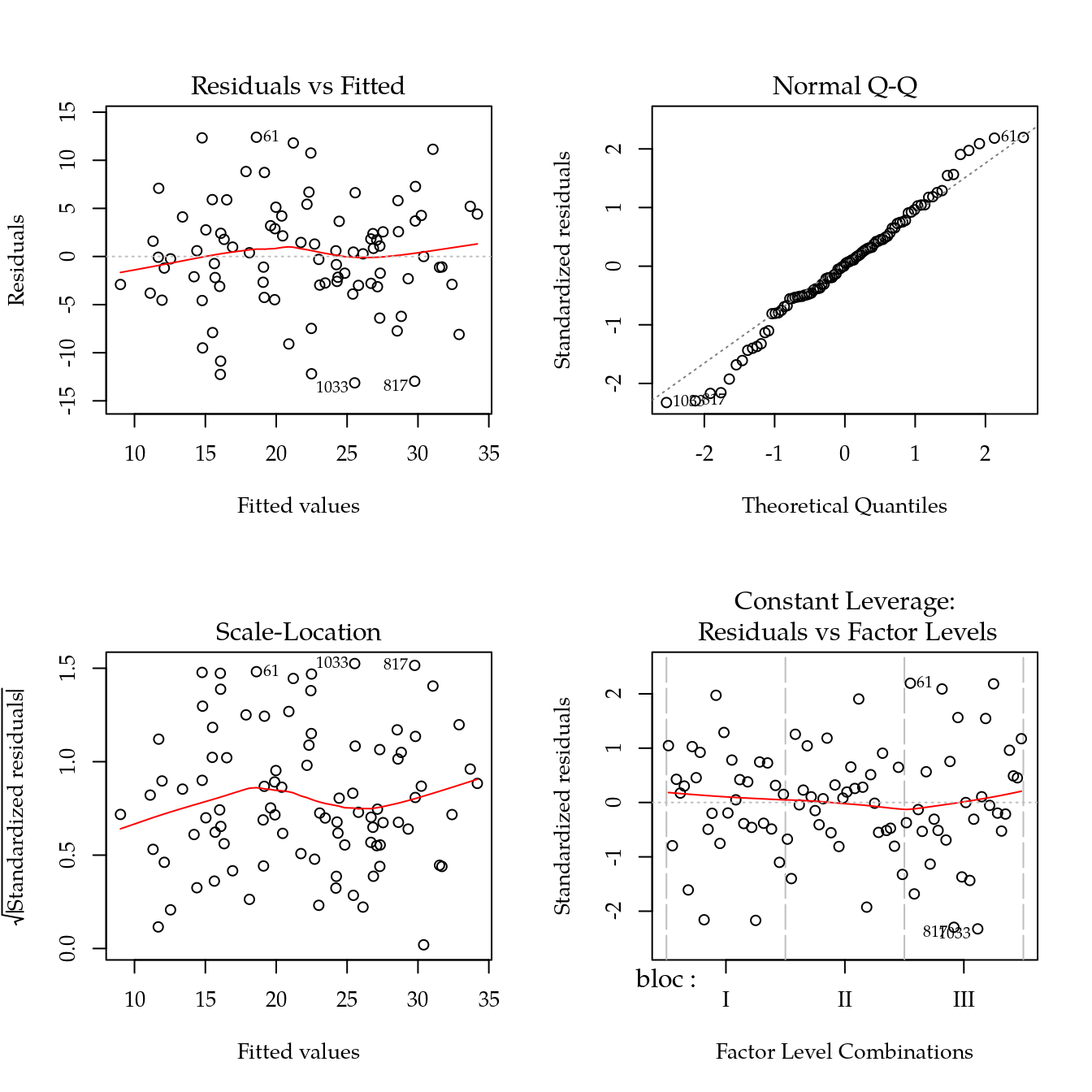
## Analysis of Variance Table
##
## Response: mfr
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 304.09 152.043 3.0715 0.053959 .
## biocar 2 549.18 274.590 5.5472 0.006245 **
## P 9 1093.90 121.544 2.4554 0.019148 *
## biocar:P 18 1558.27 86.570 1.7489 0.056187 .
## Residuals 58 2871.03 49.501
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1L <- LE_matrix(m0, effect = c("biocar", "P"))
pred <- equallevels(attr(L, "grid"), bio)
ci <- confint(glht(m0, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
str(pred)## 'data.frame': 30 obs. of 5 variables:
## $ biocar : Factor w/ 3 levels "S_BP","SP","SP_B": 1 2 3 1 2 3 1 2 3 1 ...
## $ P : Factor w/ 10 levels "0","1.5","3",..: 1 1 1 2 2 2 3 3 3 4 ...
## $ Estimate: num 13.6 24.8 18.6 18 29.6 ...
## $ lwr : num 5.47 16.67 10.44 9.87 21.5 ...
## $ upr : num 21.7 32.9 26.7 26.1 37.8 ...pch <- c(1, 4, 5)
key <- list(title = "Biocarvão",
cex.title = 1.1,
text = list(levels(bio$biocar)),
lines = list(pch = pch, lty = 1),
columns = 3,
type = "o",
divide = 1)
segplot(P ~ lwr + upr,
centers = Estimate,
data = pred,
draw = FALSE,
groups = biocar,
xlab = "Dose de P",
ylab = "Altura final (cm)",
key = key,
horizontal = FALSE,
pch = pch,
gap = 0.1,
panel = panel.groups.segplot)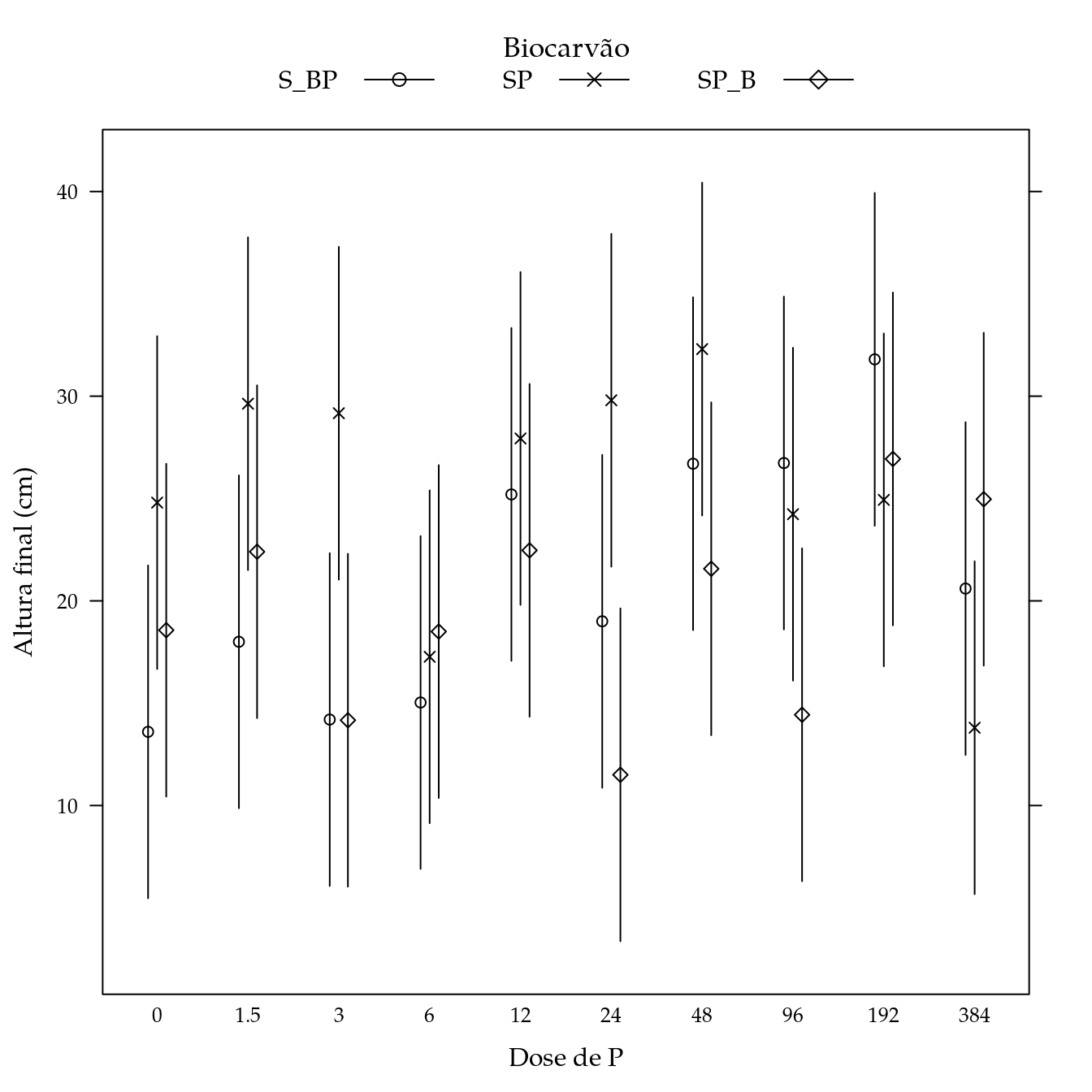
#-----------------------------------------------------------------------
# Modelo reduzido baseado em splines.
m1 <- update(m0, ~ bloc + biocar * bs(dose, df = 3))
par(mfrow = c(2, 2))
plot(m1)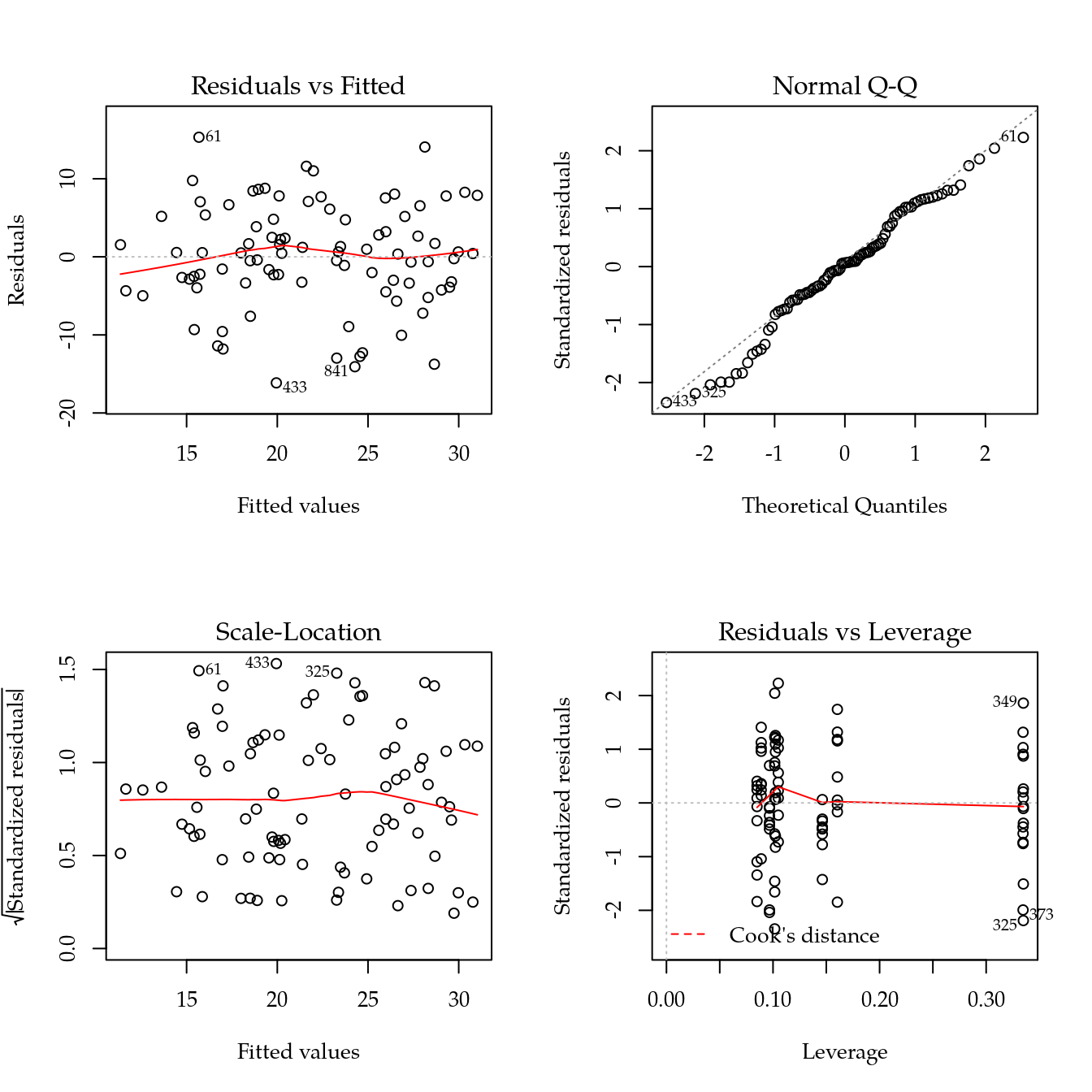
## Analysis of Variance Table
##
## Response: mfr
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 304.1 152.043 2.8826 0.062130 .
## biocar 2 549.2 274.590 5.2060 0.007605 **
## bs(dose, df = 3) 3 395.8 131.924 2.5012 0.065723 .
## biocar:bs(dose, df = 3) 6 1118.8 186.472 3.5354 0.003860 **
## Residuals 76 4008.6 52.745
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Teste para a falta de ajuste.
anova(m1, m0)## Analysis of Variance Table
##
## Model 1: mfr ~ bloc + biocar + bs(dose, df = 3) + biocar:bs(dose, df = 3)
## Model 2: mfr ~ bloc + biocar * P
## Res.Df RSS Df Sum of Sq F Pr(>F)
## 1 76 4008.6
## 2 58 2871.0 18 1137.6 1.2767 0.237# Novos valores em grid para fazer o gráfico de bandas.
pred <- with(bio,
expand.grid(bloc = factor(levels(bloc)[1],
levels = levels(bloc)),
biocar = levels(biocar),
dose = seq(min(dose),
max(dose),
length.out = 60)))
# Extrai a especificação do tempo base splines.
form <- attr(terms(formula(m1)), "term.labels")[3]
B <- eval(parse(text = form), envir = bio)
str(B)## 'bs' num [1:90, 1:3] 0 0 0 0.373 0.373 ...
## - attr(*, "dimnames")=List of 2
## ..$ : NULL
## ..$ : chr [1:3] "1" "2" "3"
## - attr(*, "degree")= int 3
## - attr(*, "knots")= num(0)
## - attr(*, "Boundary.knots")= num [1:2] 0 5.96
## - attr(*, "intercept")= logi FALSE# Gera a matriz para os valores de predição.
X <- predict(B, pred$dos)
# Cria a matriz do modelo para fazer a predição.
L <- model.matrix(~bloc + biocar * X, data = pred)
# Dá pesos para os efeitos de blocos correspondente à LSmeans.
i <- attr(L, "assign") == 1
L[, i] <- 1/(sum(i) + 1)
# Obtém os valores preditos com IC.
ci <- confint(glht(m1, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
xyplot(mfr ~ dose,
groups = biocar,
data = bio,
auto.key = list(points = FALSE,
lines = TRUE,
divide = 1,
type = "o")) +
as.layer(xyplot(Estimate ~ dose,
groups = biocar,
data = pred,
type = "l",
ly = pred$lwr,
uy = pred$upr,
cty = "bands",
alpha = 0.3,
prepanel = prepanel.cbH,
panel.groups = panel.cbH,
panel = panel.superpose))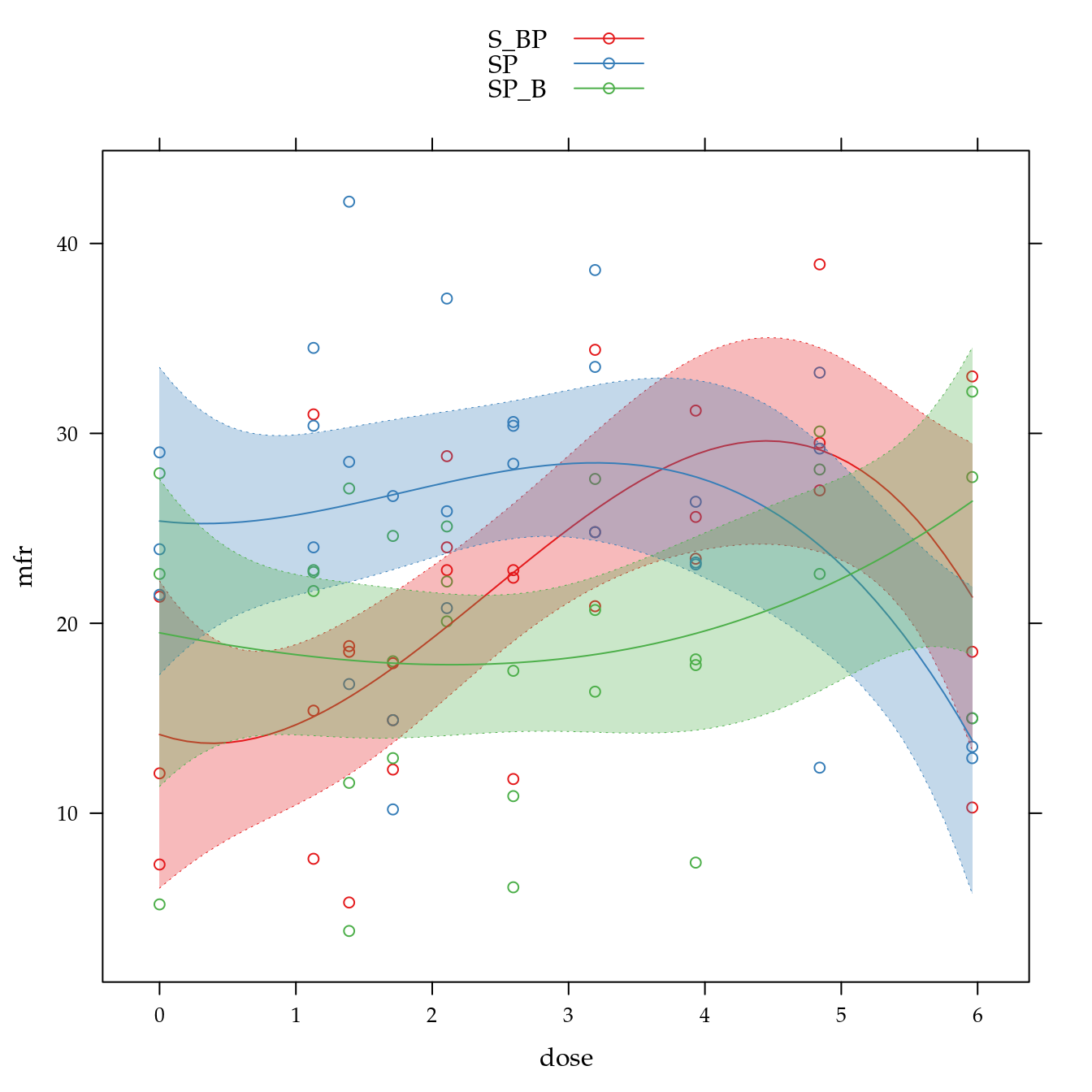
Massa seca de parte aérea (mspa)
#-----------------------------------------------------------------------
# Modelo saturado (considera dose como qualitativo).
m0 <- lm(mspa ~ bloc + biocar * P, data = bio)
par(mfrow = c(2, 2))
plot(m0)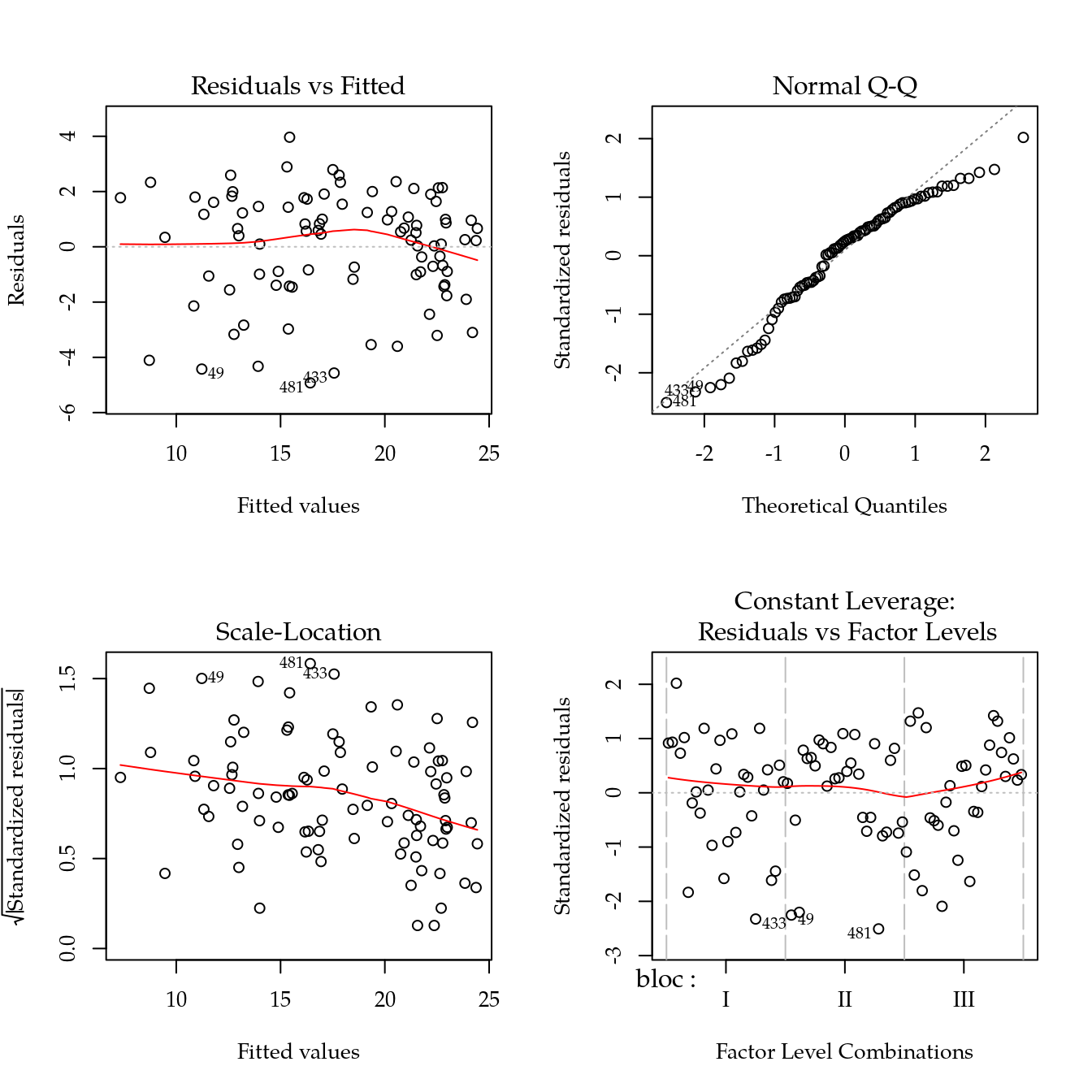
## Analysis of Variance Table
##
## Response: mspa
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 40.00 20.00 3.3432 0.042251 *
## biocar 2 1028.34 514.17 85.9398 < 2.2e-16 ***
## P 9 165.55 18.39 3.0744 0.004423 **
## biocar:P 18 533.18 29.62 4.9510 1.593e-06 ***
## Residuals 58 347.01 5.98
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1L <- LE_matrix(m0, effect = c("biocar", "P"))
pred <- equallevels(attr(L, "grid"), bio)
ci <- confint(glht(m0, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
str(pred)## 'data.frame': 30 obs. of 5 variables:
## $ biocar : Factor w/ 3 levels "S_BP","SP","SP_B": 1 2 3 1 2 3 1 2 3 1 ...
## $ P : Factor w/ 10 levels "0","1.5","3",..: 1 1 1 2 2 2 3 3 3 4 ...
## $ Estimate: num 10.4 22.2 15.7 12.2 23.4 ...
## $ lwr : num 7.57 19.37 12.91 9.34 20.57 ...
## $ upr : num 13.2 25 18.6 15 26.2 ...pch <- c(1, 4, 5)
key <- list(title = "Biocarvão",
cex.title = 1.1,
text = list(levels(bio$biocar)),
lines = list(pch = pch, lty = 1),
columns = 3,
type = "o",
divide = 1)
segplot(P ~ lwr + upr,
centers = Estimate,
data = pred,
draw = FALSE,
groups = biocar,
xlab = "Dose de P",
ylab = "Altura final (cm)",
key = key,
horizontal = FALSE,
pch = pch,
gap = 0.1,
panel = panel.groups.segplot)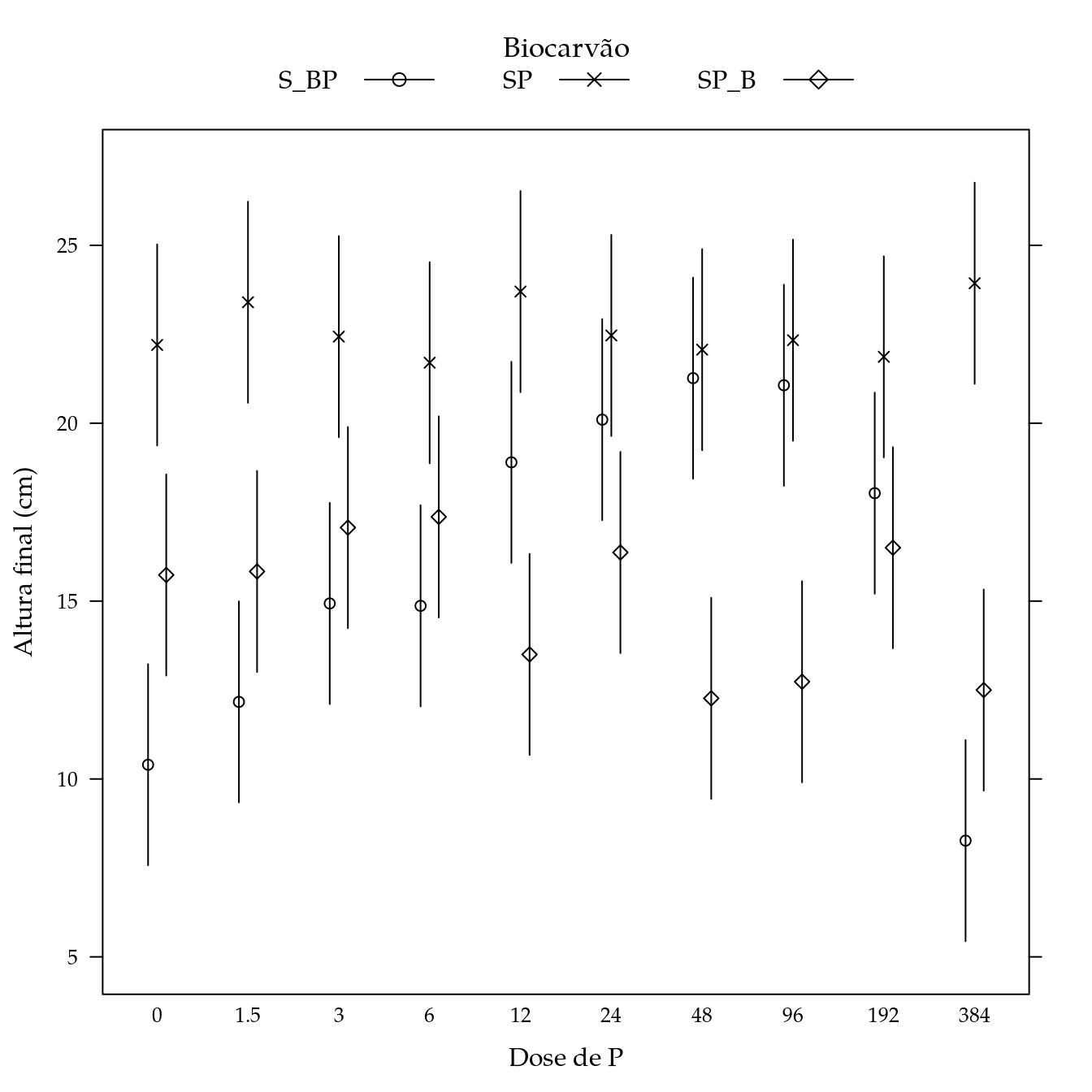
#-----------------------------------------------------------------------
# Modelo reduzido baseado em splines.
m1 <- update(m0, ~ bloc + biocar * bs(dose, df = 3))
par(mfrow = c(2, 2))
plot(m1)
## Analysis of Variance Table
##
## Response: mspa
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 40.00 20.00 3.3486 0.04039 *
## biocar 2 1028.34 514.17 86.0793 < 2.2e-16 ***
## bs(dose, df = 3) 3 148.04 49.35 8.2613 7.947e-05 ***
## biocar:bs(dose, df = 3) 6 443.73 73.96 12.3812 1.124e-09 ***
## Residuals 76 453.96 5.97
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Teste para a falta de ajuste.
anova(m1, m0)## Analysis of Variance Table
##
## Model 1: mspa ~ bloc + biocar + bs(dose, df = 3) + biocar:bs(dose, df = 3)
## Model 2: mspa ~ bloc + biocar * P
## Res.Df RSS Df Sum of Sq F Pr(>F)
## 1 76 453.96
## 2 58 347.01 18 106.96 0.9932 0.4803# Novos valores em grid para fazer o gráfico de bandas.
pred <- with(bio,
expand.grid(bloc = factor(levels(bloc)[1],
levels = levels(bloc)),
biocar = levels(biocar),
dose = seq(min(dose),
max(dose),
length.out = 60)))
# Extrai a especificação do tempo base splines.
form <- attr(terms(formula(m1)), "term.labels")[3]
B <- eval(parse(text = form), envir = bio)
str(B)## 'bs' num [1:90, 1:3] 0 0 0 0.373 0.373 ...
## - attr(*, "dimnames")=List of 2
## ..$ : NULL
## ..$ : chr [1:3] "1" "2" "3"
## - attr(*, "degree")= int 3
## - attr(*, "knots")= num(0)
## - attr(*, "Boundary.knots")= num [1:2] 0 5.96
## - attr(*, "intercept")= logi FALSE# Gera a matriz para os valores de predição.
X <- predict(B, pred$dos)
# Cria a matriz do modelo para fazer a predição.
L <- model.matrix(~bloc + biocar * X, data = pred)
# Dá pesos para os efeitos de blocos correspondente à LSmeans.
i <- attr(L, "assign") == 1
L[, i] <- 1/(sum(i) + 1)
# Obtém os valores preditos com IC.
ci <- confint(glht(m1, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
xyplot(mspa ~ dose,
groups = biocar,
data = bio,
auto.key = list(points = FALSE,
lines = TRUE,
divide = 1,
type = "o")) +
as.layer(xyplot(Estimate ~ dose,
groups = biocar,
data = pred,
type = "l",
ly = pred$lwr,
uy = pred$upr,
cty = "bands",
alpha = 0.3,
prepanel = prepanel.cbH,
panel.groups = panel.cbH,
panel = panel.superpose))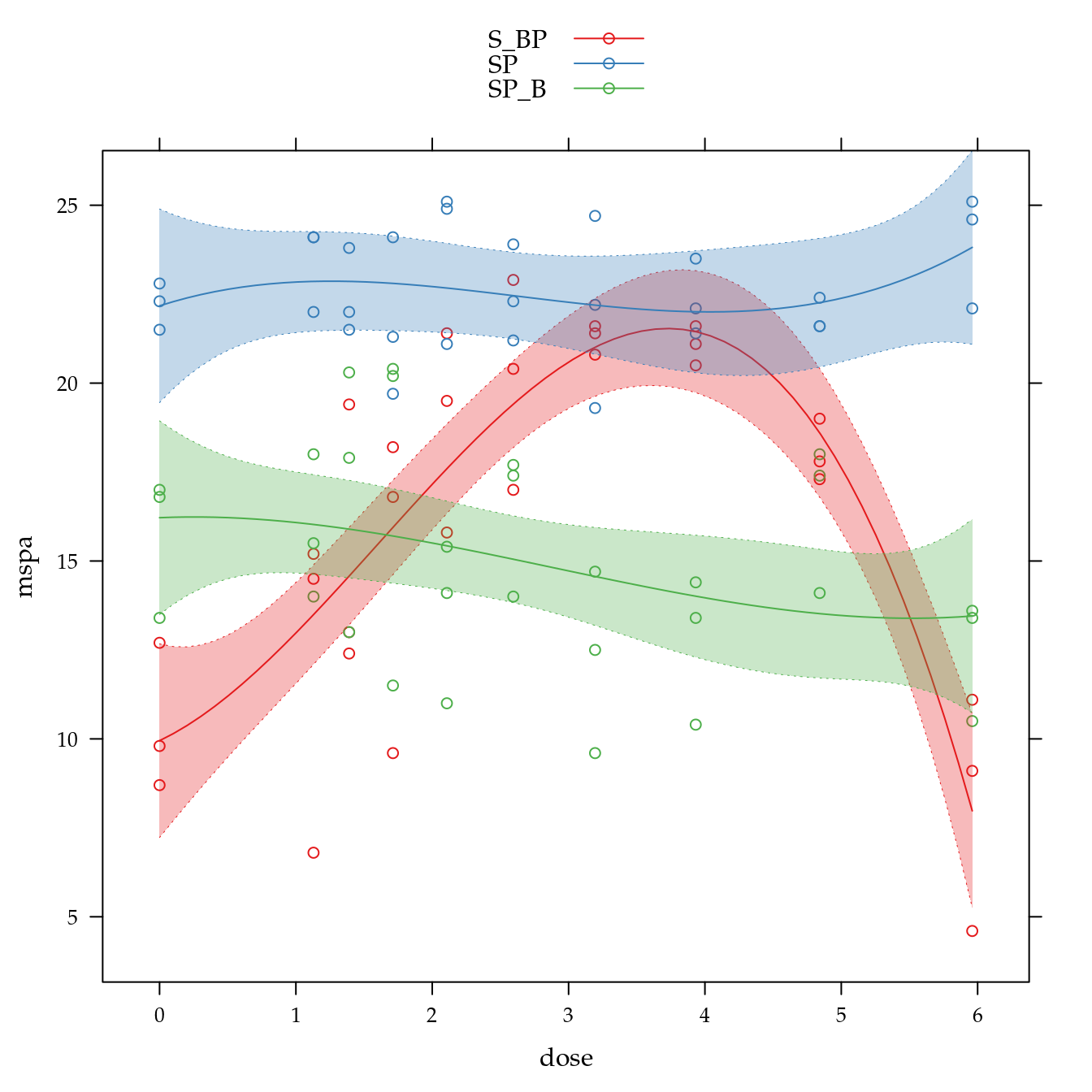
Massa seca de raízes (msr)
#-----------------------------------------------------------------------
# Modelo saturado (considera dose como qualitativo).
m0 <- lm(msr ~ bloc + biocar * P, data = bio)
par(mfrow = c(2, 2))
plot(m0)
## Analysis of Variance Table
##
## Response: msr
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 23.75 11.8748 1.4072 0.253051
## biocar 2 62.55 31.2748 3.7063 0.030566 *
## P 9 247.60 27.5107 3.2602 0.002855 **
## biocar:P 18 362.45 20.1360 2.3863 0.006532 **
## Residuals 58 489.42 8.4383
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1L <- LE_matrix(m0, effect = c("biocar", "P"))
pred <- equallevels(attr(L, "grid"), bio)
ci <- confint(glht(m0, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
str(pred)## 'data.frame': 30 obs. of 5 variables:
## $ biocar : Factor w/ 3 levels "S_BP","SP","SP_B": 1 2 3 1 2 3 1 2 3 1 ...
## $ P : Factor w/ 10 levels "0","1.5","3",..: 1 1 1 2 2 2 3 3 3 4 ...
## $ Estimate: num 7.53 11.43 10.17 7.77 14.47 ...
## $ lwr : num 4.18 8.08 6.81 4.41 11.11 ...
## $ upr : num 10.9 14.8 13.5 11.1 17.8 ...pch <- c(1, 4, 5)
key <- list(title = "Biocarvão",
cex.title = 1.1,
text = list(levels(bio$biocar)),
lines = list(pch = pch, lty = 1),
columns = 3,
type = "o",
divide = 1)
segplot(P ~ lwr + upr,
centers = Estimate,
data = pred,
draw = FALSE,
groups = biocar,
xlab = "Dose de P",
ylab = "Altura final (cm)",
key = key,
horizontal = FALSE,
pch = pch,
gap = 0.1,
panel = panel.groups.segplot)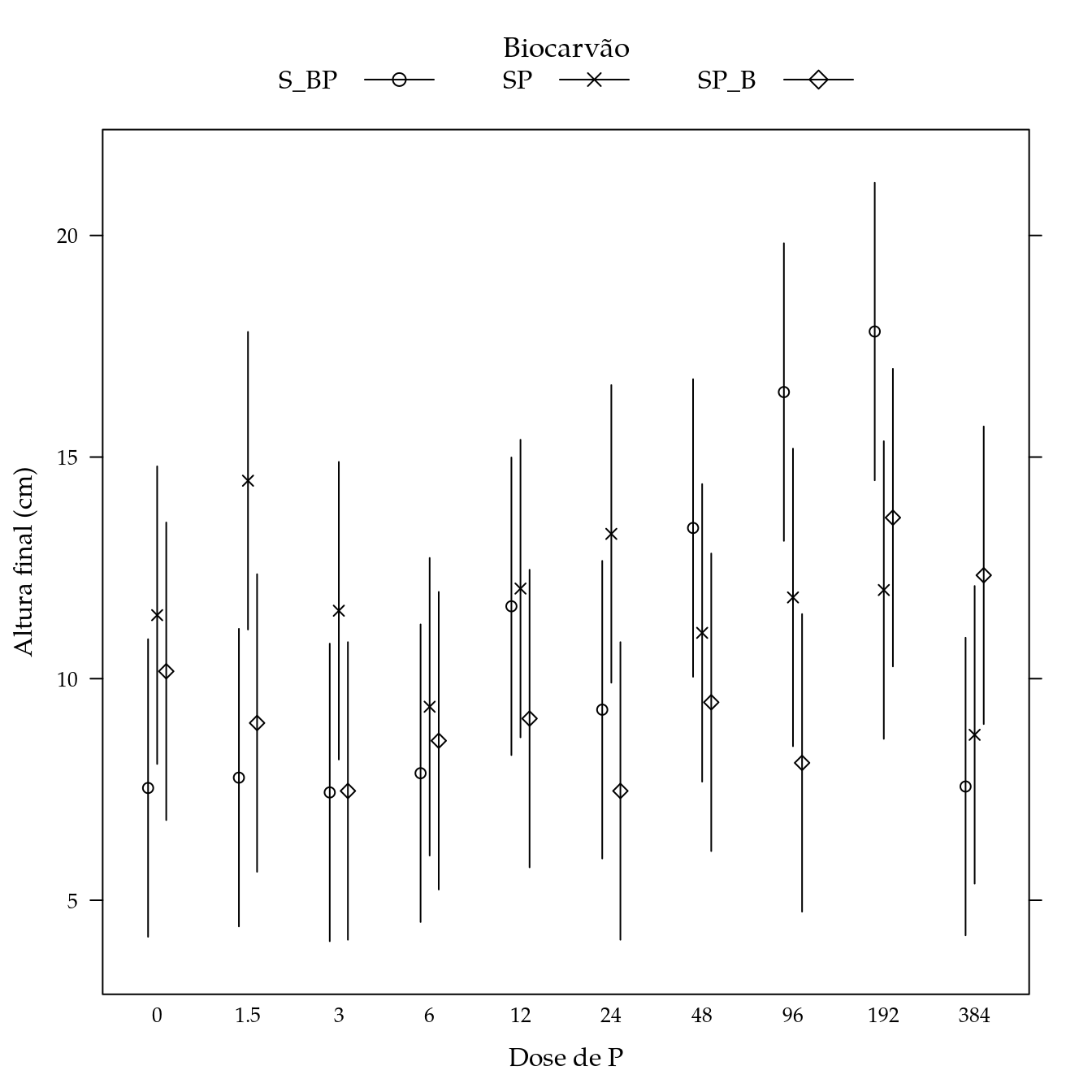
#-----------------------------------------------------------------------
# Modelo reduzido baseado em splines.
m1 <- update(m0, ~ bloc + biocar * bs(dose, df = 3))
par(mfrow = c(2, 2))
plot(m1)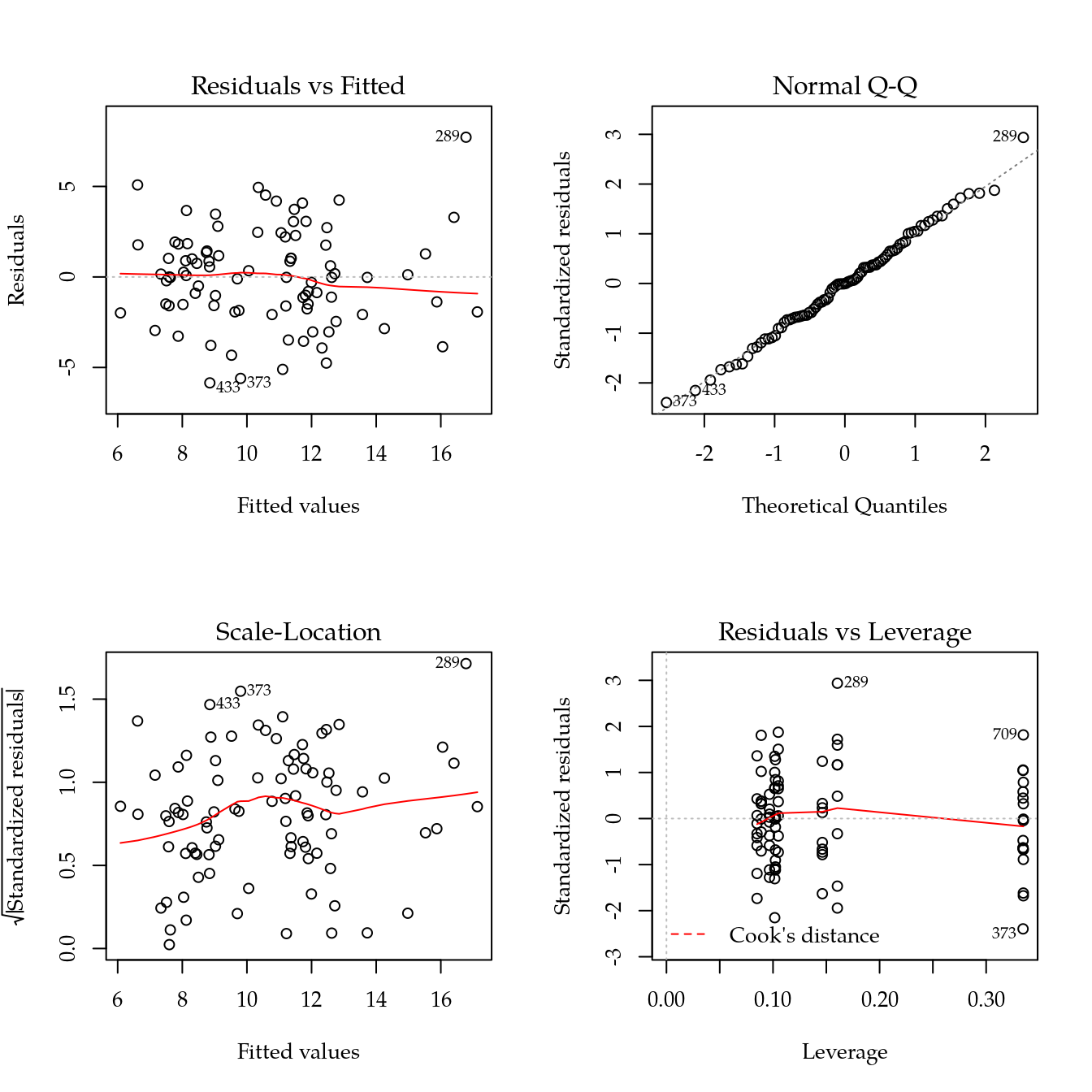
## Analysis of Variance Table
##
## Response: msr
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 23.75 11.875 1.4434 0.2425327
## biocar 2 62.55 31.275 3.8014 0.0267012 *
## bs(dose, df = 3) 3 175.22 58.408 7.0993 0.0002873 ***
## biocar:bs(dose, df = 3) 6 298.98 49.830 6.0567 3.227e-05 ***
## Residuals 76 625.27 8.227
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Teste para a falta de ajuste.
anova(m1, m0)## Analysis of Variance Table
##
## Model 1: msr ~ bloc + biocar + bs(dose, df = 3) + biocar:bs(dose, df = 3)
## Model 2: msr ~ bloc + biocar * P
## Res.Df RSS Df Sum of Sq F Pr(>F)
## 1 76 625.27
## 2 58 489.42 18 135.84 0.8944 0.5872# Novos valores em grid para fazer o gráfico de bandas.
pred <- with(bio,
expand.grid(bloc = factor(levels(bloc)[1],
levels = levels(bloc)),
biocar = levels(biocar),
dose = seq(min(dose),
max(dose),
length.out = 60)))
# Extrai a especificação do tempo base splines.
form <- attr(terms(formula(m1)), "term.labels")[3]
B <- eval(parse(text = form), envir = bio)
str(B)## 'bs' num [1:90, 1:3] 0 0 0 0.373 0.373 ...
## - attr(*, "dimnames")=List of 2
## ..$ : NULL
## ..$ : chr [1:3] "1" "2" "3"
## - attr(*, "degree")= int 3
## - attr(*, "knots")= num(0)
## - attr(*, "Boundary.knots")= num [1:2] 0 5.96
## - attr(*, "intercept")= logi FALSE# Gera a matriz para os valores de predição.
X <- predict(B, pred$dos)
# Cria a matriz do modelo para fazer a predição.
L <- model.matrix(~bloc + biocar * X, data = pred)
# Dá pesos para os efeitos de blocos correspondente à LSmeans.
i <- attr(L, "assign") == 1
L[, i] <- 1/(sum(i) + 1)
# Obtém os valores preditos com IC.
ci <- confint(glht(m1, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
xyplot(msr ~ dose,
groups = biocar,
data = bio,
auto.key = list(points = FALSE,
lines = TRUE,
divide = 1,
type = "o")) +
as.layer(xyplot(Estimate ~ dose,
groups = biocar,
data = pred,
type = "l",
ly = pred$lwr,
uy = pred$upr,
cty = "bands",
alpha = 0.3,
prepanel = prepanel.cbH,
panel.groups = panel.cbH,
panel = panel.superpose))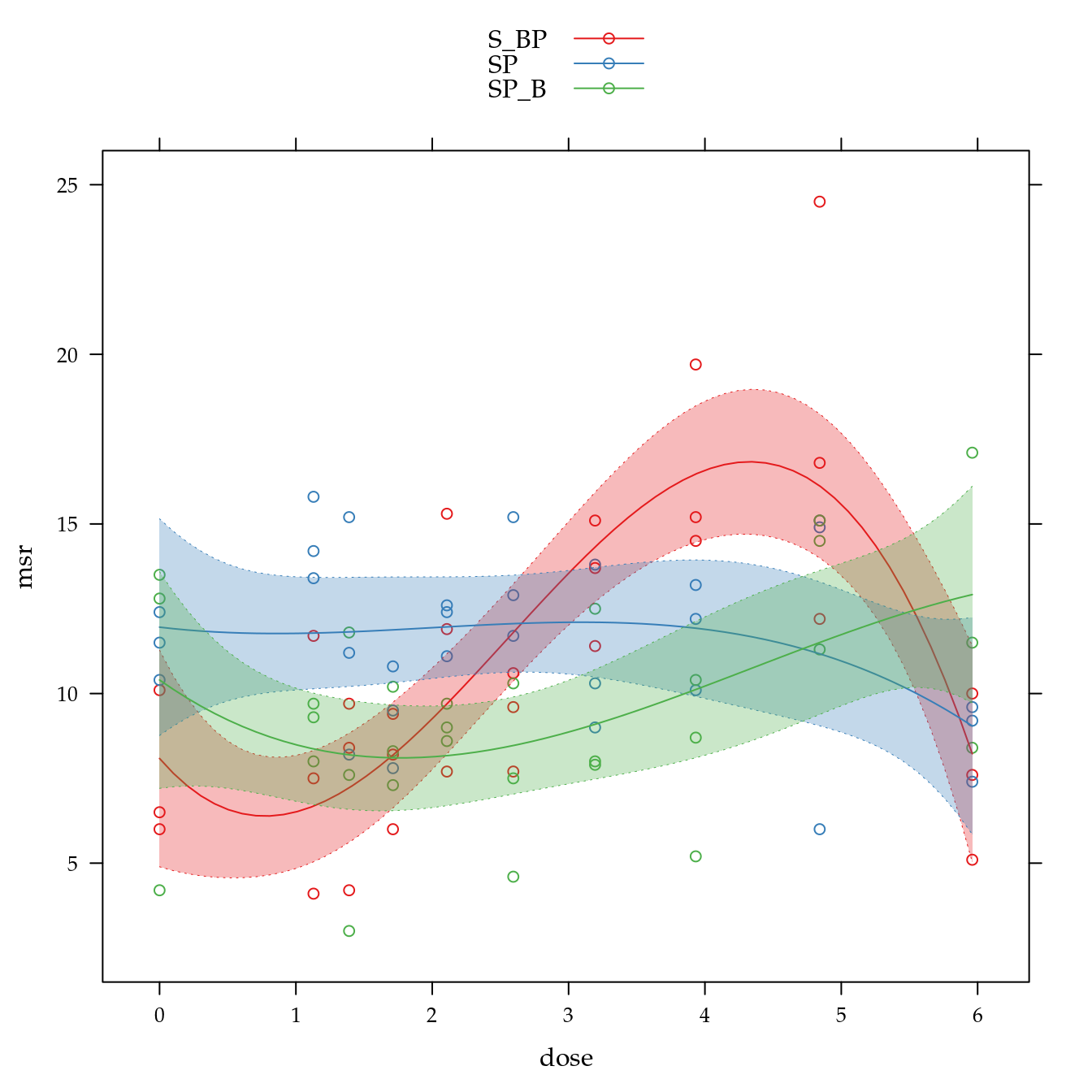
pH em água (ph)
#-----------------------------------------------------------------------
# Modelo saturado (considera dose como qualitativo).
m0 <- lm(ph ~ bloc + biocar * P, data = bio)
par(mfrow = c(2, 2))
plot(m0)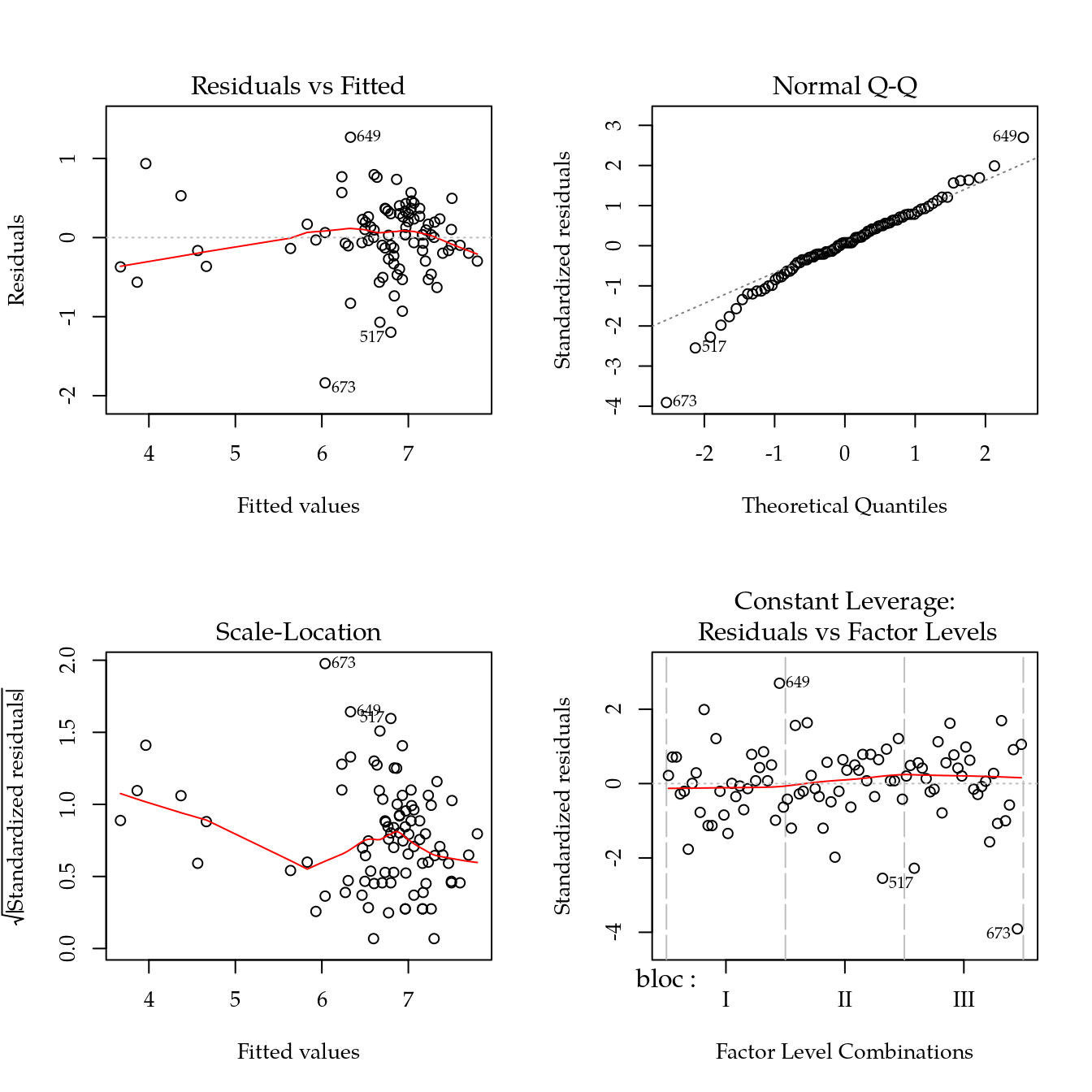
## Analysis of Variance Table
##
## Response: ph
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 1.3342 0.6671 1.9444 0.1522823
## biocar 2 12.1442 6.0721 17.6984 9.995e-07 ***
## P 9 20.5512 2.2835 6.6556 1.786e-06 ***
## biocar:P 18 21.7024 1.2057 3.5142 0.0001424 ***
## Residuals 58 19.8991 0.3431
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1L <- LE_matrix(m0, effect = c("biocar", "P"))
pred <- equallevels(attr(L, "grid"), bio)
ci <- confint(glht(m0, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
str(pred)## 'data.frame': 30 obs. of 5 variables:
## $ biocar : Factor w/ 3 levels "S_BP","SP","SP_B": 1 2 3 1 2 3 1 2 3 1 ...
## $ P : Factor w/ 10 levels "0","1.5","3",..: 1 1 1 2 2 2 3 3 3 4 ...
## $ Estimate: num 7.37 7.1 6.93 6.63 6.8 ...
## $ lwr : num 6.69 6.42 6.26 5.96 6.12 ...
## $ upr : num 8.04 7.78 7.61 7.31 7.48 ...pch <- c(1, 4, 5)
key <- list(title = "Biocarvão",
cex.title = 1.1,
text = list(levels(bio$biocar)),
lines = list(pch = pch, lty = 1),
columns = 3,
type = "o",
divide = 1)
segplot(P ~ lwr + upr,
centers = Estimate,
data = pred,
draw = FALSE,
groups = biocar,
xlab = "Dose de P",
ylab = "Altura final (cm)",
key = key,
horizontal = FALSE,
pch = pch,
gap = 0.1,
panel = panel.groups.segplot)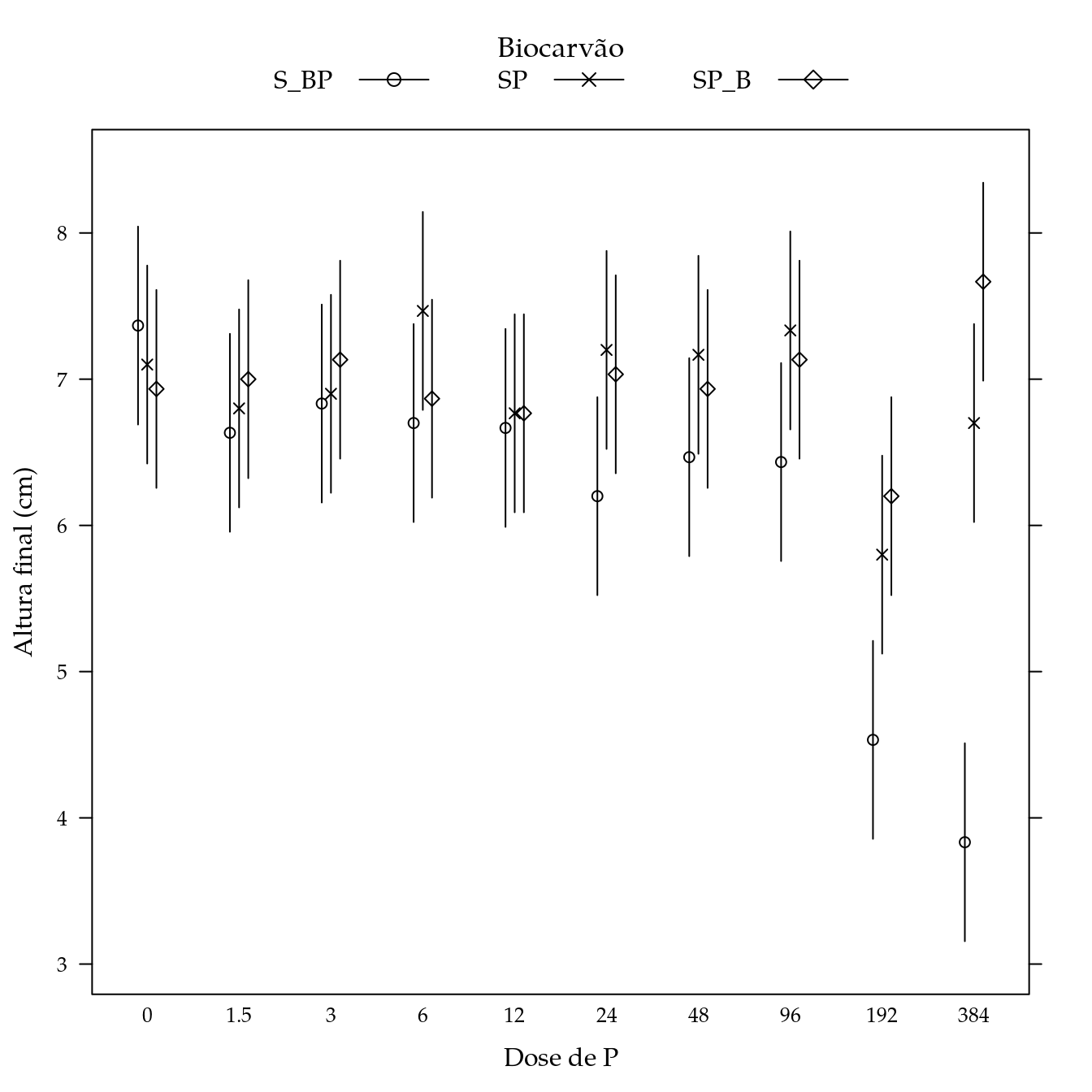
#-----------------------------------------------------------------------
# Modelo reduzido baseado em splines.
m1 <- update(m0, ~ bloc + biocar * bs(dose, df = 3))
par(mfrow = c(2, 2))
plot(m1)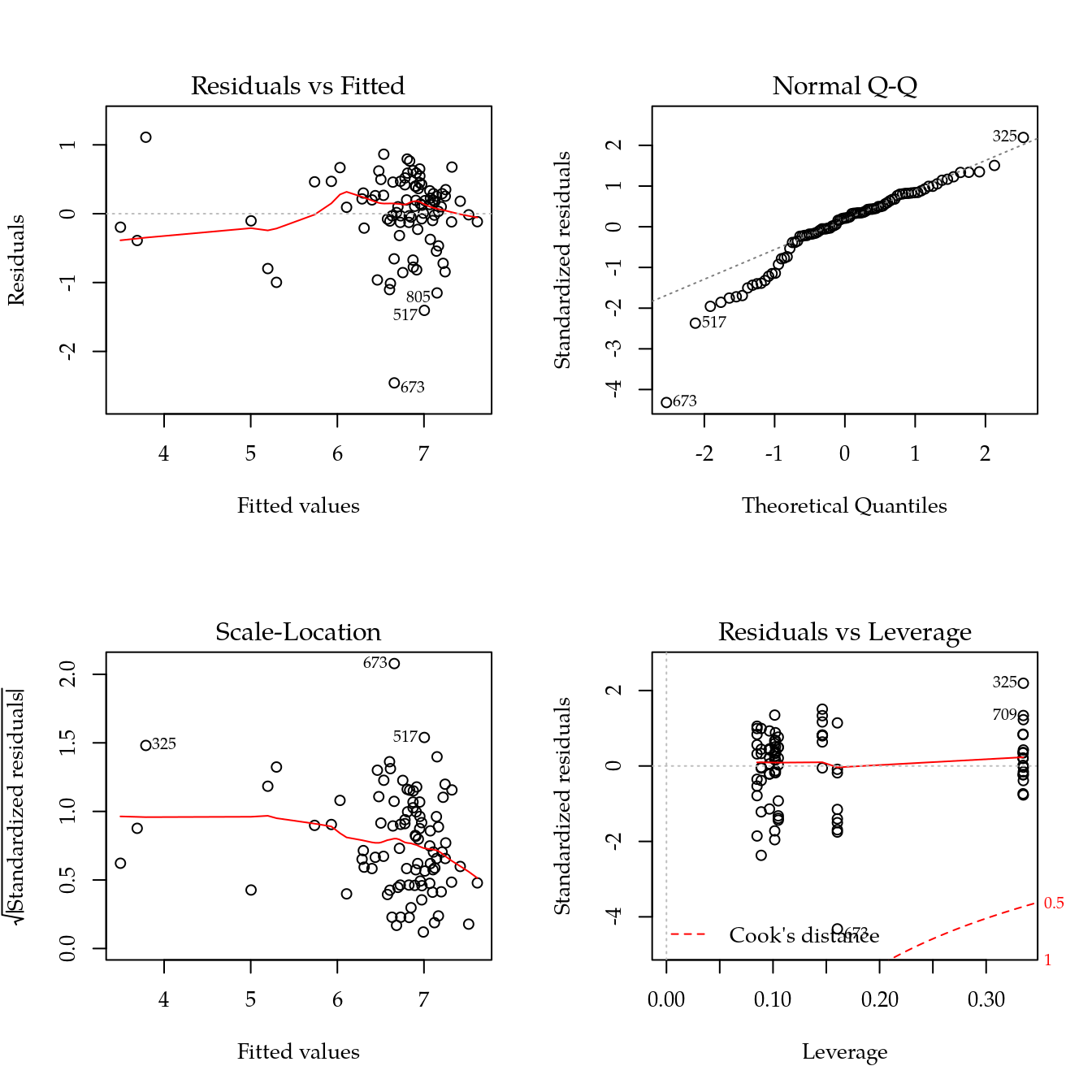
## Analysis of Variance Table
##
## Response: ph
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 1.3342 0.6671 1.7308 0.184
## biocar 2 12.1442 6.0721 15.7541 1.889e-06 ***
## bs(dose, df = 3) 3 12.5601 4.1867 10.8624 5.117e-06 ***
## biocar:bs(dose, df = 3) 6 20.3000 3.3833 8.7780 2.880e-07 ***
## Residuals 76 29.2927 0.3854
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Teste para a falta de ajuste.
anova(m1, m0)## Analysis of Variance Table
##
## Model 1: ph ~ bloc + biocar + bs(dose, df = 3) + biocar:bs(dose, df = 3)
## Model 2: ph ~ bloc + biocar * P
## Res.Df RSS Df Sum of Sq F Pr(>F)
## 1 76 29.293
## 2 58 19.899 18 9.3936 1.5211 0.1158# Novos valores em grid para fazer o gráfico de bandas.
pred <- with(bio,
expand.grid(bloc = factor(levels(bloc)[1],
levels = levels(bloc)),
biocar = levels(biocar),
dose = seq(min(dose),
max(dose),
length.out = 60)))
# Extrai a especificação do tempo base splines.
form <- attr(terms(formula(m1)), "term.labels")[3]
B <- eval(parse(text = form), envir = bio)
str(B)## 'bs' num [1:90, 1:3] 0 0 0 0.373 0.373 ...
## - attr(*, "dimnames")=List of 2
## ..$ : NULL
## ..$ : chr [1:3] "1" "2" "3"
## - attr(*, "degree")= int 3
## - attr(*, "knots")= num(0)
## - attr(*, "Boundary.knots")= num [1:2] 0 5.96
## - attr(*, "intercept")= logi FALSE# Gera a matriz para os valores de predição.
X <- predict(B, pred$dos)
# Cria a matriz do modelo para fazer a predição.
L <- model.matrix(~bloc + biocar * X, data = pred)
# Dá pesos para os efeitos de blocos correspondente à LSmeans.
i <- attr(L, "assign") == 1
L[, i] <- 1/(sum(i) + 1)
# Obtém os valores preditos com IC.
ci <- confint(glht(m1, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
xyplot(ph ~ dose,
groups = biocar,
data = bio,
auto.key = list(points = FALSE,
lines = TRUE,
divide = 1,
type = "o")) +
as.layer(xyplot(Estimate ~ dose,
groups = biocar,
data = pred,
type = "l",
ly = pred$lwr,
uy = pred$upr,
cty = "bands",
alpha = 0.3,
prepanel = prepanel.cbH,
panel.groups = panel.cbH,
panel = panel.superpose))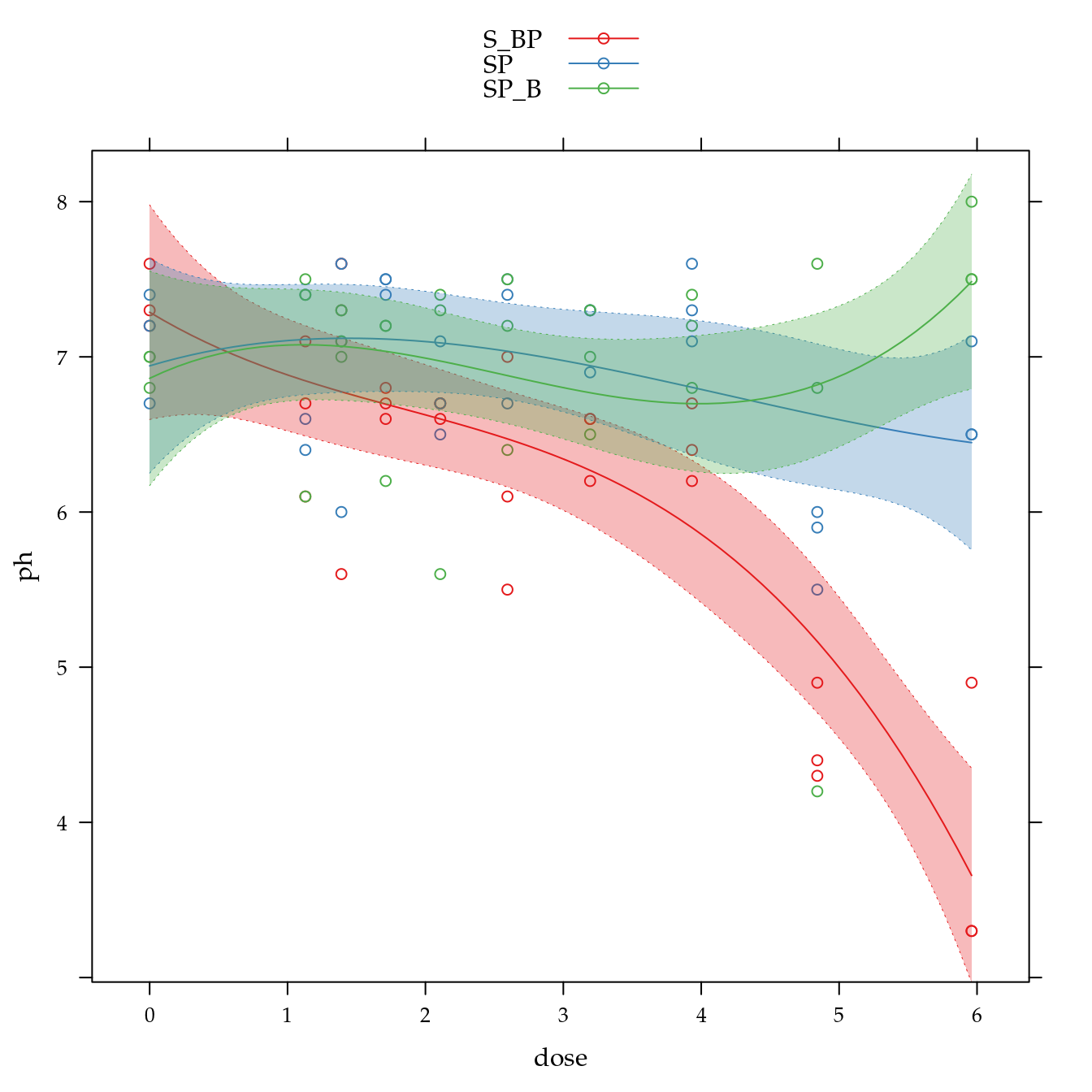
pH em KCl (phkcl)
#-----------------------------------------------------------------------
# Modelo saturado (considera dose como qualitativo).
m0 <- lm(phkcl ~ bloc + biocar * P, data = bio)
par(mfrow = c(2, 2))
plot(m0)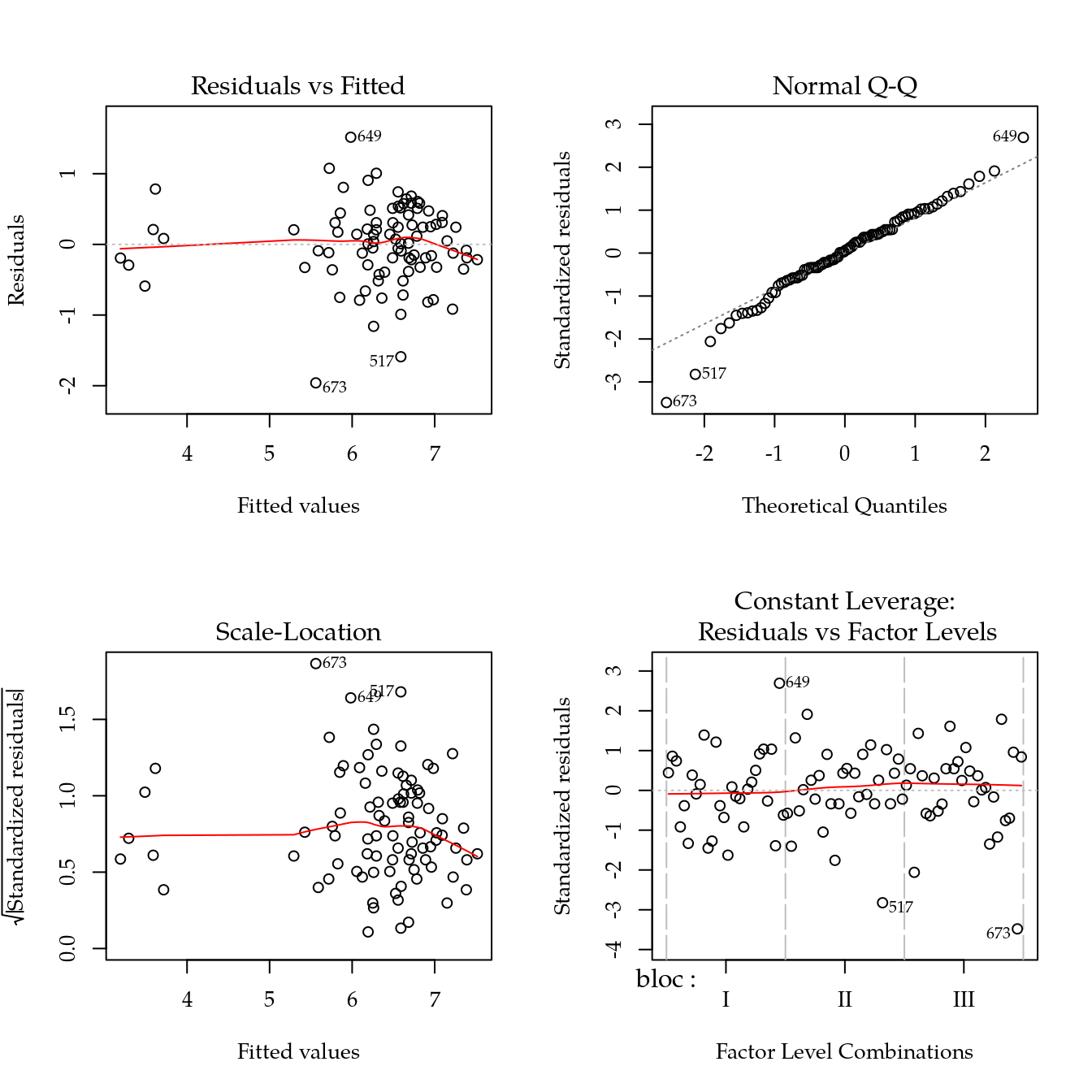
## Analysis of Variance Table
##
## Response: phkcl
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 2.8327 1.4163 2.8776 0.064338 .
## biocar 2 18.5447 9.2723 18.8387 4.965e-07 ***
## P 9 25.2849 2.8094 5.7080 1.221e-05 ***
## biocar:P 18 23.6864 1.3159 2.6736 0.002433 **
## Residuals 58 28.5473 0.4922
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1L <- LE_matrix(m0, effect = c("biocar", "P"))
pred <- equallevels(attr(L, "grid"), bio)
ci <- confint(glht(m0, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
str(pred)## 'data.frame': 30 obs. of 5 variables:
## $ biocar : Factor w/ 3 levels "S_BP","SP","SP_B": 1 2 3 1 2 3 1 2 3 1 ...
## $ P : Factor w/ 10 levels "0","1.5","3",..: 1 1 1 2 2 2 3 3 3 4 ...
## $ Estimate: num 6.77 6.73 6.5 6.03 6.43 ...
## $ lwr : num 5.96 5.92 5.69 5.22 5.62 ...
## $ upr : num 7.58 7.54 7.31 6.84 7.24 ...pch <- c(1, 4, 5)
key <- list(title = "Biocarvão",
cex.title = 1.1,
text = list(levels(bio$biocar)),
lines = list(pch = pch, lty = 1),
columns = 3,
type = "o",
divide = 1)
segplot(P ~ lwr + upr,
centers = Estimate,
data = pred,
draw = FALSE,
groups = biocar,
xlab = "Dose de P",
ylab = "Altura final (cm)",
key = key,
horizontal = FALSE,
pch = pch,
gap = 0.1,
panel = panel.groups.segplot)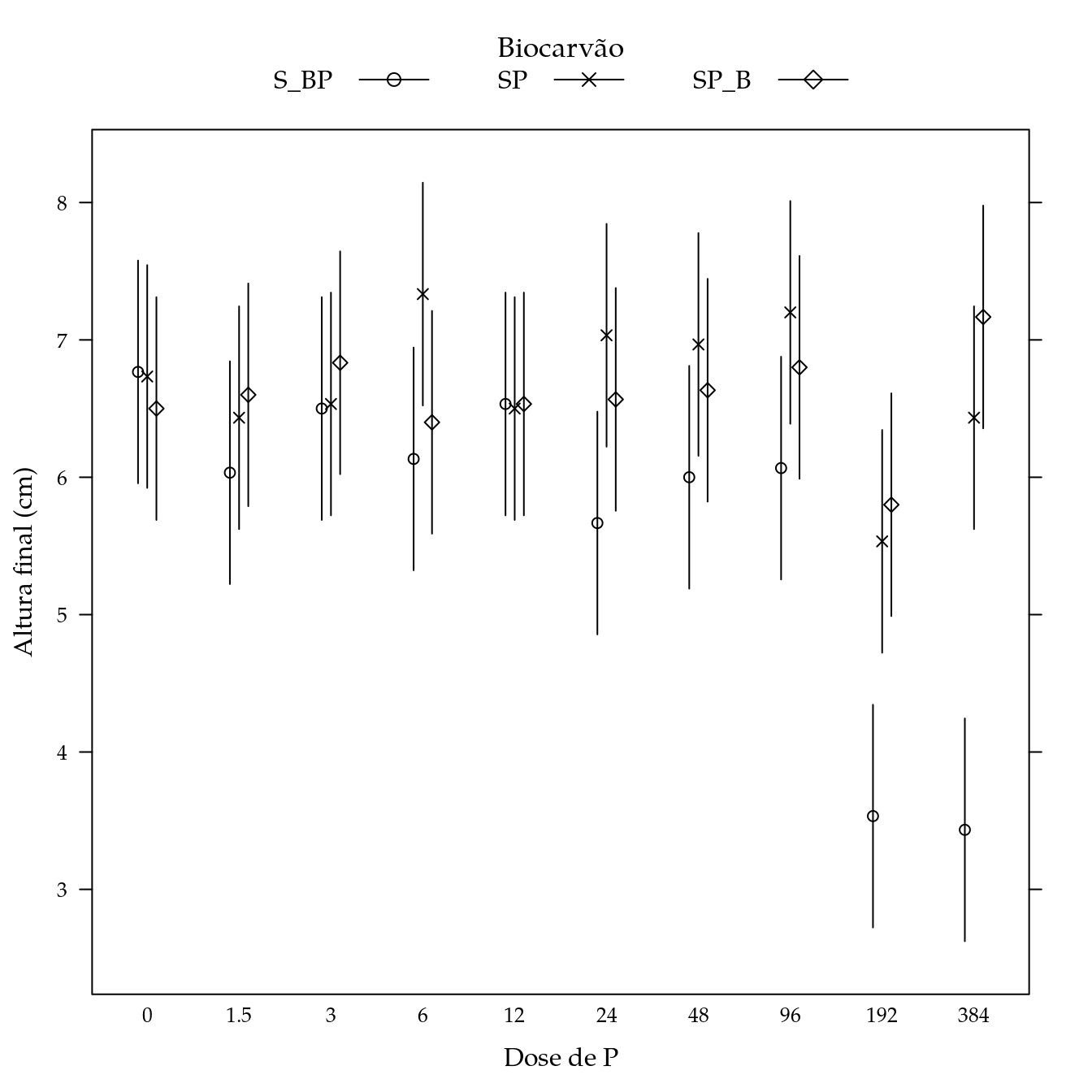
#-----------------------------------------------------------------------
# Modelo reduzido baseado em splines.
m1 <- update(m0, ~ bloc + biocar * bs(dose, df = 3))
par(mfrow = c(2, 2))
plot(m1)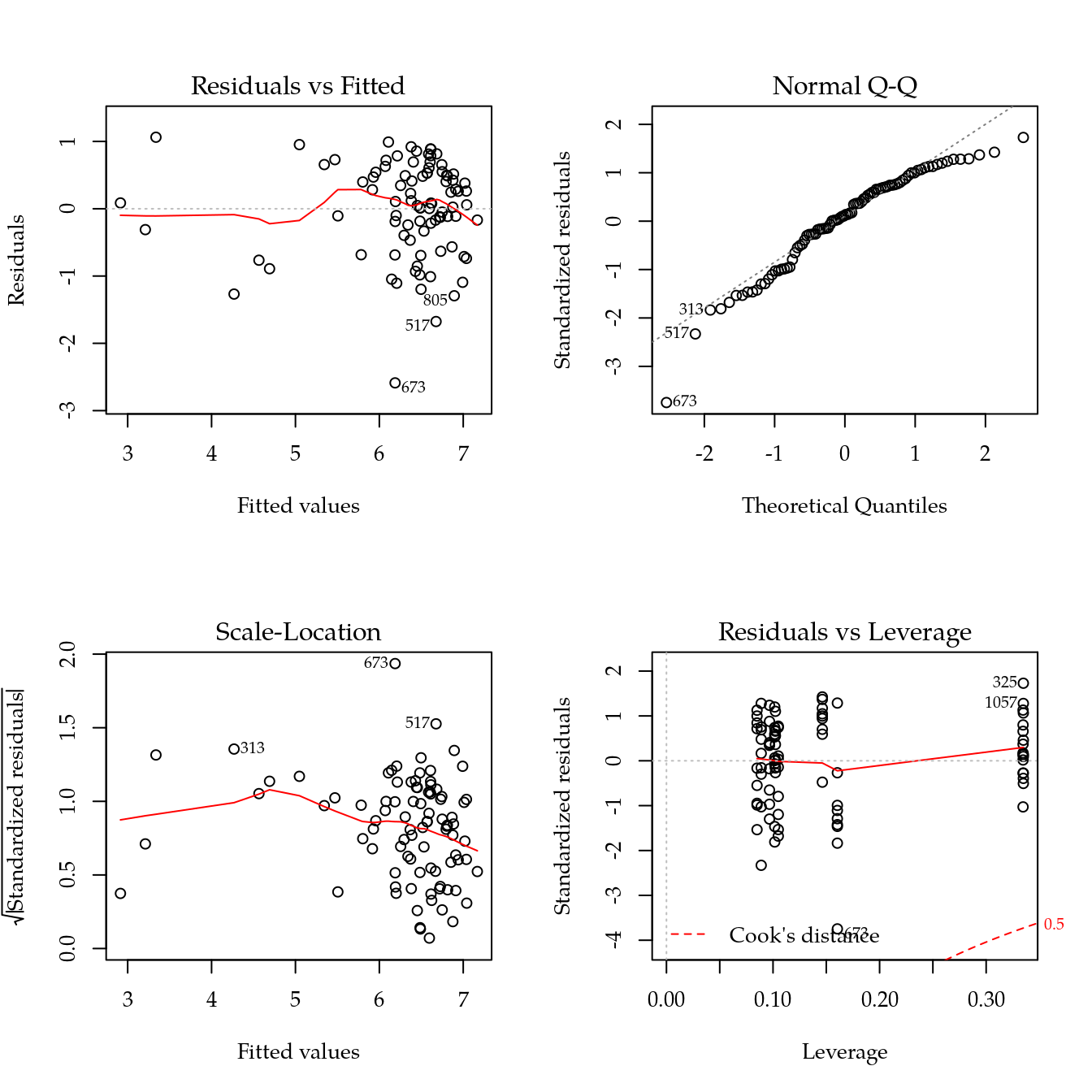
## Analysis of Variance Table
##
## Response: phkcl
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 2.833 1.4163 2.4938 0.08933 .
## biocar 2 18.545 9.2723 16.3262 1.263e-06 ***
## bs(dose, df = 3) 3 13.855 4.6182 8.1315 9.157e-05 ***
## biocar:bs(dose, df = 3) 6 20.501 3.4168 6.0160 3.476e-05 ***
## Residuals 76 43.163 0.5679
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Teste para a falta de ajuste.
anova(m1, m0)## Analysis of Variance Table
##
## Model 1: phkcl ~ bloc + biocar + bs(dose, df = 3) + biocar:bs(dose, df = 3)
## Model 2: phkcl ~ bloc + biocar * P
## Res.Df RSS Df Sum of Sq F Pr(>F)
## 1 76 43.163
## 2 58 28.547 18 14.616 1.6498 0.07734 .
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Novos valores em grid para fazer o gráfico de bandas.
pred <- with(bio,
expand.grid(bloc = factor(levels(bloc)[1],
levels = levels(bloc)),
biocar = levels(biocar),
dose = seq(min(dose),
max(dose),
length.out = 60)))
# Extrai a especificação do tempo base splines.
form <- attr(terms(formula(m1)), "term.labels")[3]
B <- eval(parse(text = form), envir = bio)
str(B)## 'bs' num [1:90, 1:3] 0 0 0 0.373 0.373 ...
## - attr(*, "dimnames")=List of 2
## ..$ : NULL
## ..$ : chr [1:3] "1" "2" "3"
## - attr(*, "degree")= int 3
## - attr(*, "knots")= num(0)
## - attr(*, "Boundary.knots")= num [1:2] 0 5.96
## - attr(*, "intercept")= logi FALSE# Gera a matriz para os valores de predição.
X <- predict(B, pred$dos)
# Cria a matriz do modelo para fazer a predição.
L <- model.matrix(~bloc + biocar * X, data = pred)
# Dá pesos para os efeitos de blocos correspondente à LSmeans.
i <- attr(L, "assign") == 1
L[, i] <- 1/(sum(i) + 1)
# Obtém os valores preditos com IC.
ci <- confint(glht(m1, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
xyplot(phkcl ~ dose,
groups = biocar,
data = bio,
auto.key = list(points = FALSE,
lines = TRUE,
divide = 1,
type = "o")) +
as.layer(xyplot(Estimate ~ dose,
groups = biocar,
data = pred,
type = "l",
ly = pred$lwr,
uy = pred$upr,
cty = "bands",
alpha = 0.3,
prepanel = prepanel.cbH,
panel.groups = panel.cbH,
panel = panel.superpose))
Ponto de carga zero (pcz)
#-----------------------------------------------------------------------
# Modelo saturado (considera dose como qualitativo).
m0 <- lm(pcz ~ bloc + biocar * P, data = bio)
par(mfrow = c(2, 2))
plot(m0)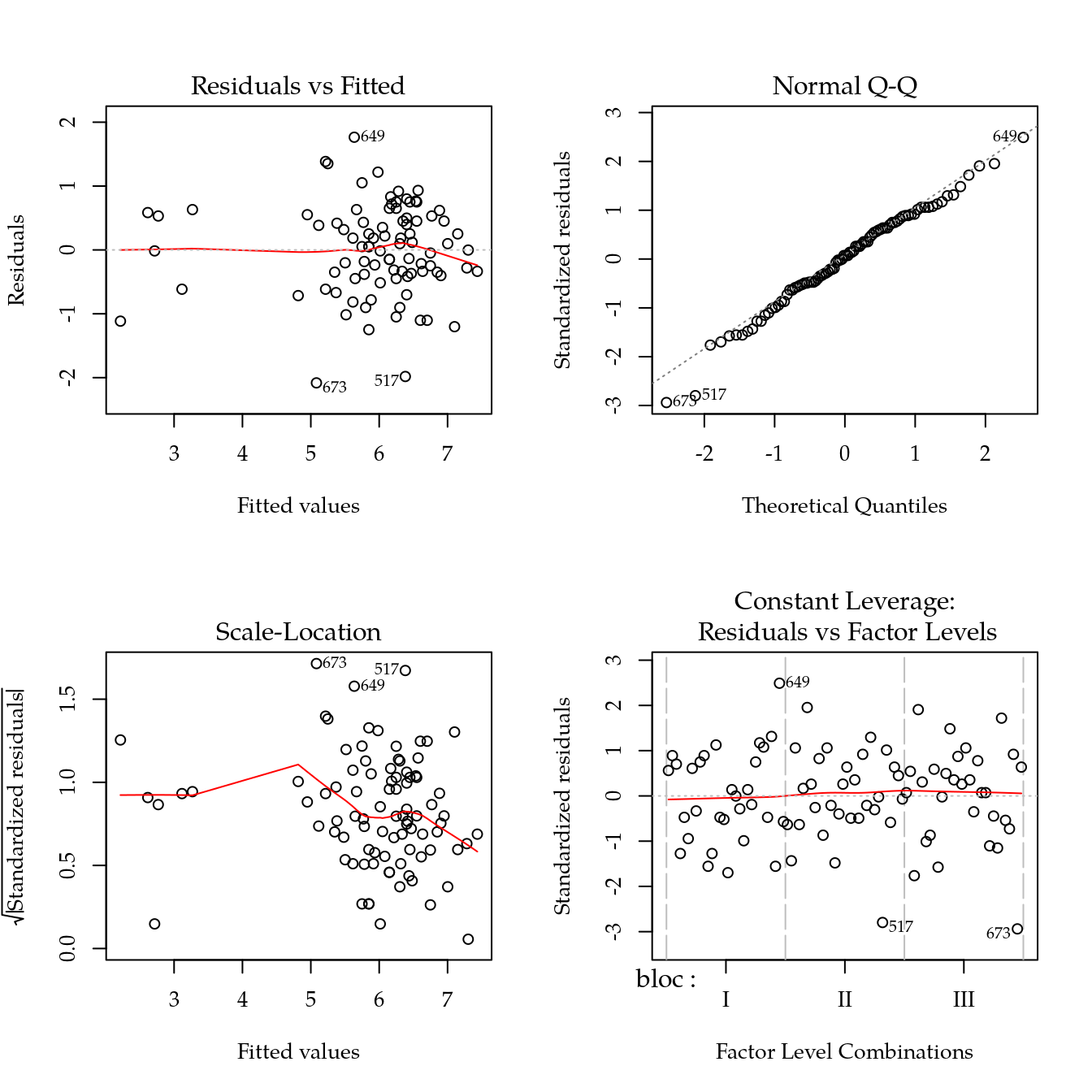
## Analysis of Variance Table
##
## Response: pcz
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 4.897 2.4484 3.1434 0.050566 .
## biocar 2 26.772 13.3861 17.1858 1.377e-06 ***
## P 9 31.616 3.5129 4.5101 0.000162 ***
## biocar:P 18 27.759 1.5422 1.9799 0.026116 *
## Residuals 58 45.176 0.7789
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1L <- LE_matrix(m0, effect = c("biocar", "P"))
pred <- equallevels(attr(L, "grid"), bio)
ci <- confint(glht(m0, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
str(pred)## 'data.frame': 30 obs. of 5 variables:
## $ biocar : Factor w/ 3 levels "S_BP","SP","SP_B": 1 2 3 1 2 3 1 2 3 1 ...
## $ P : Factor w/ 10 levels "0","1.5","3",..: 1 1 1 2 2 2 3 3 3 4 ...
## $ Estimate: num 6.17 6.37 6.07 5.43 6.07 ...
## $ lwr : num 5.15 5.35 5.05 4.41 5.05 ...
## $ upr : num 7.19 7.39 7.09 6.45 7.09 ...pch <- c(1, 4, 5)
key <- list(title = "Biocarvão",
cex.title = 1.1,
text = list(levels(bio$biocar)),
lines = list(pch = pch, lty = 1),
columns = 3,
type = "o",
divide = 1)
segplot(P ~ lwr + upr,
centers = Estimate,
data = pred,
draw = FALSE,
groups = biocar,
xlab = "Dose de P",
ylab = "Altura final (cm)",
key = key,
horizontal = FALSE,
pch = pch,
gap = 0.1,
panel = panel.groups.segplot)
#-----------------------------------------------------------------------
# Modelo reduzido baseado em splines.
m1 <- update(m0, ~ bloc + biocar * bs(dose, df = 3))
par(mfrow = c(2, 2))
plot(m1)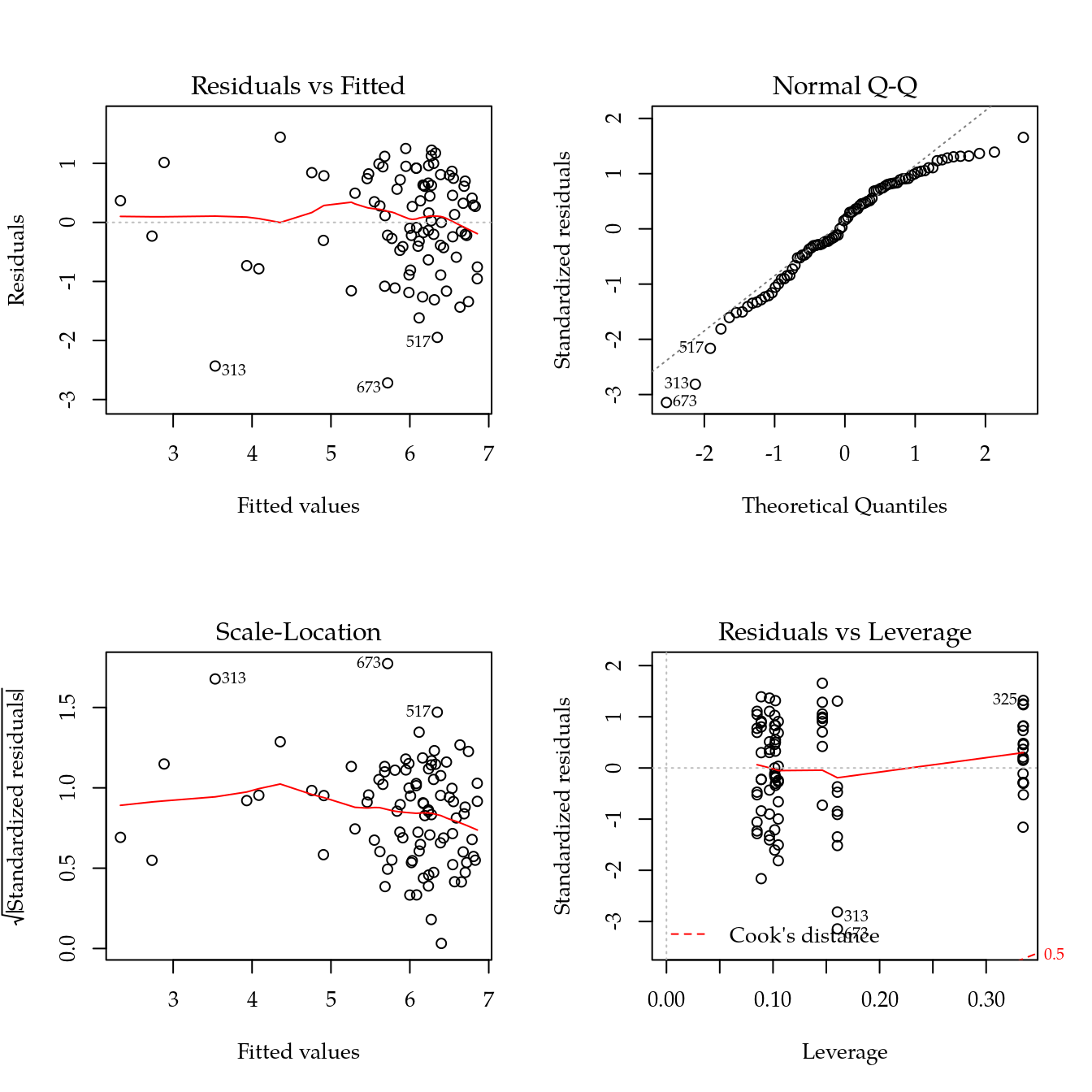
## Analysis of Variance Table
##
## Response: pcz
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 4.897 2.4484 2.7513 0.070212 .
## biocar 2 26.772 13.3861 15.0417 3.136e-06 ***
## bs(dose, df = 3) 3 15.630 5.2101 5.8545 0.001187 **
## biocar:bs(dose, df = 3) 6 21.286 3.5477 3.9865 0.001603 **
## Residuals 76 67.635 0.8899
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Teste para a falta de ajuste.
anova(m1, m0)## Analysis of Variance Table
##
## Model 1: pcz ~ bloc + biocar + bs(dose, df = 3) + biocar:bs(dose, df = 3)
## Model 2: pcz ~ bloc + biocar * P
## Res.Df RSS Df Sum of Sq F Pr(>F)
## 1 76 67.635
## 2 58 45.176 18 22.458 1.6019 0.09003 .
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Novos valores em grid para fazer o gráfico de bandas.
pred <- with(bio,
expand.grid(bloc = factor(levels(bloc)[1],
levels = levels(bloc)),
biocar = levels(biocar),
dose = seq(min(dose),
max(dose),
length.out = 60)))
# Extrai a especificação do tempo base splines.
form <- attr(terms(formula(m1)), "term.labels")[3]
B <- eval(parse(text = form), envir = bio)
str(B)## 'bs' num [1:90, 1:3] 0 0 0 0.373 0.373 ...
## - attr(*, "dimnames")=List of 2
## ..$ : NULL
## ..$ : chr [1:3] "1" "2" "3"
## - attr(*, "degree")= int 3
## - attr(*, "knots")= num(0)
## - attr(*, "Boundary.knots")= num [1:2] 0 5.96
## - attr(*, "intercept")= logi FALSE# Gera a matriz para os valores de predição.
X <- predict(B, pred$dos)
# Cria a matriz do modelo para fazer a predição.
L <- model.matrix(~bloc + biocar * X, data = pred)
# Dá pesos para os efeitos de blocos correspondente à LSmeans.
i <- attr(L, "assign") == 1
L[, i] <- 1/(sum(i) + 1)
# Obtém os valores preditos com IC.
ci <- confint(glht(m1, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
xyplot(pcz ~ dose,
groups = biocar,
data = bio,
auto.key = list(points = FALSE,
lines = TRUE,
divide = 1,
type = "o")) +
as.layer(xyplot(Estimate ~ dose,
groups = biocar,
data = pred,
type = "l",
ly = pred$lwr,
uy = pred$upr,
cty = "bands",
alpha = 0.3,
prepanel = prepanel.cbH,
panel.groups = panel.cbH,
panel = panel.superpose))
Fósforo (p)
#-----------------------------------------------------------------------
# Modelo saturado (considera dose como qualitativo).
m0 <- lm(log(p) ~ bloc + biocar * P, data = bio)
par(mfrow = c(2, 2))
plot(m0)
## Analysis of Variance Table
##
## Response: log(p)
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 0.0174 0.0087 0.1333 0.8755
## biocar 2 12.1580 6.0790 93.3584 < 2.2e-16 ***
## P 9 25.6059 2.8451 43.6939 < 2.2e-16 ***
## biocar:P 18 4.7591 0.2644 4.0604 2.61e-05 ***
## Residuals 57 3.7115 0.0651
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1L <- LE_matrix(m0, effect = c("biocar", "P"))
pred <- equallevels(attr(L, "grid"), bio)
ci <- confint(glht(m0, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
str(pred)## 'data.frame': 30 obs. of 5 variables:
## $ biocar : Factor w/ 3 levels "S_BP","SP","SP_B": 1 2 3 1 2 3 1 2 3 1 ...
## $ P : Factor w/ 10 levels "0","1.5","3",..: 1 1 1 2 2 2 3 3 3 4 ...
## $ Estimate: num 3.63 3.29 3.8 4.14 3.64 ...
## $ lwr : num 3.33 2.99 3.51 3.85 3.34 ...
## $ upr : num 3.92 3.58 4.1 4.44 3.93 ...pch <- c(1, 4, 5)
key <- list(title = "Biocarvão",
cex.title = 1.1,
text = list(levels(bio$biocar)),
lines = list(pch = pch, lty = 1),
columns = 3,
type = "o",
divide = 1)
segplot(P ~ lwr + upr,
centers = Estimate,
data = pred,
draw = FALSE,
groups = biocar,
xlab = "Dose de P",
ylab = "Altura final (cm)",
key = key,
horizontal = FALSE,
pch = pch,
gap = 0.1,
panel = panel.groups.segplot)
#-----------------------------------------------------------------------
# Modelo reduzido baseado em splines.
m1 <- update(m0, ~ bloc + biocar * bs(dose, df = 3))
par(mfrow = c(2, 2))
plot(m1)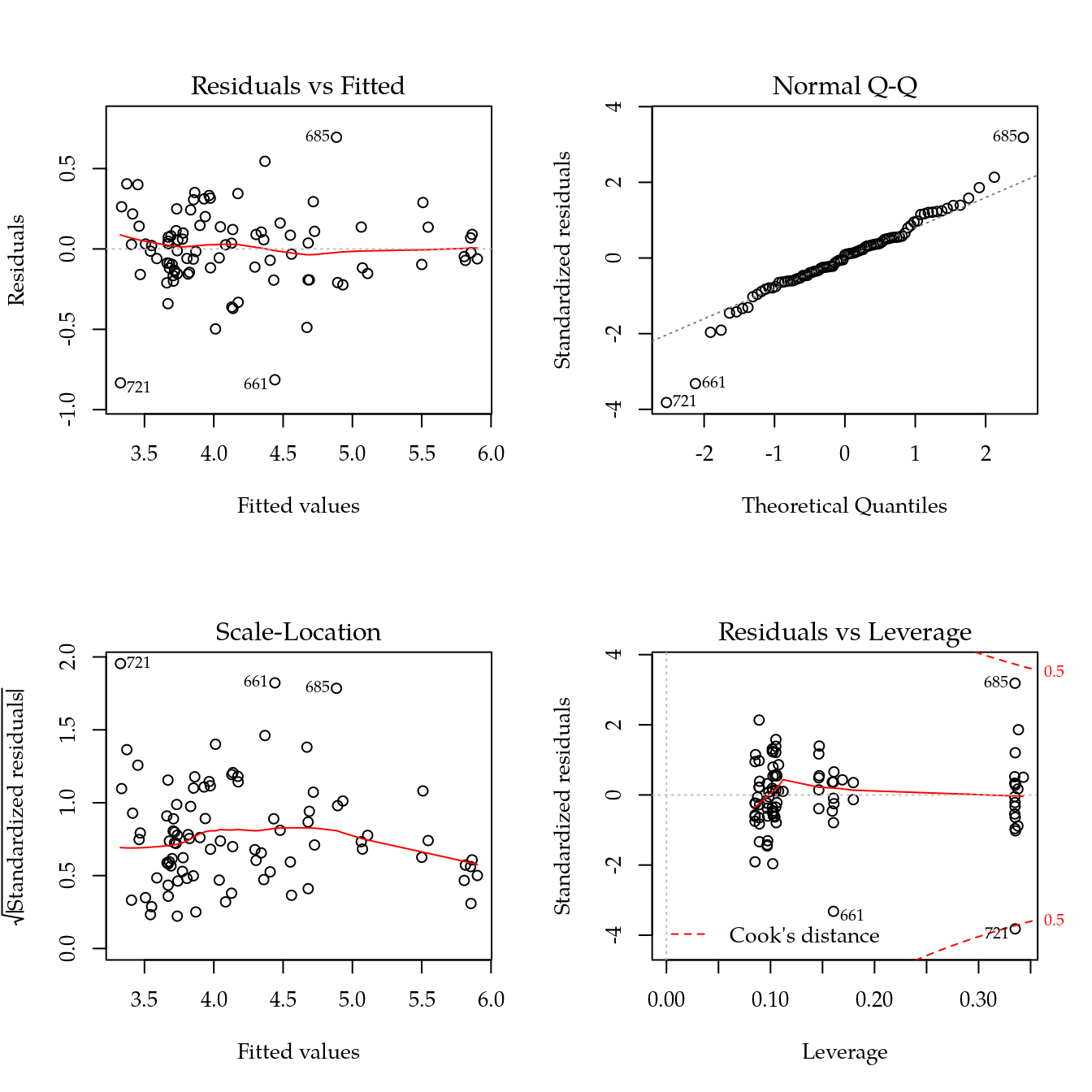
## Analysis of Variance Table
##
## Response: log(p)
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 0.0174 0.0087 0.1212 0.886
## biocar 2 12.1580 6.0790 84.9123 < 2.2e-16 ***
## bs(dose, df = 3) 3 25.0476 8.3492 116.6233 < 2.2e-16 ***
## biocar:bs(dose, df = 3) 6 3.6595 0.6099 8.5195 4.629e-07 ***
## Residuals 75 5.3693 0.0716
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Teste para a falta de ajuste.
anova(m1, m0)## Analysis of Variance Table
##
## Model 1: log(p) ~ bloc + biocar + bs(dose, df = 3) + biocar:bs(dose, df = 3)
## Model 2: log(p) ~ bloc + biocar * P
## Res.Df RSS Df Sum of Sq F Pr(>F)
## 1 75 5.3693
## 2 57 3.7115 18 1.6578 1.4145 0.1605# Novos valores em grid para fazer o gráfico de bandas.
pred <- with(bio,
expand.grid(bloc = factor(levels(bloc)[1],
levels = levels(bloc)),
biocar = levels(biocar),
dose = seq(min(dose),
max(dose),
length.out = 60)))
# Extrai a especificação do tempo base splines.
form <- attr(terms(formula(m1)), "term.labels")[3]
B <- eval(parse(text = form), envir = bio)
str(B)## 'bs' num [1:90, 1:3] 0 0 0 0.373 0.373 ...
## - attr(*, "dimnames")=List of 2
## ..$ : NULL
## ..$ : chr [1:3] "1" "2" "3"
## - attr(*, "degree")= int 3
## - attr(*, "knots")= num(0)
## - attr(*, "Boundary.knots")= num [1:2] 0 5.96
## - attr(*, "intercept")= logi FALSE# Gera a matriz para os valores de predição.
X <- predict(B, pred$dos)
# Cria a matriz do modelo para fazer a predição.
L <- model.matrix(~bloc + biocar * X, data = pred)
# Dá pesos para os efeitos de blocos correspondente à LSmeans.
i <- attr(L, "assign") == 1
L[, i] <- 1/(sum(i) + 1)
# Obtém os valores preditos com IC.
ci <- confint(glht(m1, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
xyplot(log(p) ~ dose,
groups = biocar,
data = bio,
auto.key = list(points = FALSE,
lines = TRUE,
divide = 1,
type = "o")) +
as.layer(xyplot(Estimate ~ dose,
groups = biocar,
data = pred,
type = "l",
ly = pred$lwr,
uy = pred$upr,
cty = "bands",
alpha = 0.3,
prepanel = prepanel.cbH,
panel.groups = panel.cbH,
panel = panel.superpose))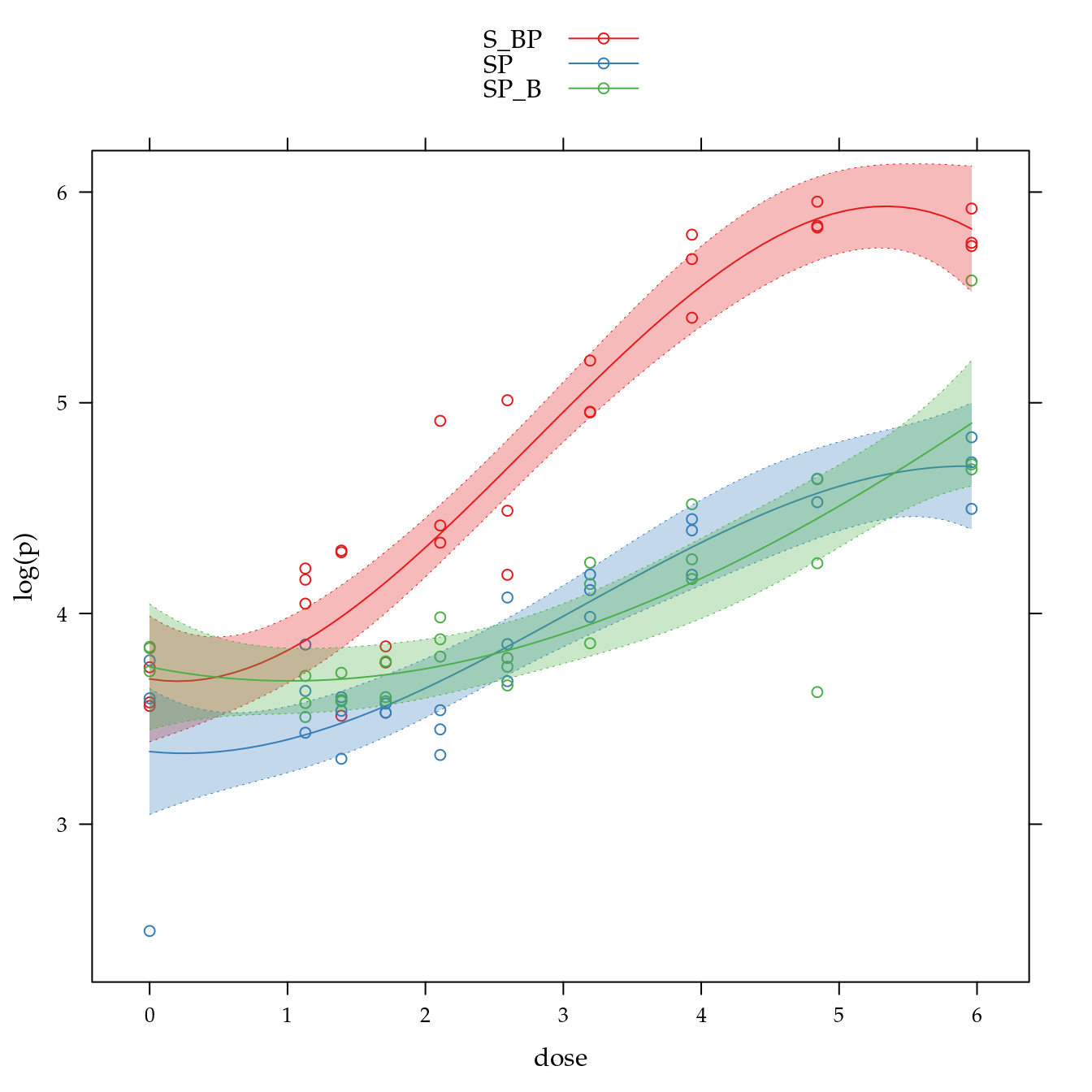
Potássio (k)
#-----------------------------------------------------------------------
# Modelo saturado (considera dose como qualitativo).
m0 <- lm(k ~ bloc + biocar * P, data = bio)
par(mfrow = c(2, 2))
plot(m0)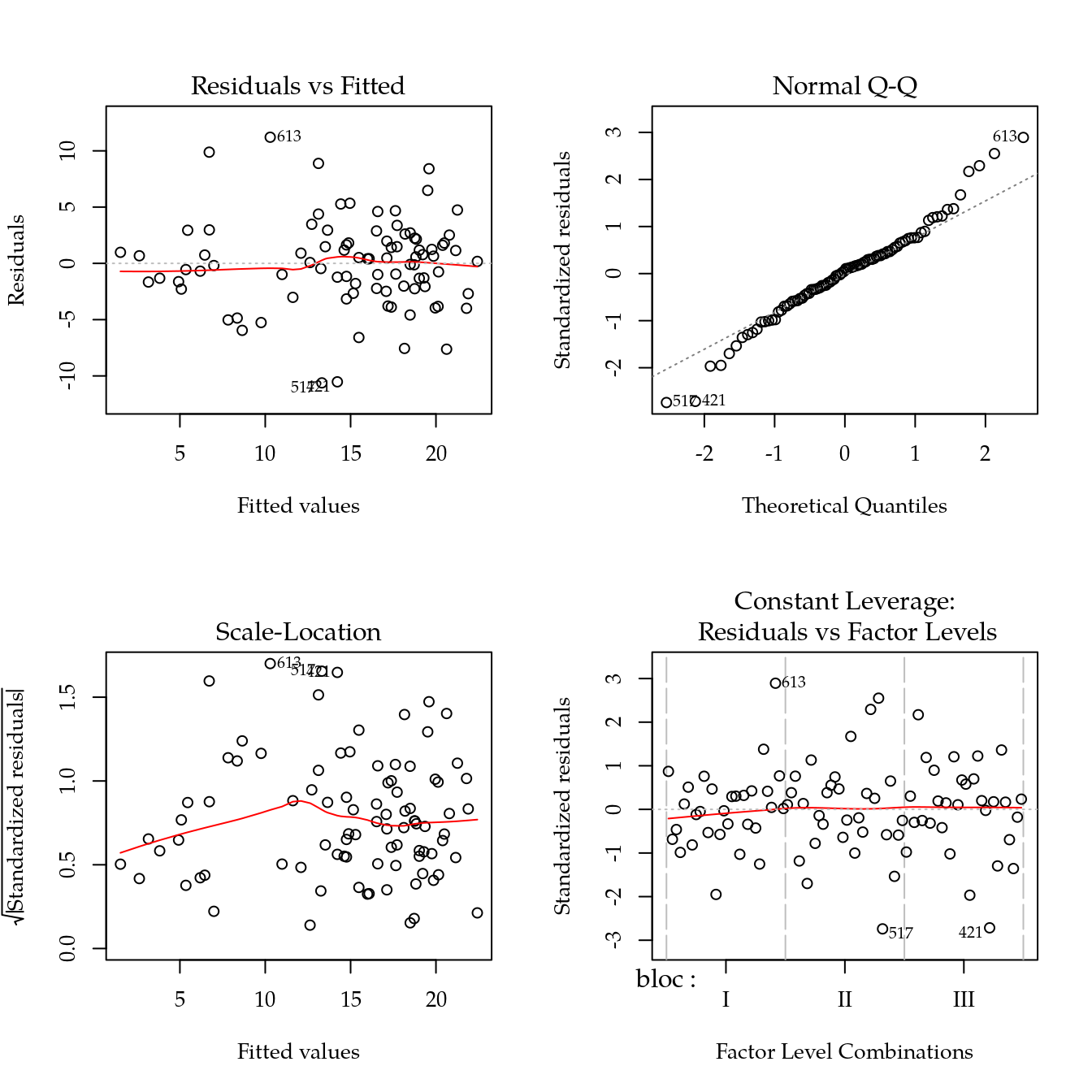
## Analysis of Variance Table
##
## Response: k
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 41.99 20.99 0.9007 0.411880
## biocar 2 652.48 326.24 13.9972 1.096e-05 ***
## P 9 681.39 75.71 3.2483 0.002936 **
## biocar:P 18 1022.55 56.81 2.4373 0.005480 **
## Residuals 58 1351.84 23.31
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1L <- LE_matrix(m0, effect = c("biocar", "P"))
pred <- equallevels(attr(L, "grid"), bio)
ci <- confint(glht(m0, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
str(pred)## 'data.frame': 30 obs. of 5 variables:
## $ biocar : Factor w/ 3 levels "S_BP","SP","SP_B": 1 2 3 1 2 3 1 2 3 1 ...
## $ P : Factor w/ 10 levels "0","1.5","3",..: 1 1 1 2 2 2 3 3 3 4 ...
## $ Estimate: num 17 18.6 18.3 14.4 19.8 ...
## $ lwr : num 11.42 13.05 12.72 8.85 14.19 ...
## $ upr : num 22.6 24.2 23.9 20 25.3 ...pch <- c(1, 4, 5)
key <- list(title = "Biocarvão",
cex.title = 1.1,
text = list(levels(bio$biocar)),
lines = list(pch = pch, lty = 1),
columns = 3,
type = "o",
divide = 1)
segplot(P ~ lwr + upr,
centers = Estimate,
data = pred,
draw = FALSE,
groups = biocar,
xlab = "Dose de P",
ylab = "Altura final (cm)",
key = key,
horizontal = FALSE,
pch = pch,
gap = 0.1,
panel = panel.groups.segplot)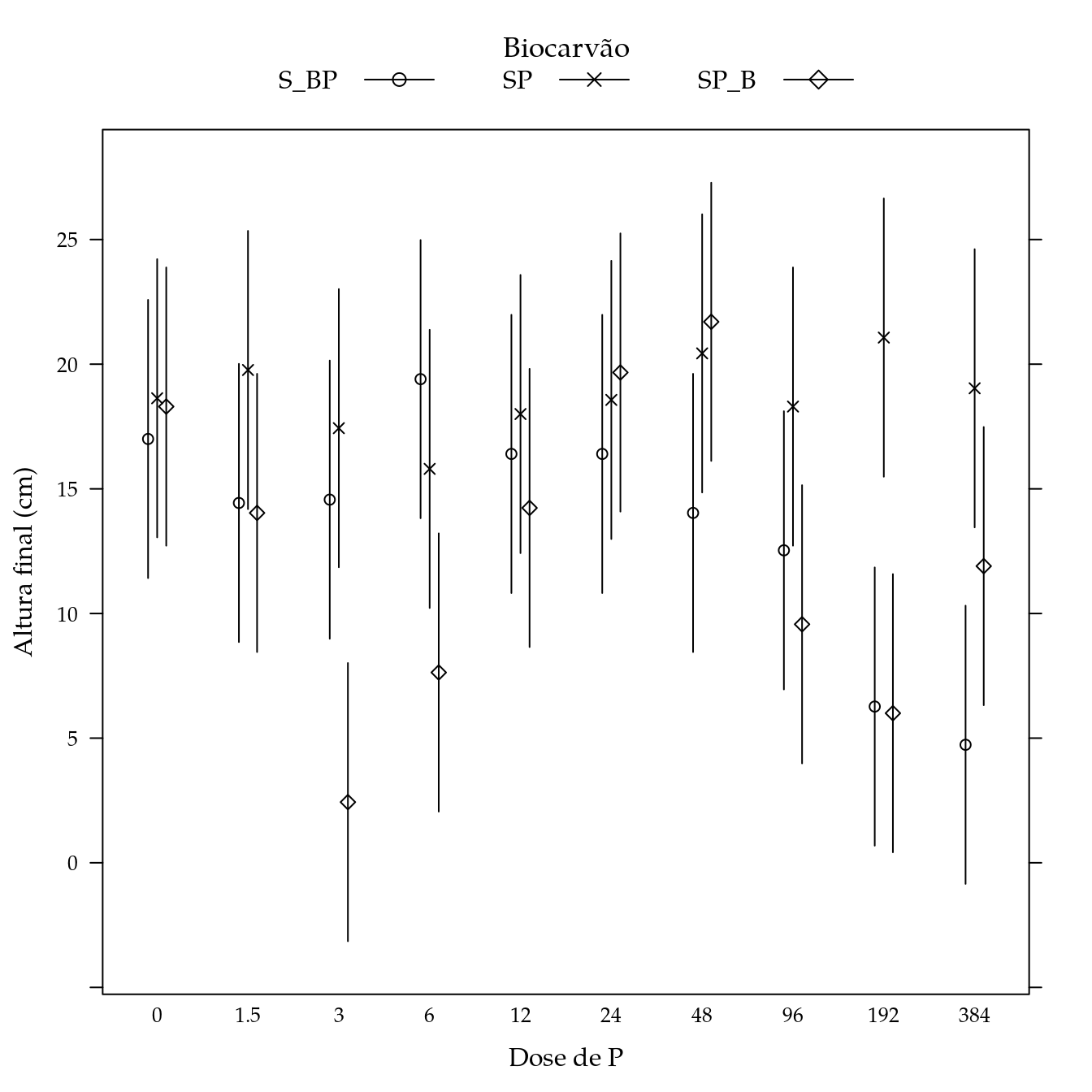
#-----------------------------------------------------------------------
# Modelo reduzido baseado em splines.
m1 <- update(m0, ~ bloc + biocar * bs(dose, df = 3))
par(mfrow = c(2, 2))
plot(m1)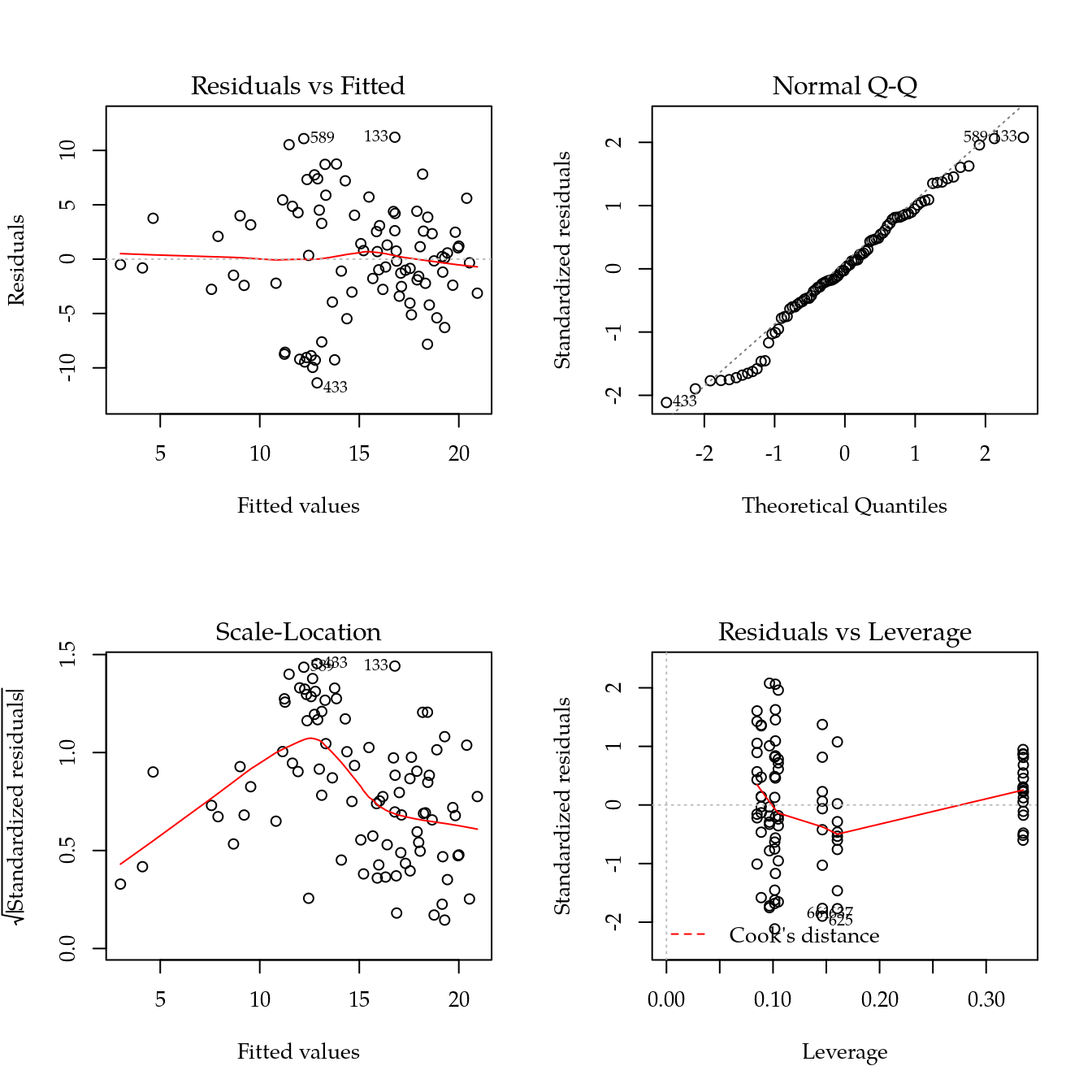
## Analysis of Variance Table
##
## Response: k
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 41.99 20.99 0.6511 0.5243733
## biocar 2 652.48 326.24 10.1175 0.0001271 ***
## bs(dose, df = 3) 3 226.44 75.48 2.3408 0.0799454 .
## biocar:bs(dose, df = 3) 6 378.70 63.12 1.9574 0.0823265 .
## Residuals 76 2450.64 32.25
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Teste para a falta de ajuste.
anova(m1, m0)## Analysis of Variance Table
##
## Model 1: k ~ bloc + biocar + bs(dose, df = 3) + biocar:bs(dose, df = 3)
## Model 2: k ~ bloc + biocar * P
## Res.Df RSS Df Sum of Sq F Pr(>F)
## 1 76 2450.6
## 2 58 1351.8 18 1098.8 2.6191 0.002933 **
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Novos valores em grid para fazer o gráfico de bandas.
pred <- with(bio,
expand.grid(bloc = factor(levels(bloc)[1],
levels = levels(bloc)),
biocar = levels(biocar),
dose = seq(min(dose),
max(dose),
length.out = 60)))
# Extrai a especificação do tempo base splines.
form <- attr(terms(formula(m1)), "term.labels")[3]
B <- eval(parse(text = form), envir = bio)
str(B)## 'bs' num [1:90, 1:3] 0 0 0 0.373 0.373 ...
## - attr(*, "dimnames")=List of 2
## ..$ : NULL
## ..$ : chr [1:3] "1" "2" "3"
## - attr(*, "degree")= int 3
## - attr(*, "knots")= num(0)
## - attr(*, "Boundary.knots")= num [1:2] 0 5.96
## - attr(*, "intercept")= logi FALSE# Gera a matriz para os valores de predição.
X <- predict(B, pred$dos)
# Cria a matriz do modelo para fazer a predição.
L <- model.matrix(~bloc + biocar * X, data = pred)
# Dá pesos para os efeitos de blocos correspondente à LSmeans.
i <- attr(L, "assign") == 1
L[, i] <- 1/(sum(i) + 1)
# Obtém os valores preditos com IC.
ci <- confint(glht(m1, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
xyplot(k ~ dose,
groups = biocar,
data = bio,
auto.key = list(points = FALSE,
lines = TRUE,
divide = 1,
type = "o")) +
as.layer(xyplot(Estimate ~ dose,
groups = biocar,
data = pred,
type = "l",
ly = pred$lwr,
uy = pred$upr,
cty = "bands",
alpha = 0.3,
prepanel = prepanel.cbH,
panel.groups = panel.cbH,
panel = panel.superpose))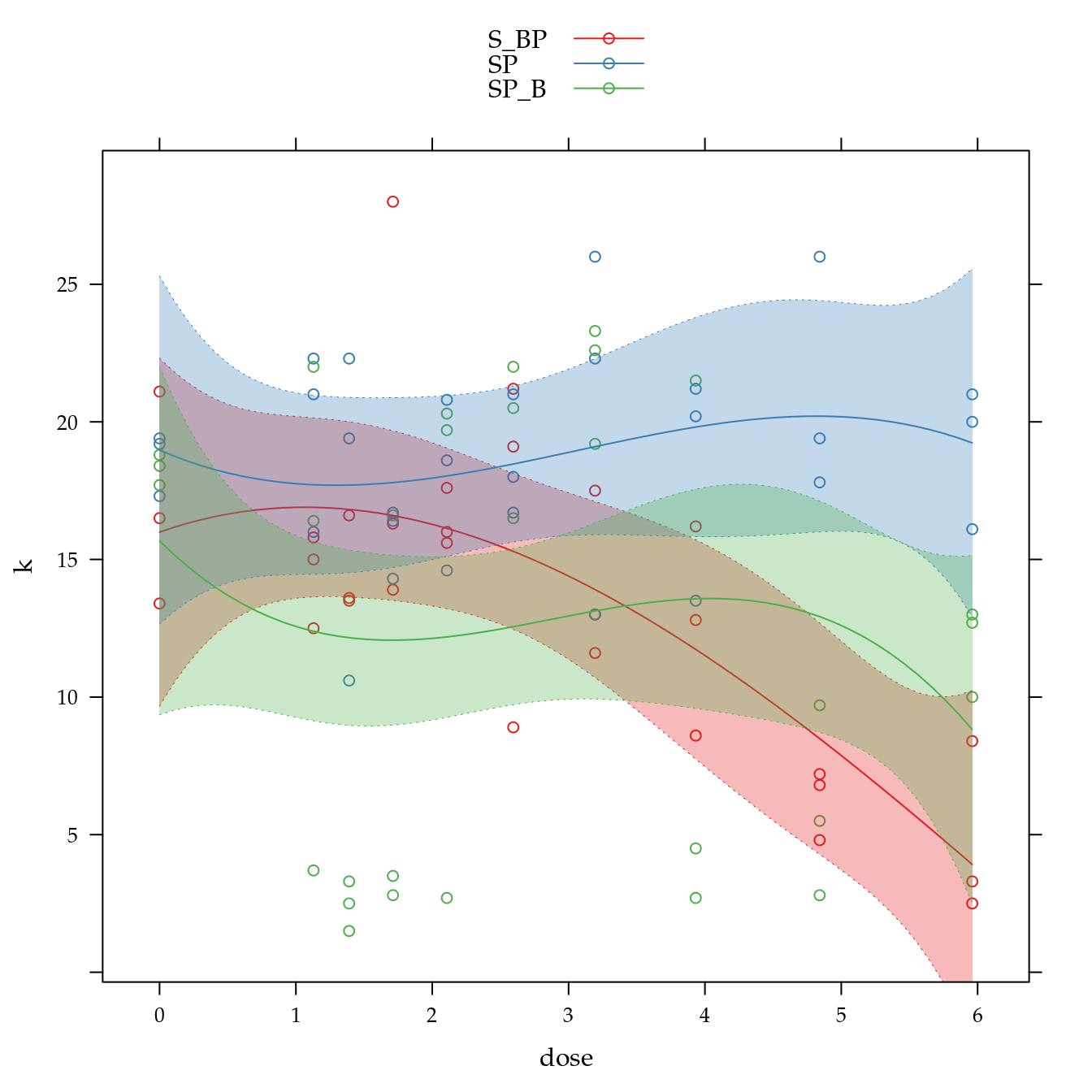
Cálcio (Ca)
#-----------------------------------------------------------------------
# Modelo saturado (considera dose como qualitativo).
m0 <- lm(ca ~ bloc + biocar * P, data = bio)
par(mfrow = c(2, 2))
plot(m0)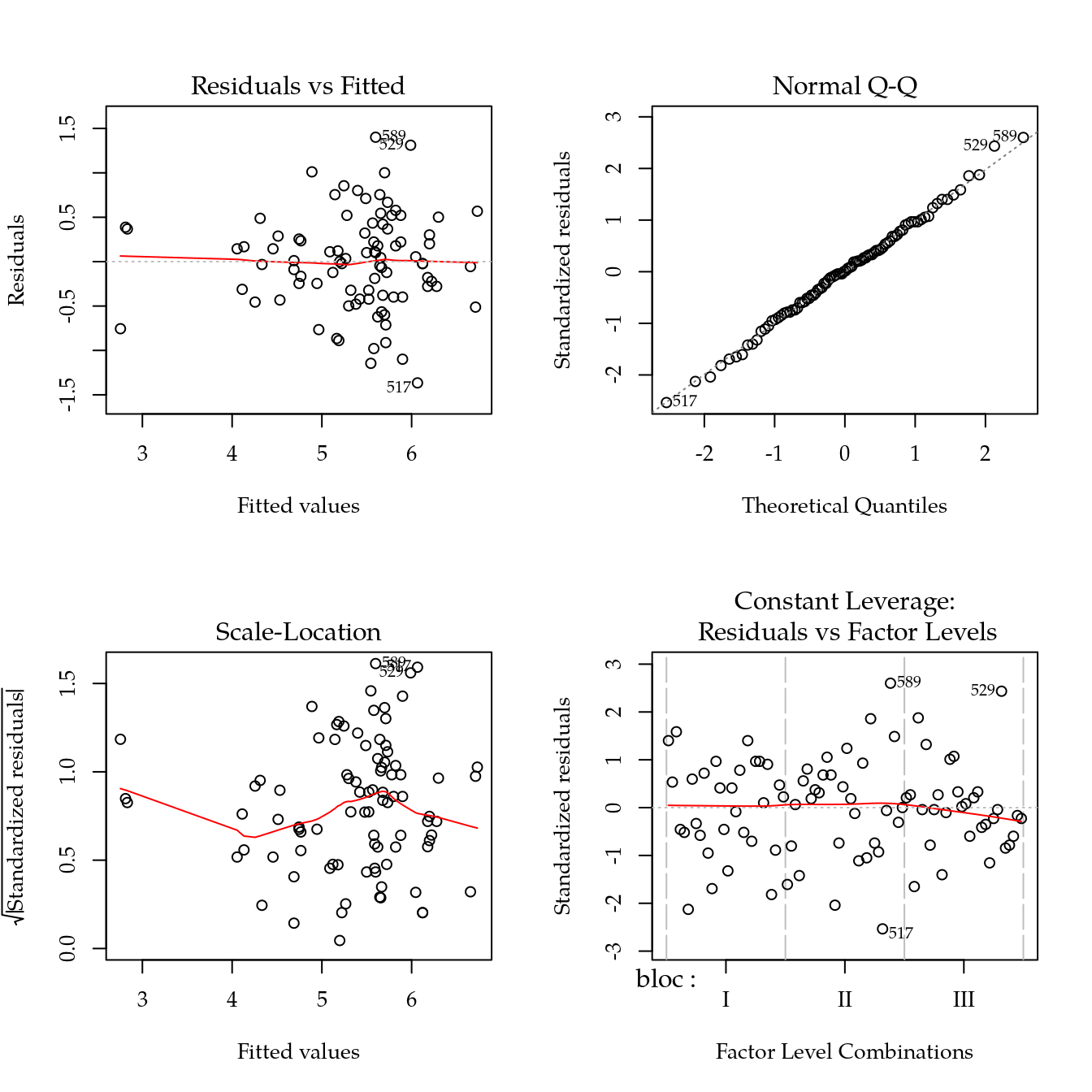
## Analysis of Variance Table
##
## Response: ca
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 0.0949 0.0474 0.1054 0.9001744
## biocar 2 8.8436 4.4218 9.8192 0.0002124 ***
## P 9 10.9534 1.2170 2.7026 0.0106650 *
## biocar:P 18 30.6809 1.7045 3.7851 5.877e-05 ***
## Residuals 58 26.1184 0.4503
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1L <- LE_matrix(m0, effect = c("biocar", "P"))
pred <- equallevels(attr(L, "grid"), bio)
ci <- confint(glht(m0, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
str(pred)## 'data.frame': 30 obs. of 5 variables:
## $ biocar : Factor w/ 3 levels "S_BP","SP","SP_B": 1 2 3 1 2 3 1 2 3 1 ...
## $ P : Factor w/ 10 levels "0","1.5","3",..: 1 1 1 2 2 2 3 3 3 4 ...
## $ Estimate: num 5.13 6.7 5.63 4.5 5.7 ...
## $ lwr : num 4.36 5.92 4.86 3.72 4.92 ...
## $ upr : num 5.91 7.48 6.41 5.28 6.48 ...pch <- c(1, 4, 5)
key <- list(title = "Biocarvão",
cex.title = 1.1,
text = list(levels(bio$biocar)),
lines = list(pch = pch, lty = 1),
columns = 3,
type = "o",
divide = 1)
segplot(P ~ lwr + upr,
centers = Estimate,
data = pred,
draw = FALSE,
groups = biocar,
xlab = "Dose de P",
ylab = "Altura final (cm)",
key = key,
horizontal = FALSE,
pch = pch,
gap = 0.1,
panel = panel.groups.segplot)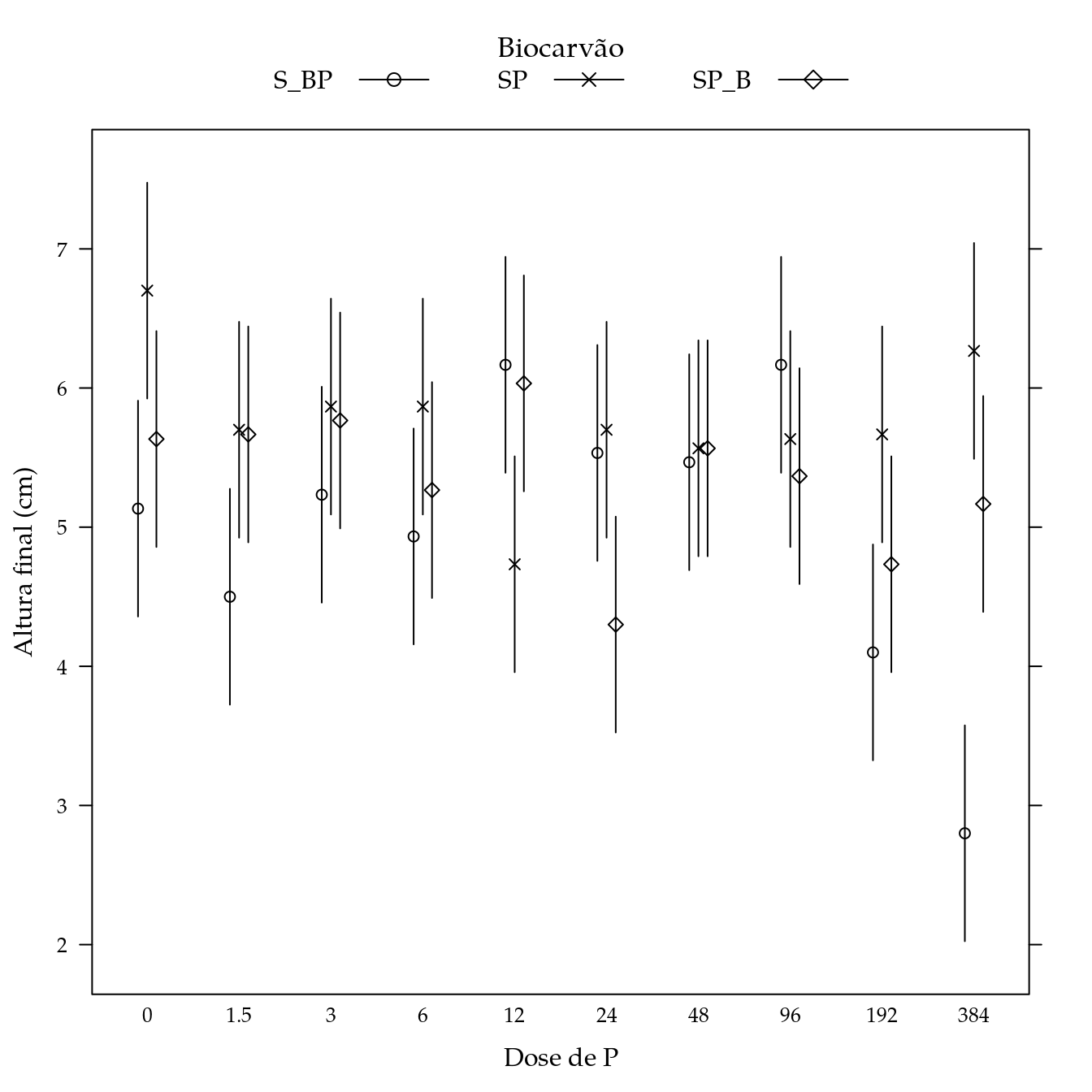
#-----------------------------------------------------------------------
# Modelo reduzido baseado em splines.
m1 <- update(m0, ~ bloc + biocar * bs(dose, df = 3))
par(mfrow = c(2, 2))
plot(m1)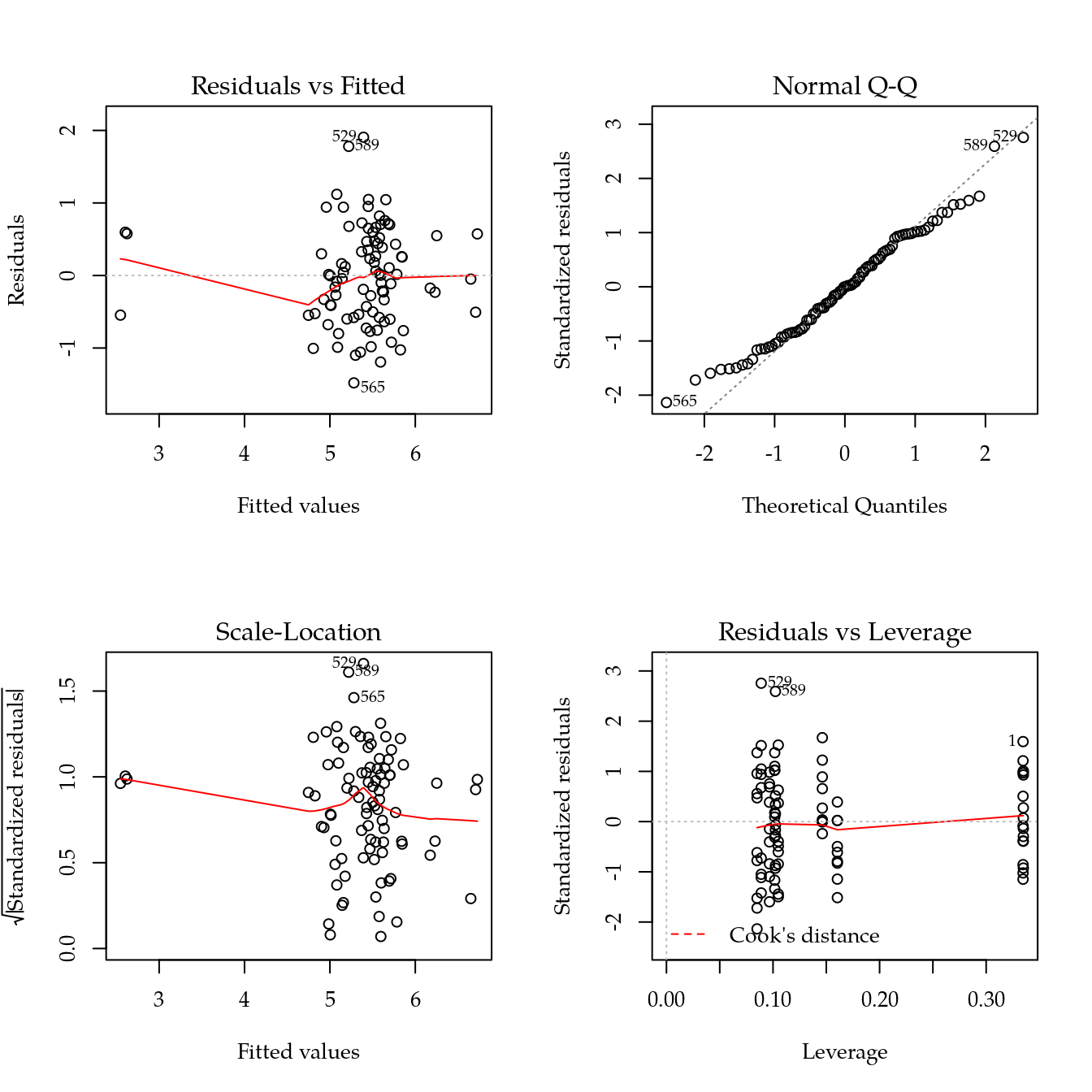
## Analysis of Variance Table
##
## Response: ca
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 0.095 0.0474 0.0903 0.9137185
## biocar 2 8.844 4.4218 8.4196 0.0004979 ***
## bs(dose, df = 3) 3 6.872 2.2907 4.3617 0.0068824 **
## biocar:bs(dose, df = 3) 6 20.967 3.4945 6.6540 1.099e-05 ***
## Residuals 76 39.913 0.5252
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Teste para a falta de ajuste.
anova(m1, m0)## Analysis of Variance Table
##
## Model 1: ca ~ bloc + biocar + bs(dose, df = 3) + biocar:bs(dose, df = 3)
## Model 2: ca ~ bloc + biocar * P
## Res.Df RSS Df Sum of Sq F Pr(>F)
## 1 76 39.913
## 2 58 26.118 18 13.795 1.7019 0.06543 .
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Novos valores em grid para fazer o gráfico de bandas.
pred <- with(bio,
expand.grid(bloc = factor(levels(bloc)[1],
levels = levels(bloc)),
biocar = levels(biocar),
dose = seq(min(dose),
max(dose),
length.out = 60)))
# Extrai a especificação do tempo base splines.
form <- attr(terms(formula(m1)), "term.labels")[3]
B <- eval(parse(text = form), envir = bio)
str(B)## 'bs' num [1:90, 1:3] 0 0 0 0.373 0.373 ...
## - attr(*, "dimnames")=List of 2
## ..$ : NULL
## ..$ : chr [1:3] "1" "2" "3"
## - attr(*, "degree")= int 3
## - attr(*, "knots")= num(0)
## - attr(*, "Boundary.knots")= num [1:2] 0 5.96
## - attr(*, "intercept")= logi FALSE# Gera a matriz para os valores de predição.
X <- predict(B, pred$dos)
# Cria a matriz do modelo para fazer a predição.
L <- model.matrix(~bloc + biocar * X, data = pred)
# Dá pesos para os efeitos de blocos correspondente à LSmeans.
i <- attr(L, "assign") == 1
L[, i] <- 1/(sum(i) + 1)
# Obtém os valores preditos com IC.
ci <- confint(glht(m1, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
xyplot(ca ~ dose,
groups = biocar,
data = bio,
auto.key = list(points = FALSE,
lines = TRUE,
divide = 1,
type = "o")) +
as.layer(xyplot(Estimate ~ dose,
groups = biocar,
data = pred,
type = "l",
ly = pred$lwr,
uy = pred$upr,
cty = "bands",
alpha = 0.3,
prepanel = prepanel.cbH,
panel.groups = panel.cbH,
panel = panel.superpose))
Magnésio (Mg)
#-----------------------------------------------------------------------
# Modelo saturado (considera dose como qualitativo).
m0 <- lm(mg ~ bloc + biocar * P, data = bio)
par(mfrow = c(2, 2))
plot(m0)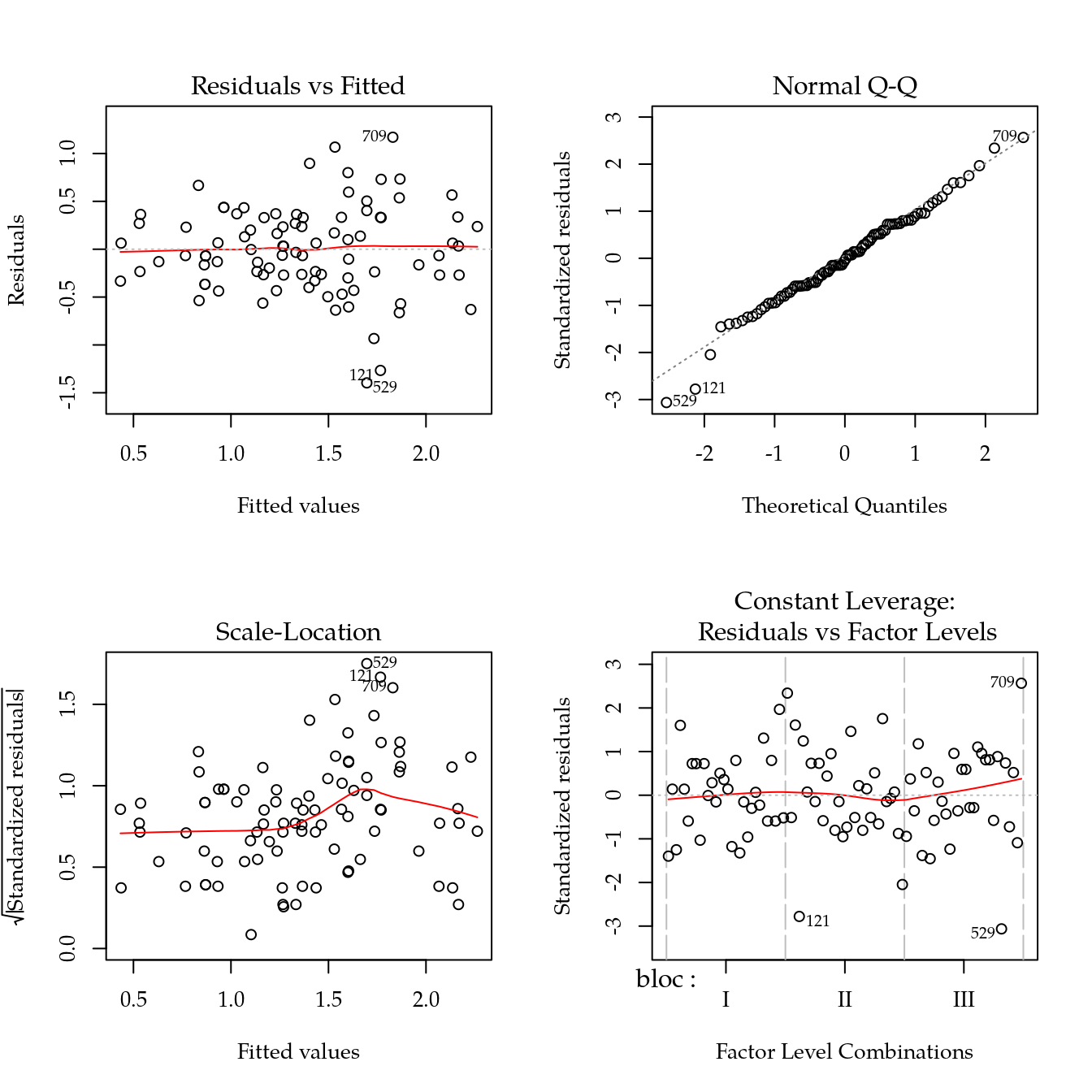
## Analysis of Variance Table
##
## Response: mg
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 0.1807 0.0903 0.2801 0.7567
## biocar 2 8.7007 4.3503 13.4887 1.547e-05 ***
## P 9 2.7773 0.3086 0.9568 0.4846
## biocar:P 18 6.0393 0.3355 1.0403 0.4324
## Residuals 58 18.7060 0.3225
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1L <- LE_matrix(m0, effect = c("biocar", "P"))
pred <- equallevels(attr(L, "grid"), bio)
ci <- confint(glht(m0, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
str(pred)## 'data.frame': 30 obs. of 5 variables:
## $ biocar : Factor w/ 3 levels "S_BP","SP","SP_B": 1 2 3 1 2 3 1 2 3 1 ...
## $ P : Factor w/ 10 levels "0","1.5","3",..: 1 1 1 2 2 2 3 3 3 4 ...
## $ Estimate: num 1.567 1.133 0.967 1.467 1.1 ...
## $ lwr : num 0.91 0.477 0.31 0.81 0.444 ...
## $ upr : num 2.22 1.79 1.62 2.12 1.76 ...pch <- c(1, 4, 5)
key <- list(title = "Biocarvão",
cex.title = 1.1,
text = list(levels(bio$biocar)),
lines = list(pch = pch, lty = 1),
columns = 3,
type = "o",
divide = 1)
segplot(P ~ lwr + upr,
centers = Estimate,
data = pred,
draw = FALSE,
groups = biocar,
xlab = "Dose de P",
ylab = "Altura final (cm)",
key = key,
horizontal = FALSE,
pch = pch,
gap = 0.1,
panel = panel.groups.segplot)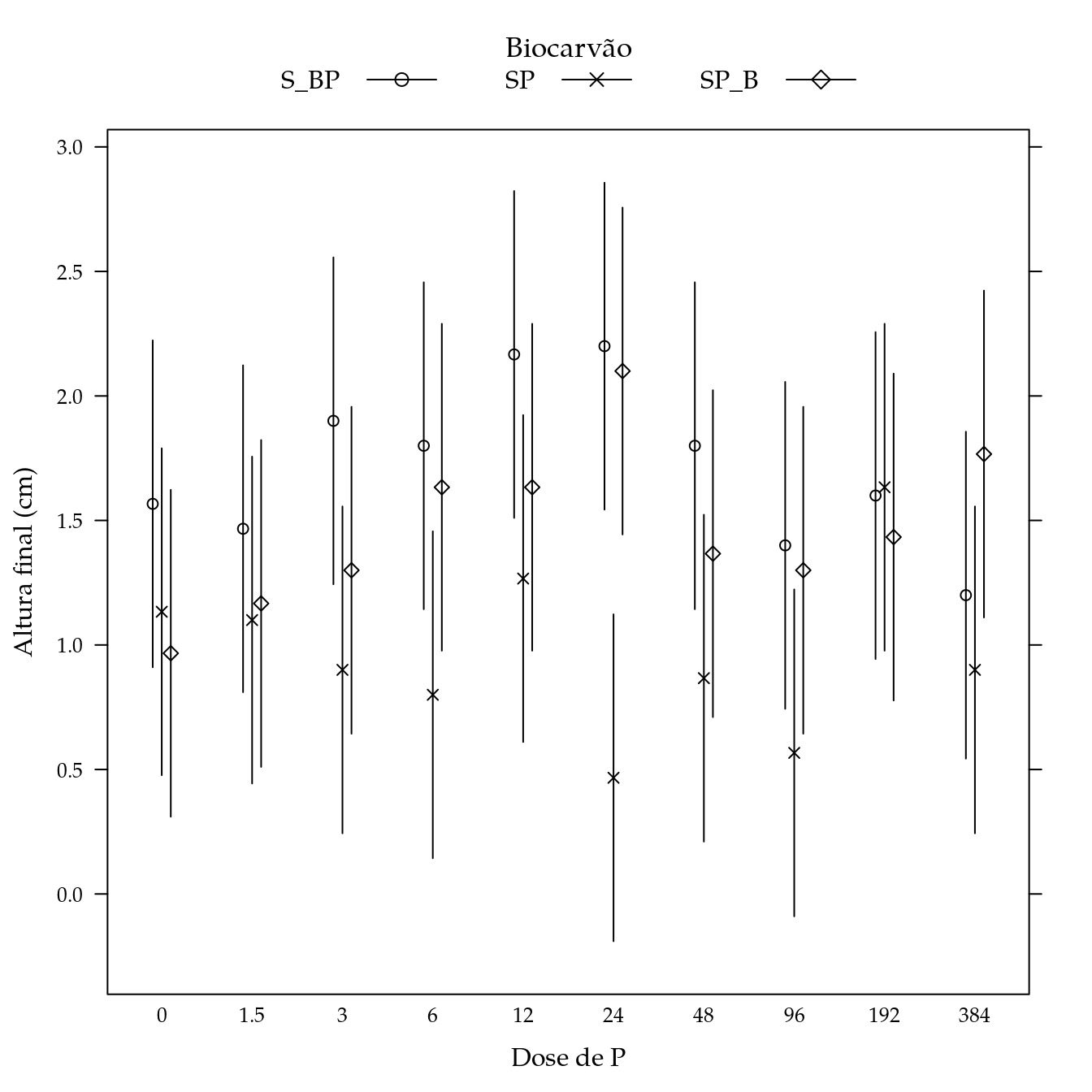
#-----------------------------------------------------------------------
# Modelo reduzido baseado em splines.
m1 <- update(m0, ~ bloc + biocar * bs(dose, df = 3))
par(mfrow = c(2, 2))
plot(m1)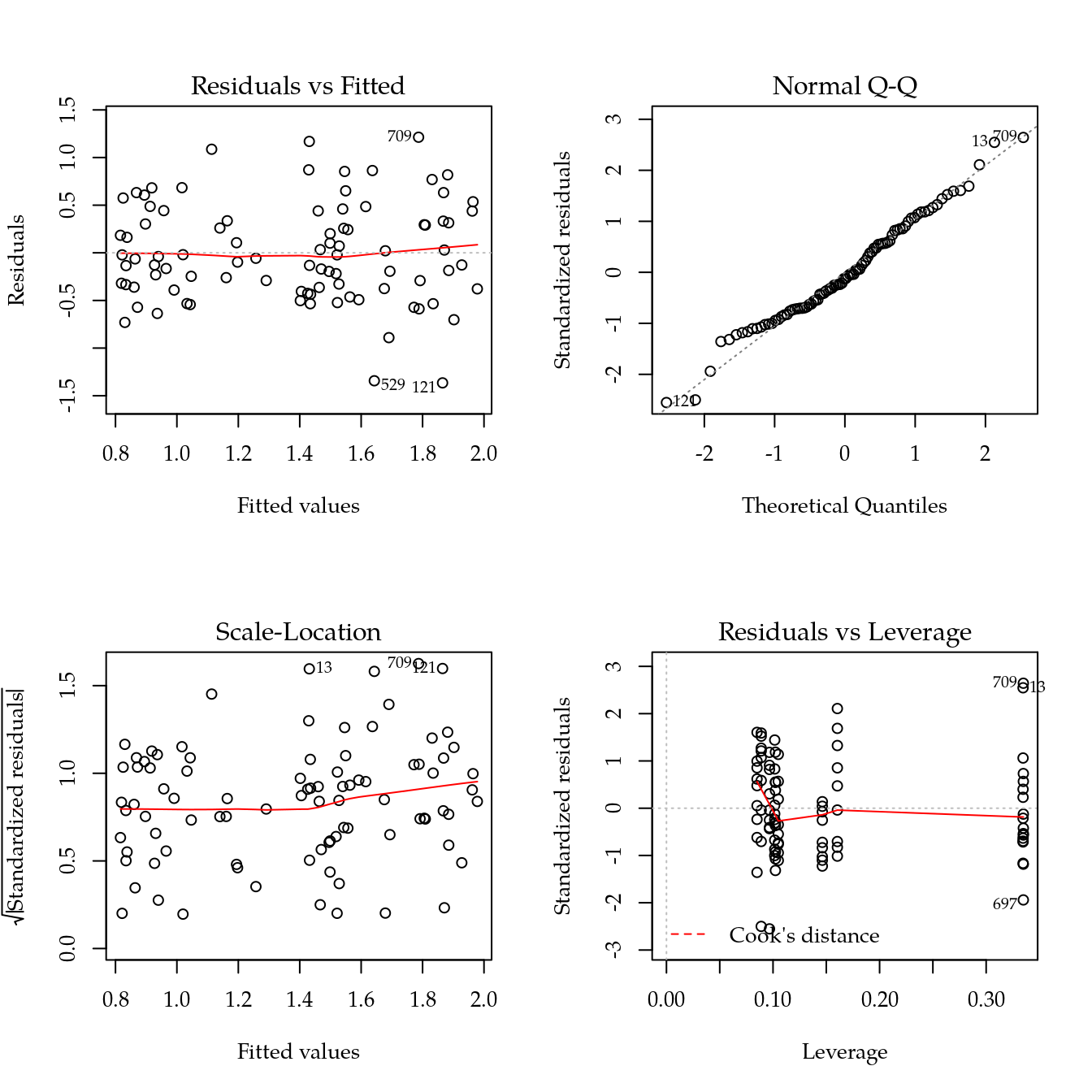
## Analysis of Variance Table
##
## Response: mg
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 0.1807 0.0903 0.2854 0.7525
## biocar 2 8.7007 4.3503 13.7440 8.037e-06 ***
## bs(dose, df = 3) 3 0.5458 0.1819 0.5748 0.6333
## biocar:bs(dose, df = 3) 6 2.9209 0.4868 1.5380 0.1773
## Residuals 76 24.0559 0.3165
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Teste para a falta de ajuste.
anova(m1, m0)## Analysis of Variance Table
##
## Model 1: mg ~ bloc + biocar + bs(dose, df = 3) + biocar:bs(dose, df = 3)
## Model 2: mg ~ bloc + biocar * P
## Res.Df RSS Df Sum of Sq F Pr(>F)
## 1 76 24.056
## 2 58 18.706 18 5.3499 0.9216 0.5571# Novos valores em grid para fazer o gráfico de bandas.
pred <- with(bio,
expand.grid(bloc = factor(levels(bloc)[1],
levels = levels(bloc)),
biocar = levels(biocar),
dose = seq(min(dose),
max(dose),
length.out = 60)))
# Extrai a especificação do tempo base splines.
form <- attr(terms(formula(m1)), "term.labels")[3]
B <- eval(parse(text = form), envir = bio)
str(B)## 'bs' num [1:90, 1:3] 0 0 0 0.373 0.373 ...
## - attr(*, "dimnames")=List of 2
## ..$ : NULL
## ..$ : chr [1:3] "1" "2" "3"
## - attr(*, "degree")= int 3
## - attr(*, "knots")= num(0)
## - attr(*, "Boundary.knots")= num [1:2] 0 5.96
## - attr(*, "intercept")= logi FALSE# Gera a matriz para os valores de predição.
X <- predict(B, pred$dos)
# Cria a matriz do modelo para fazer a predição.
L <- model.matrix(~bloc + biocar * X, data = pred)
# Dá pesos para os efeitos de blocos correspondente à LSmeans.
i <- attr(L, "assign") == 1
L[, i] <- 1/(sum(i) + 1)
# Obtém os valores preditos com IC.
ci <- confint(glht(m1, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
xyplot(mg ~ dose,
groups = biocar,
data = bio,
auto.key = list(points = FALSE,
lines = TRUE,
divide = 1,
type = "o")) +
as.layer(xyplot(Estimate ~ dose,
groups = biocar,
data = pred,
type = "l",
ly = pred$lwr,
uy = pred$upr,
cty = "bands",
alpha = 0.3,
prepanel = prepanel.cbH,
panel.groups = panel.cbH,
panel = panel.superpose))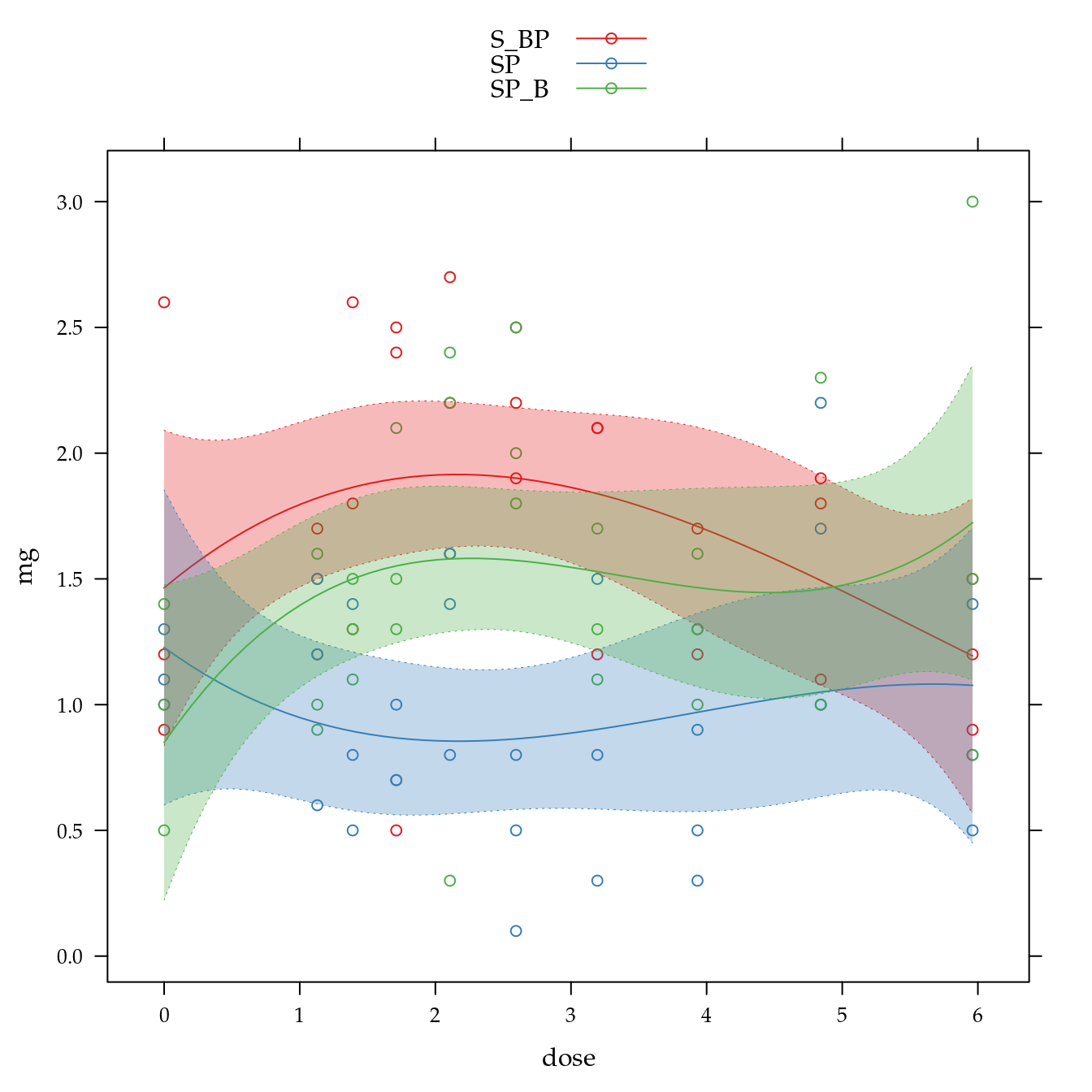
Hidrogênio + alumínio (hal)
#-----------------------------------------------------------------------
# Modelo saturado (considera dose como qualitativo).
m0 <- lm((hal)^0.33 ~ bloc + biocar * P, data = bio)
par(mfrow = c(2, 2))
plot(m0)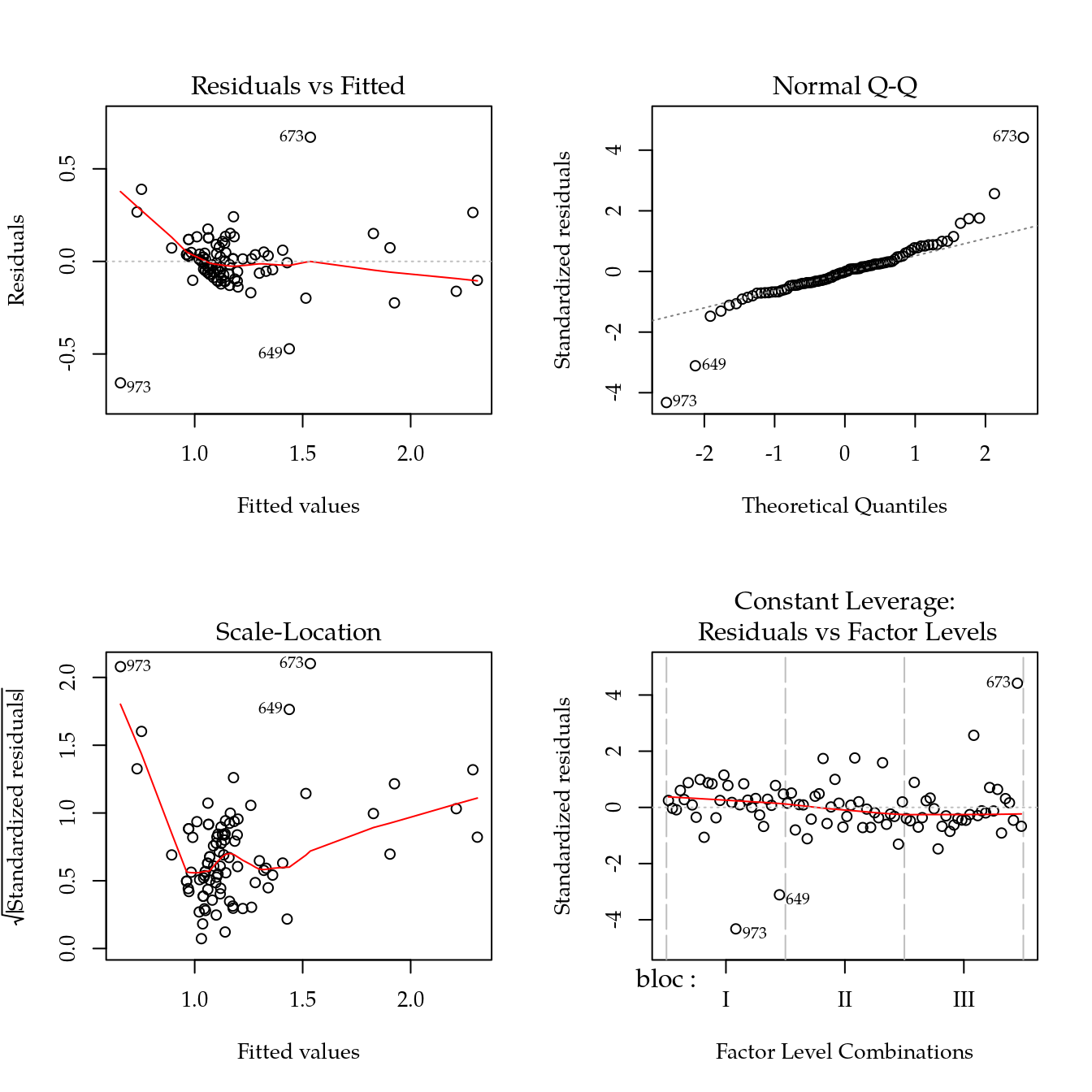
## Analysis of Variance Table
##
## Response: (hal)^0.33
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 0.1585 0.07925 2.2174 0.118
## biocar 2 1.4719 0.73593 20.5915 1.749e-07 ***
## P 9 2.5445 0.28272 7.9106 1.641e-07 ***
## biocar:P 18 3.0231 0.16795 4.6992 3.375e-06 ***
## Residuals 58 2.0729 0.03574
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1L <- LE_matrix(m0, effect = c("biocar", "P"))
pred <- equallevels(attr(L, "grid"), bio)
ci <- confint(glht(m0, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
str(pred)## 'data.frame': 30 obs. of 5 variables:
## $ biocar : Factor w/ 3 levels "S_BP","SP","SP_B": 1 2 3 1 2 3 1 2 3 1 ...
## $ P : Factor w/ 10 levels "0","1.5","3",..: 1 1 1 2 2 2 3 3 3 4 ...
## $ Estimate: num 1.02 1.07 1.08 1.1 1.12 ...
## $ lwr : num 0.802 0.85 0.862 0.877 0.904 ...
## $ upr : num 1.24 1.29 1.3 1.31 1.34 ...pch <- c(1, 4, 5)
key <- list(title = "Biocarvão",
cex.title = 1.1,
text = list(levels(bio$biocar)),
lines = list(pch = pch, lty = 1),
columns = 3,
type = "o",
divide = 1)
segplot(P ~ lwr + upr,
centers = Estimate,
data = pred,
draw = FALSE,
groups = biocar,
xlab = "Dose de P",
ylab = "Altura final (cm)",
key = key,
horizontal = FALSE,
pch = pch,
gap = 0.1,
panel = panel.groups.segplot)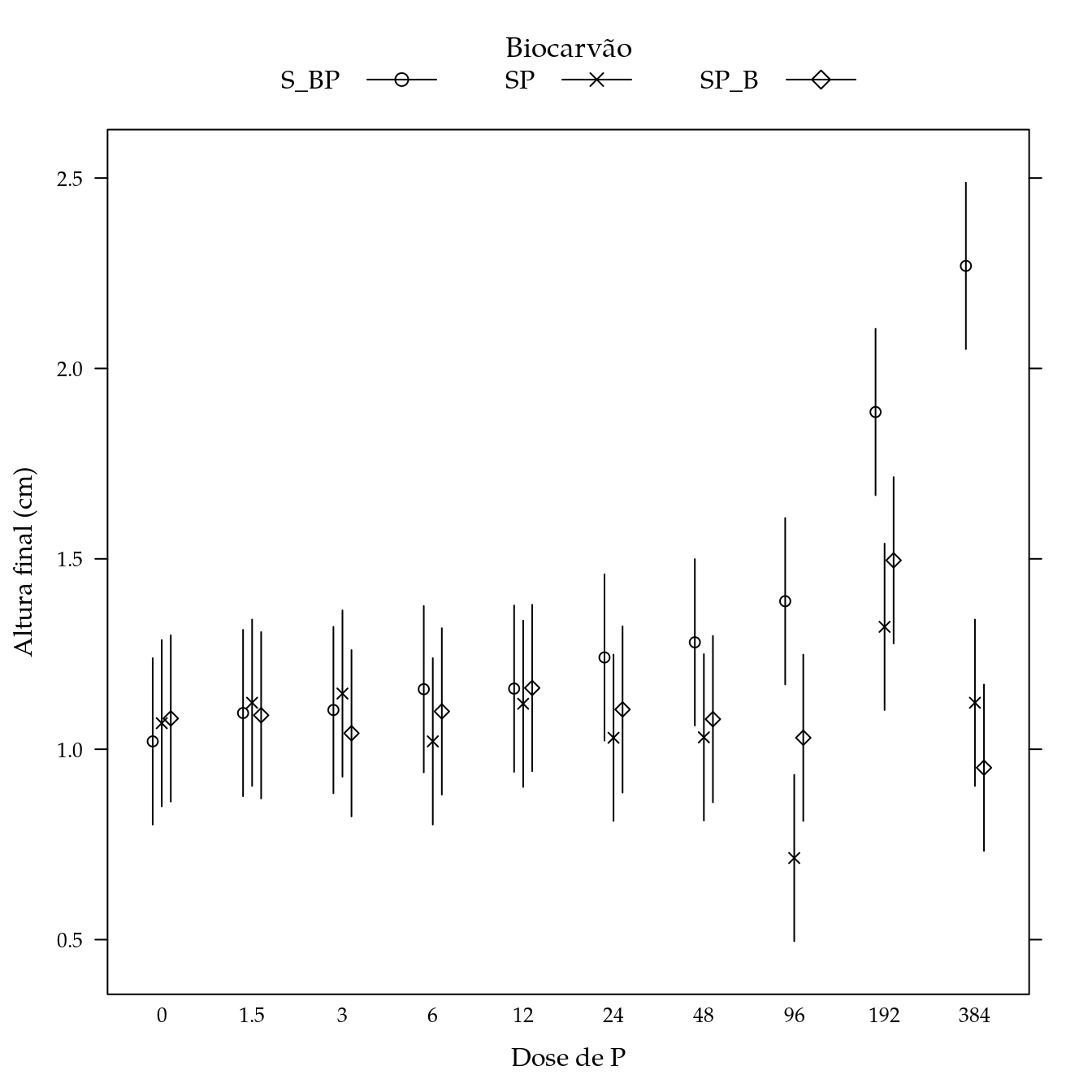
#-----------------------------------------------------------------------
# Modelo reduzido baseado em splines.
m1 <- update(m0, ~ bloc + biocar * bs(dose, df = 3))
par(mfrow = c(2, 2))
plot(m1)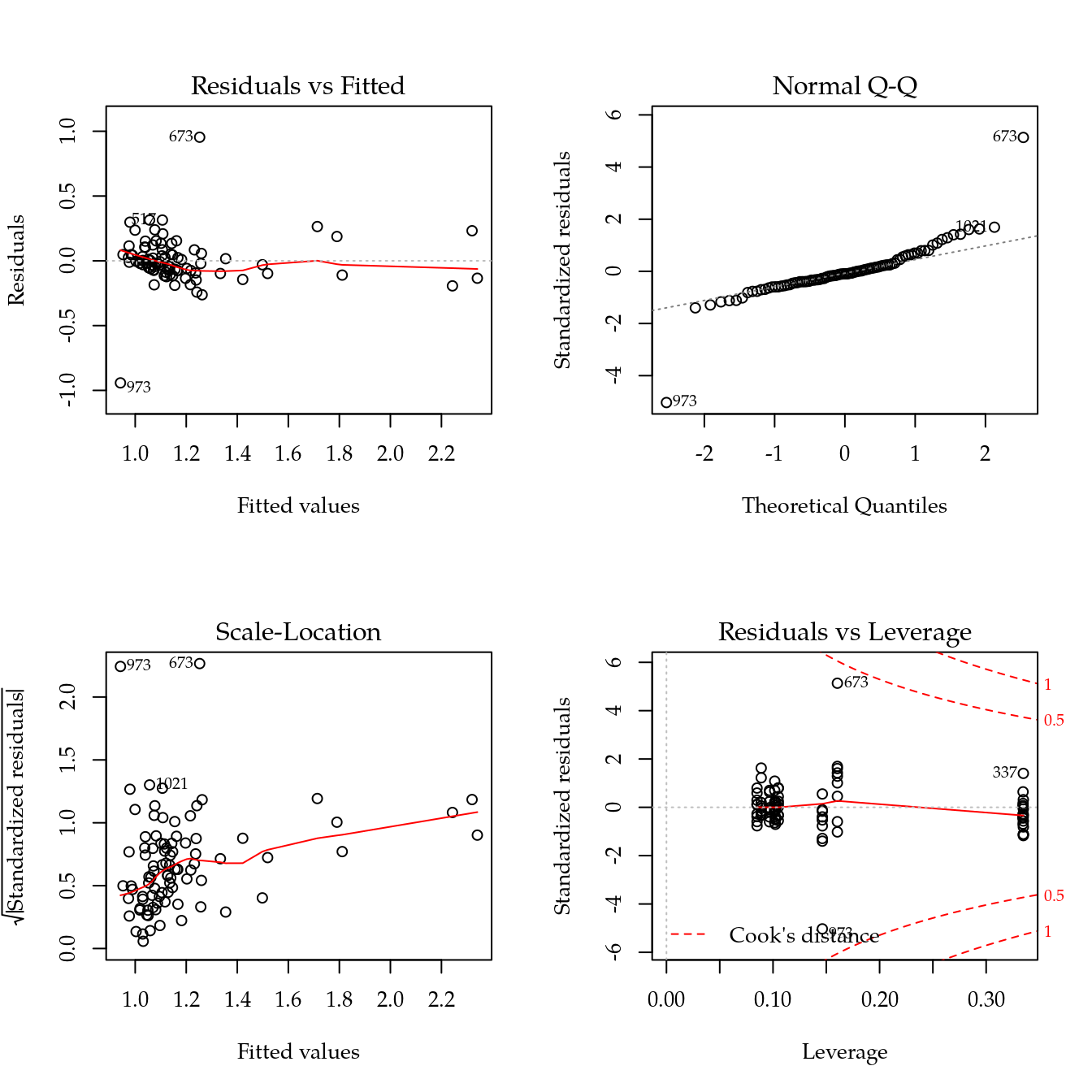
## Analysis of Variance Table
##
## Response: (hal)^0.33
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 0.1585 0.07925 1.9291 0.1523
## biocar 2 1.4719 0.73593 17.9144 4.225e-07 ***
## bs(dose, df = 3) 3 1.6763 0.55877 13.6019 3.432e-07 ***
## biocar:bs(dose, df = 3) 6 2.8420 0.47367 11.5304 3.898e-09 ***
## Residuals 76 3.1221 0.04108
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Teste para a falta de ajuste.
anova(m1, m0)## Analysis of Variance Table
##
## Model 1: (hal)^0.33 ~ bloc + biocar + bs(dose, df = 3) + biocar:bs(dose,
## df = 3)
## Model 2: (hal)^0.33 ~ bloc + biocar * P
## Res.Df RSS Df Sum of Sq F Pr(>F)
## 1 76 3.1221
## 2 58 2.0729 18 1.0492 1.631 0.08212 .
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Novos valores em grid para fazer o gráfico de bandas.
pred <- with(bio,
expand.grid(bloc = factor(levels(bloc)[1],
levels = levels(bloc)),
biocar = levels(biocar),
dose = seq(min(dose),
max(dose),
length.out = 60)))
# Extrai a especificação do tempo base splines.
form <- attr(terms(formula(m1)), "term.labels")[3]
B <- eval(parse(text = form), envir = bio)
str(B)## 'bs' num [1:90, 1:3] 0 0 0 0.373 0.373 ...
## - attr(*, "dimnames")=List of 2
## ..$ : NULL
## ..$ : chr [1:3] "1" "2" "3"
## - attr(*, "degree")= int 3
## - attr(*, "knots")= num(0)
## - attr(*, "Boundary.knots")= num [1:2] 0 5.96
## - attr(*, "intercept")= logi FALSE# Gera a matriz para os valores de predição.
X <- predict(B, pred$dos)
# Cria a matriz do modelo para fazer a predição.
L <- model.matrix(~bloc + biocar * X, data = pred)
# Dá pesos para os efeitos de blocos correspondente à LSmeans.
i <- attr(L, "assign") == 1
L[, i] <- 1/(sum(i) + 1)
# Obtém os valores preditos com IC.
ci <- confint(glht(m1, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
xyplot(hal^0.33 ~ dose,
groups = biocar,
data = bio,
auto.key = list(points = FALSE,
lines = TRUE,
divide = 1,
type = "o")) +
as.layer(xyplot(Estimate ~ dose,
groups = biocar,
data = pred,
type = "l",
ly = pred$lwr,
uy = pred$upr,
cty = "bands",
alpha = 0.3,
prepanel = prepanel.cbH,
panel.groups = panel.cbH,
panel = panel.superpose))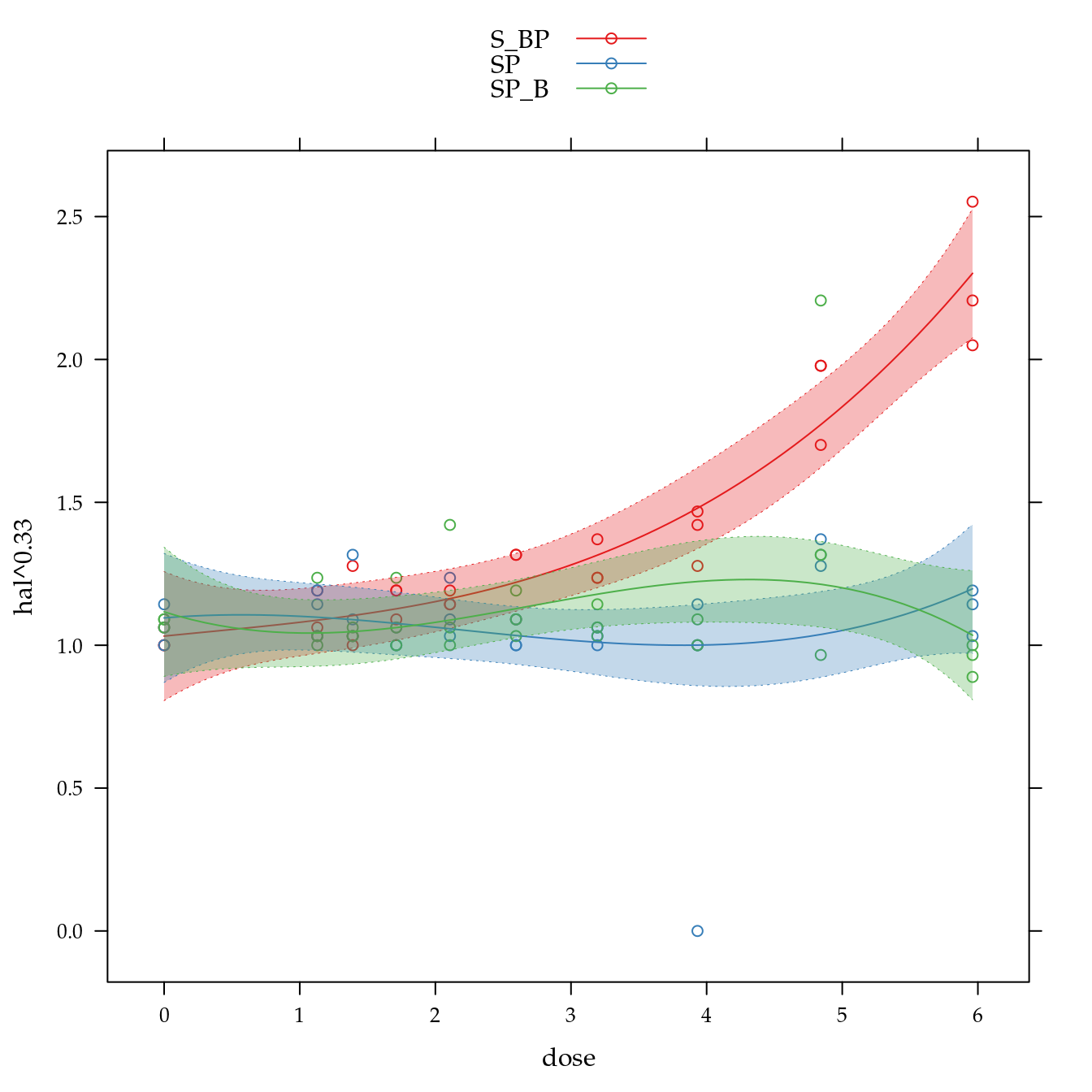
Teor de areia (are)
#-----------------------------------------------------------------------
# Modelo saturado (considera dose como qualitativo).
m0 <- lm(are ~ bloc + biocar * P, data = bio)
par(mfrow = c(2, 2))
plot(m0)
## Analysis of Variance Table
##
## Response: are
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 0.589 0.29433 0.3676 0.69402
## biocar 2 2.121 1.06033 1.3241 0.27395
## P 9 15.436 1.71511 2.1418 0.04000 *
## biocar:P 18 27.046 1.50256 1.8764 0.03692 *
## Residuals 58 46.445 0.80077
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1L <- LE_matrix(m0, effect = c("biocar", "P"))
pred <- equallevels(attr(L, "grid"), bio)
ci <- confint(glht(m0, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
str(pred)## 'data.frame': 30 obs. of 5 variables:
## $ biocar : Factor w/ 3 levels "S_BP","SP","SP_B": 1 2 3 1 2 3 1 2 3 1 ...
## $ P : Factor w/ 10 levels "0","1.5","3",..: 1 1 1 2 2 2 3 3 3 4 ...
## $ Estimate: num 70.3 70.4 70.2 70.6 70.7 ...
## $ lwr : num 69.3 69.3 69.2 69.6 69.7 ...
## $ upr : num 71.4 71.4 71.3 71.6 71.7 ...pch <- c(1, 4, 5)
key <- list(title = "Biocarvão",
cex.title = 1.1,
text = list(levels(bio$biocar)),
lines = list(pch = pch, lty = 1),
columns = 3,
type = "o",
divide = 1)
segplot(P ~ lwr + upr,
centers = Estimate,
data = pred,
draw = FALSE,
groups = biocar,
xlab = "Dose de P",
ylab = "Altura final (cm)",
key = key,
horizontal = FALSE,
pch = pch,
gap = 0.1,
panel = panel.groups.segplot)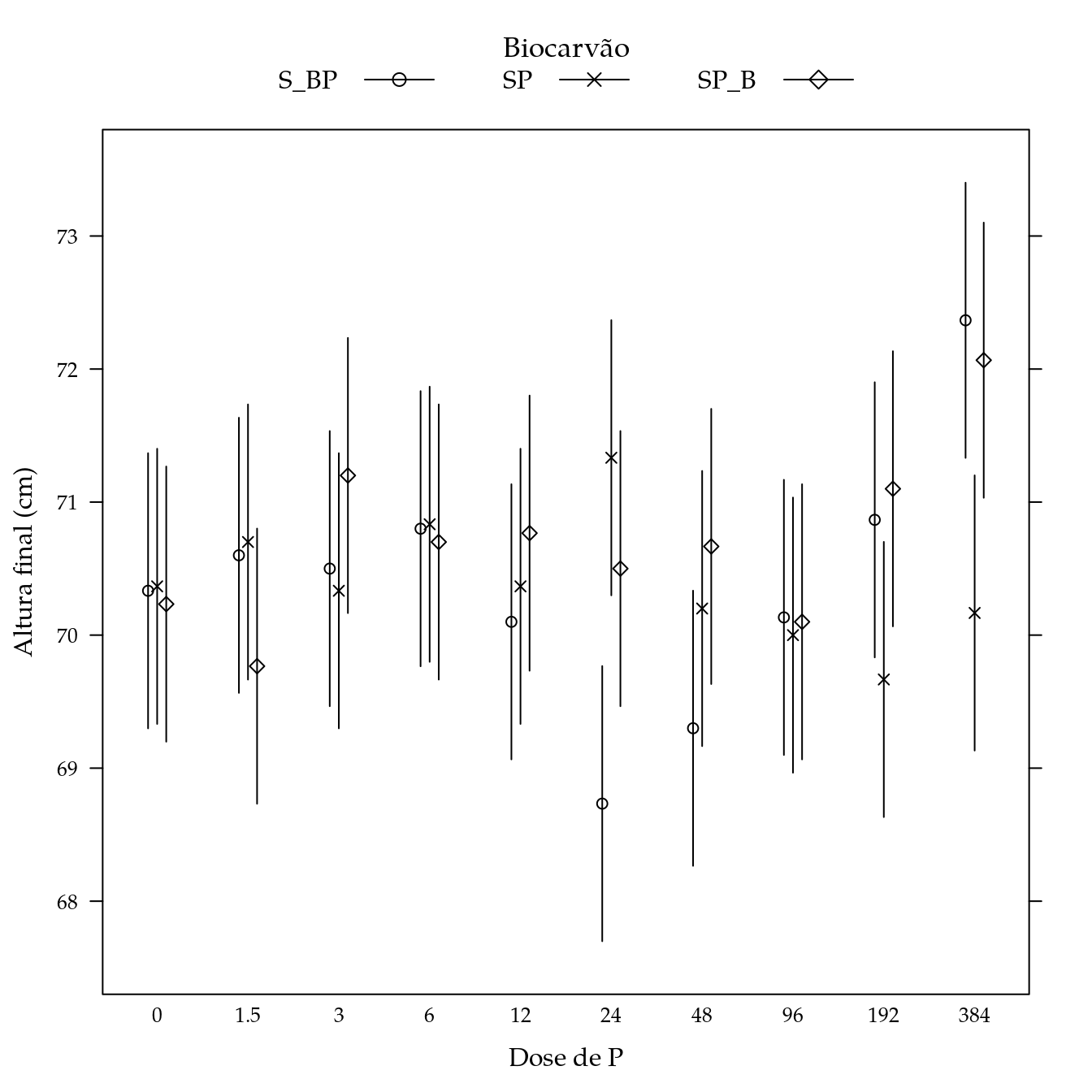
#-----------------------------------------------------------------------
# Modelo reduzido baseado em splines.
m1 <- update(m0, ~ bloc + biocar * bs(dose, df = 3))
par(mfrow = c(2, 2))
plot(m1)
## Analysis of Variance Table
##
## Response: are
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 0.589 0.2943 0.3728 0.690074
## biocar 2 2.121 1.0603 1.3429 0.267209
## bs(dose, df = 3) 3 13.803 4.6009 5.8271 0.001226 **
## biocar:bs(dose, df = 3) 6 15.116 2.5193 3.1907 0.007577 **
## Residuals 76 60.008 0.7896
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Teste para a falta de ajuste.
anova(m1, m0)## Analysis of Variance Table
##
## Model 1: are ~ bloc + biocar + bs(dose, df = 3) + biocar:bs(dose, df = 3)
## Model 2: are ~ bloc + biocar * P
## Res.Df RSS Df Sum of Sq F Pr(>F)
## 1 76 60.008
## 2 58 46.445 18 13.563 0.941 0.5359# Novos valores em grid para fazer o gráfico de bandas.
pred <- with(bio,
expand.grid(bloc = factor(levels(bloc)[1],
levels = levels(bloc)),
biocar = levels(biocar),
dose = seq(min(dose),
max(dose),
length.out = 60)))
# Extrai a especificação do tempo base splines.
form <- attr(terms(formula(m1)), "term.labels")[3]
B <- eval(parse(text = form), envir = bio)
str(B)## 'bs' num [1:90, 1:3] 0 0 0 0.373 0.373 ...
## - attr(*, "dimnames")=List of 2
## ..$ : NULL
## ..$ : chr [1:3] "1" "2" "3"
## - attr(*, "degree")= int 3
## - attr(*, "knots")= num(0)
## - attr(*, "Boundary.knots")= num [1:2] 0 5.96
## - attr(*, "intercept")= logi FALSE# Gera a matriz para os valores de predição.
X <- predict(B, pred$dos)
# Cria a matriz do modelo para fazer a predição.
L <- model.matrix(~bloc + biocar * X, data = pred)
# Dá pesos para os efeitos de blocos correspondente à LSmeans.
i <- attr(L, "assign") == 1
L[, i] <- 1/(sum(i) + 1)
# Obtém os valores preditos com IC.
ci <- confint(glht(m1, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
xyplot(are ~ dose,
groups = biocar,
data = bio,
auto.key = list(points = FALSE,
lines = TRUE,
divide = 1,
type = "o")) +
as.layer(xyplot(Estimate ~ dose,
groups = biocar,
data = pred,
type = "l",
ly = pred$lwr,
uy = pred$upr,
cty = "bands",
alpha = 0.3,
prepanel = prepanel.cbH,
panel.groups = panel.cbH,
panel = panel.superpose))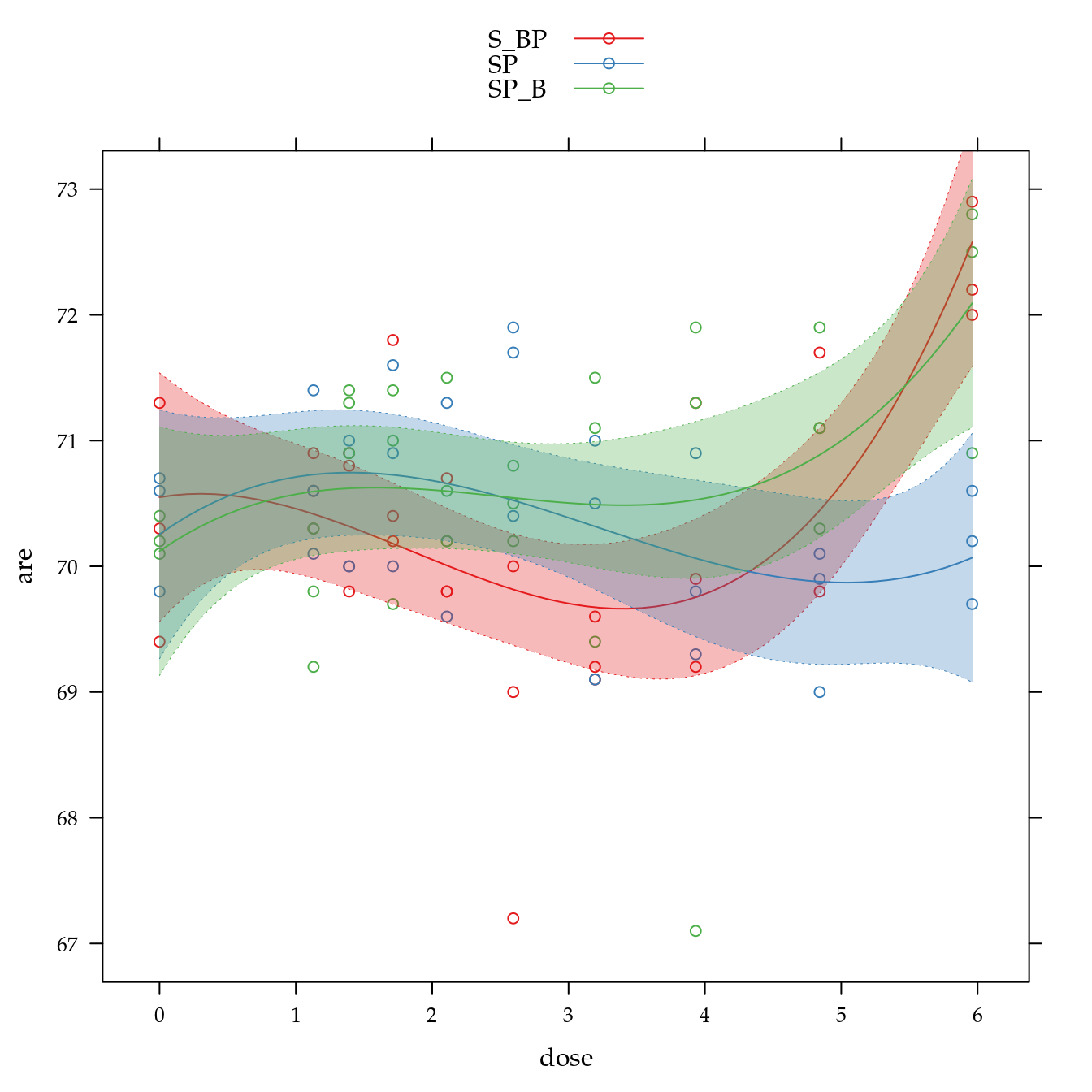
Teor de silte (sil)
#-----------------------------------------------------------------------
# Modelo saturado (considera dose como qualitativo).
m0 <- lm(sil ~ bloc + biocar * P, data = bio[bio$sil < 30, ])
par(mfrow = c(2, 2))
plot(m0)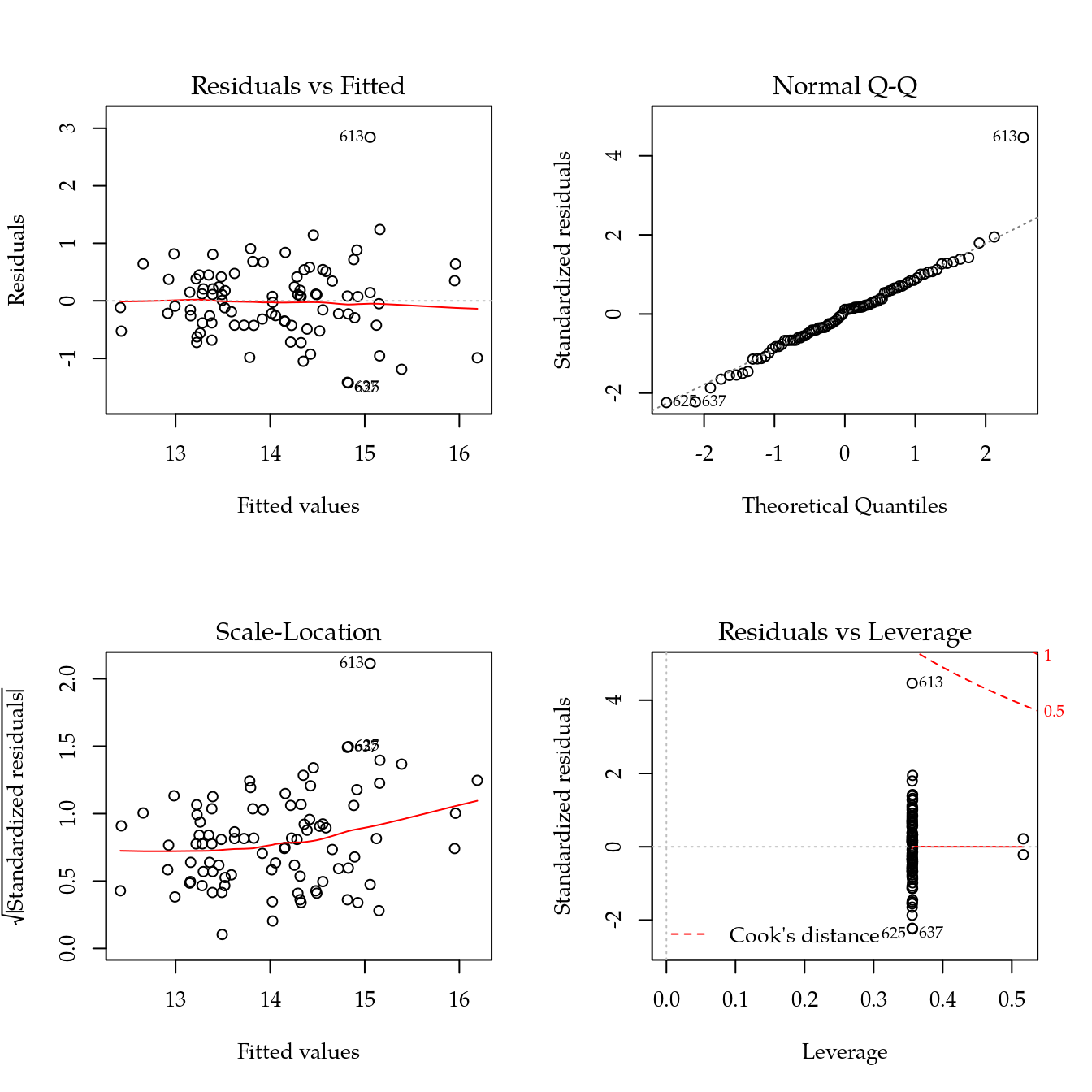
## Analysis of Variance Table
##
## Response: sil
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 1.012 0.5060 0.8037 0.452678
## biocar 2 22.046 11.0231 17.5080 1.181e-06 ***
## P 9 18.810 2.0900 3.3195 0.002538 **
## biocar:P 18 12.927 0.7182 1.1407 0.340153
## Residuals 57 35.887 0.6296
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1L <- LE_matrix(m0, effect = c("biocar", "P"))
pred <- equallevels(attr(L, "grid"), bio)
ci <- confint(glht(m0, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
str(pred)## 'data.frame': 30 obs. of 5 variables:
## $ biocar : Factor w/ 3 levels "S_BP","SP","SP_B": 1 2 3 1 2 3 1 2 3 1 ...
## $ P : Factor w/ 10 levels "0","1.5","3",..: 1 1 1 2 2 2 3 3 3 4 ...
## $ Estimate: num 14.4 13.9 14.4 14.1 13.4 ...
## $ lwr : num 13.5 13 13.5 13.2 12.4 ...
## $ upr : num 15.3 14.8 15.3 15 14.3 ...pch <- c(1, 4, 5)
key <- list(title = "Biocarvão",
cex.title = 1.1,
text = list(levels(bio$biocar)),
lines = list(pch = pch, lty = 1),
columns = 3,
type = "o",
divide = 1)
segplot(P ~ lwr + upr,
centers = Estimate,
data = pred,
draw = FALSE,
groups = biocar,
xlab = "Dose de P",
ylab = "Altura final (cm)",
key = key,
horizontal = FALSE,
pch = pch,
gap = 0.1,
panel = panel.groups.segplot)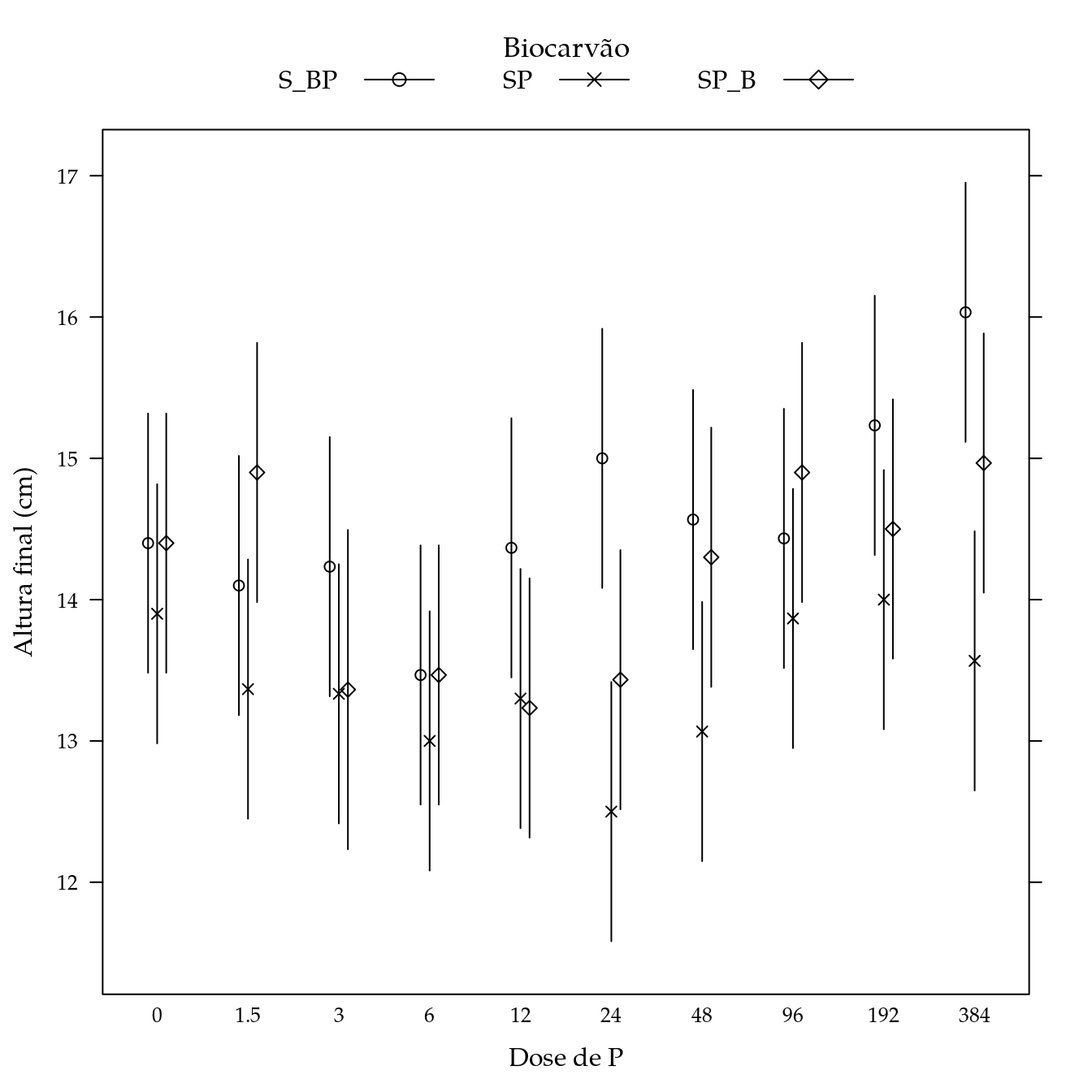
#-----------------------------------------------------------------------
# Modelo reduzido baseado em splines.
m1 <- update(m0, ~ bloc + biocar * bs(dose, df = 3))
par(mfrow = c(2, 2))
plot(m1)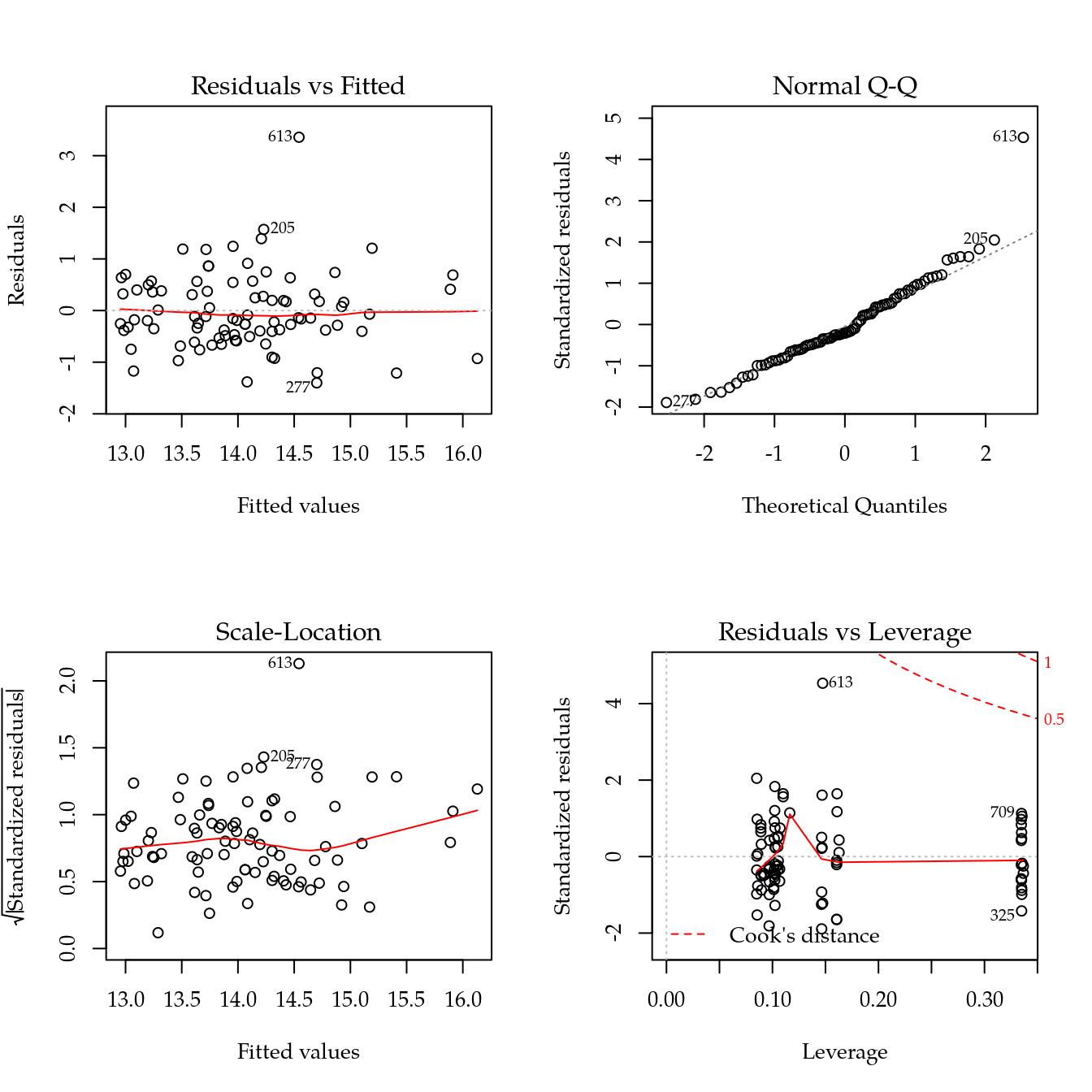
## Analysis of Variance Table
##
## Response: sil
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 1.012 0.5060 0.7867 0.4591
## biocar 2 22.046 11.0231 17.1370 7.420e-07 ***
## bs(dose, df = 3) 3 15.861 5.2871 8.2197 8.456e-05 ***
## biocar:bs(dose, df = 3) 6 3.520 0.5867 0.9122 0.4910
## Residuals 75 48.242 0.6432
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Teste para a falta de ajuste.
anova(m1, m0)## Analysis of Variance Table
##
## Model 1: sil ~ bloc + biocar + bs(dose, df = 3) + biocar:bs(dose, df = 3)
## Model 2: sil ~ bloc + biocar * P
## Res.Df RSS Df Sum of Sq F Pr(>F)
## 1 75 48.242
## 2 57 35.887 18 12.355 1.0902 0.385# Novos valores em grid para fazer o gráfico de bandas.
pred <- with(bio,
expand.grid(bloc = factor(levels(bloc)[1],
levels = levels(bloc)),
biocar = levels(biocar),
dose = seq(min(dose),
max(dose),
length.out = 60)))
# Extrai a especificação do tempo base splines.
form <- attr(terms(formula(m1)), "term.labels")[3]
B <- eval(parse(text = form), envir = bio)
str(B)## 'bs' num [1:90, 1:3] 0 0 0 0.373 0.373 ...
## - attr(*, "dimnames")=List of 2
## ..$ : NULL
## ..$ : chr [1:3] "1" "2" "3"
## - attr(*, "degree")= int 3
## - attr(*, "knots")= num(0)
## - attr(*, "Boundary.knots")= num [1:2] 0 5.96
## - attr(*, "intercept")= logi FALSE# Gera a matriz para os valores de predição.
X <- predict(B, pred$dos)
# Cria a matriz do modelo para fazer a predição.
L <- model.matrix(~bloc + biocar * X, data = pred)
# Dá pesos para os efeitos de blocos correspondente à LSmeans.
i <- attr(L, "assign") == 1
L[, i] <- 1/(sum(i) + 1)
# Obtém os valores preditos com IC.
ci <- confint(glht(m1, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
xyplot(sil ~ dose,
groups = biocar,
data = bio[bio$sil < 30, ],
auto.key = list(points = FALSE,
lines = TRUE,
divide = 1,
type = "o")) +
as.layer(xyplot(Estimate ~ dose,
groups = biocar,
data = pred,
type = "l",
ly = pred$lwr,
uy = pred$upr,
cty = "bands",
alpha = 0.3,
prepanel = prepanel.cbH,
panel.groups = panel.cbH,
panel = panel.superpose))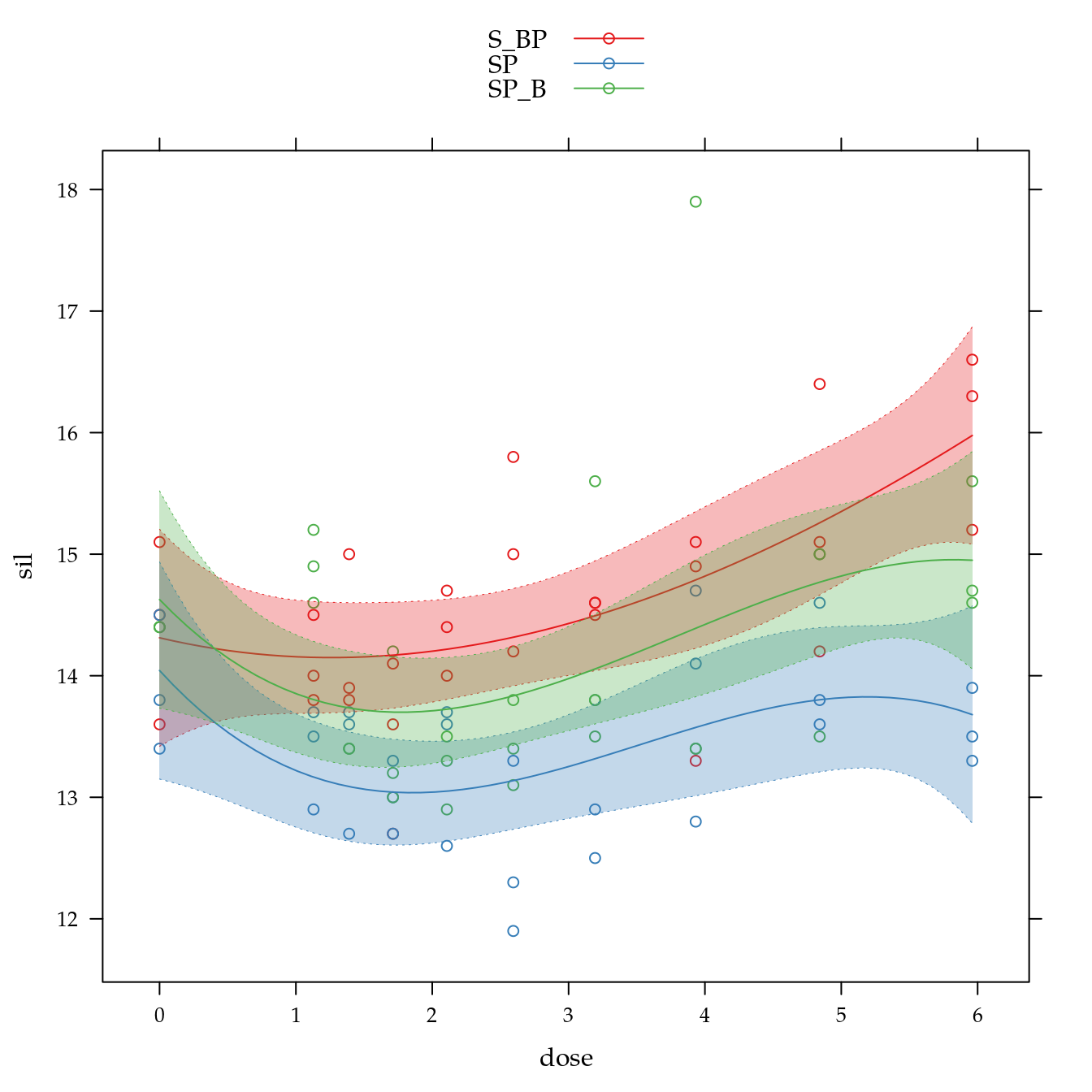
Teor de argila (arg)
#-----------------------------------------------------------------------
# Modelo saturado (considera dose como qualitativo).
m0 <- lm(arg ~ bloc + biocar * P, data = bio)
par(mfrow = c(2, 2))
plot(m0)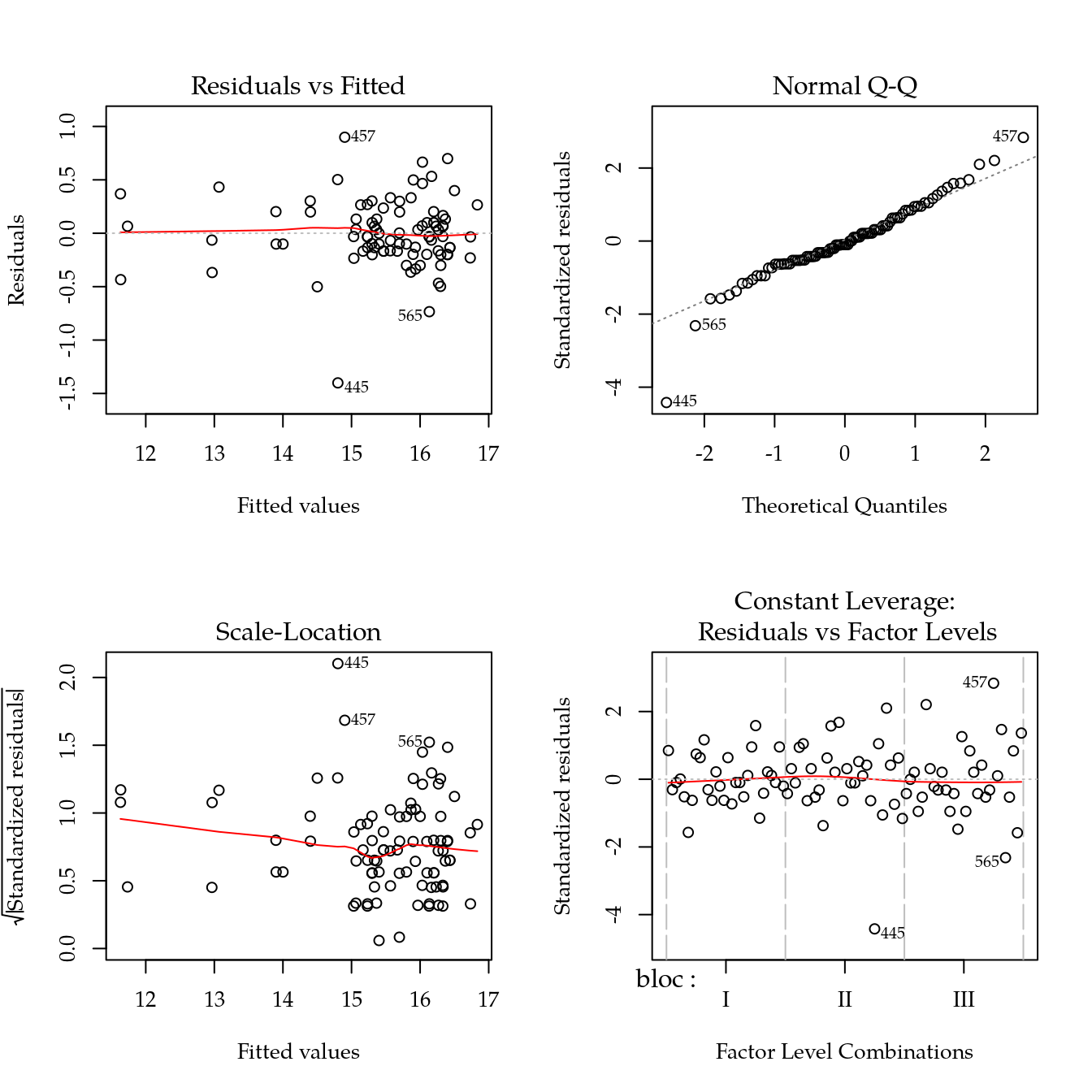
## Analysis of Variance Table
##
## Response: arg
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 0.207 0.1034 0.6637 0.5188
## biocar 2 26.774 13.3871 85.8929 < 2.2e-16 ***
## P 9 44.603 4.9559 31.7976 < 2.2e-16 ***
## biocar:P 18 29.666 1.6481 10.5743 2.41e-12 ***
## Residuals 58 9.040 0.1559
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1L <- LE_matrix(m0, effect = c("biocar", "P"))
pred <- equallevels(attr(L, "grid"), bio)
ci <- confint(glht(m0, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
str(pred)## 'data.frame': 30 obs. of 5 variables:
## $ biocar : Factor w/ 3 levels "S_BP","SP","SP_B": 1 2 3 1 2 3 1 2 3 1 ...
## $ P : Factor w/ 10 levels "0","1.5","3",..: 1 1 1 2 2 2 3 3 3 4 ...
## $ Estimate: num 15.3 15.7 15.4 15.3 15.9 ...
## $ lwr : num 14.8 15.3 14.9 14.9 15.5 ...
## $ upr : num 15.7 16.2 15.9 15.8 16.4 ...pch <- c(1, 4, 5)
key <- list(title = "Biocarvão",
cex.title = 1.1,
text = list(levels(bio$biocar)),
lines = list(pch = pch, lty = 1),
columns = 3,
type = "o",
divide = 1)
segplot(P ~ lwr + upr,
centers = Estimate,
data = pred,
draw = FALSE,
groups = biocar,
xlab = "Dose de P",
ylab = "Altura final (cm)",
key = key,
horizontal = FALSE,
pch = pch,
gap = 0.1,
panel = panel.groups.segplot)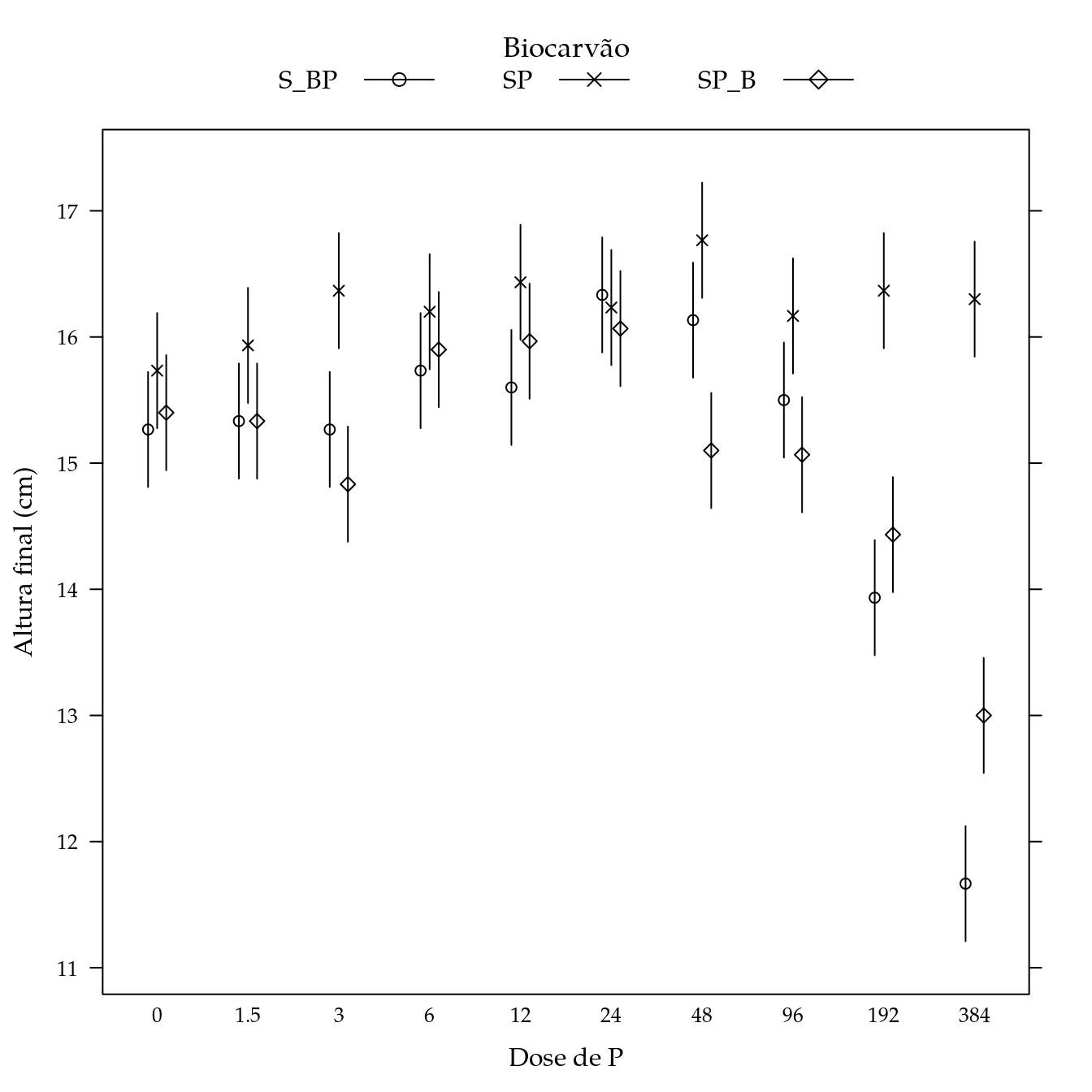
#-----------------------------------------------------------------------
# Modelo reduzido baseado em splines.
m1 <- update(m0, ~ bloc + biocar * bs(dose, df = 3))
par(mfrow = c(2, 2))
plot(m1)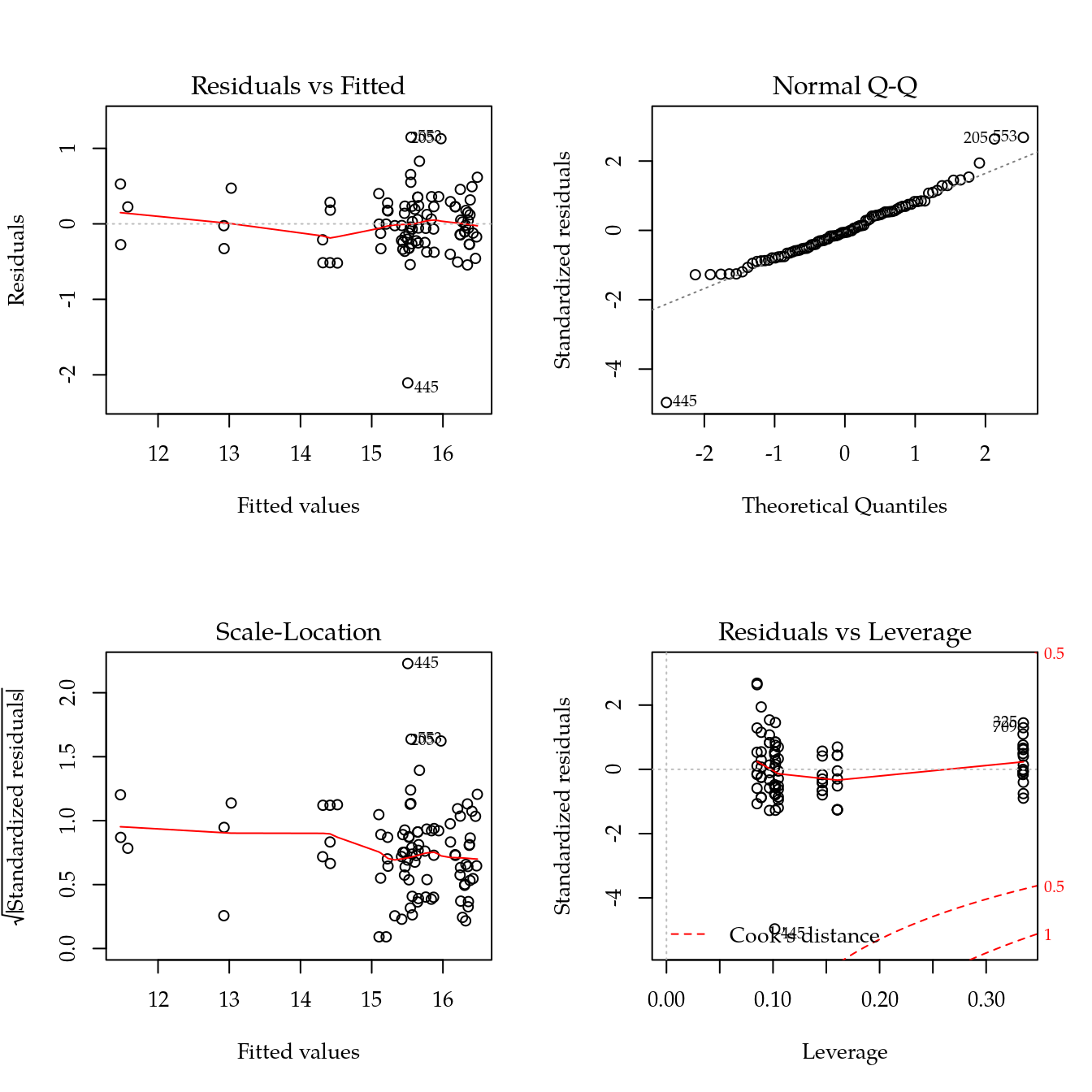
## Analysis of Variance Table
##
## Response: arg
## Df Sum Sq Mean Sq F value Pr(>F)
## bloc 2 0.207 0.1034 0.5164 0.5988
## biocar 2 26.774 13.3871 66.8246 < 2.2e-16 ***
## bs(dose, df = 3) 3 42.453 14.1509 70.6370 < 2.2e-16 ***
## biocar:bs(dose, df = 3) 6 25.631 4.2718 21.3237 1.622e-14 ***
## Residuals 76 15.225 0.2003
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Teste para a falta de ajuste.
anova(m1, m0)## Analysis of Variance Table
##
## Model 1: arg ~ bloc + biocar + bs(dose, df = 3) + biocar:bs(dose, df = 3)
## Model 2: arg ~ bloc + biocar * P
## Res.Df RSS Df Sum of Sq F Pr(>F)
## 1 76 15.2252
## 2 58 9.0398 18 6.1855 2.2048 0.01217 *
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Novos valores em grid para fazer o gráfico de bandas.
pred <- with(bio,
expand.grid(bloc = factor(levels(bloc)[1],
levels = levels(bloc)),
biocar = levels(biocar),
dose = seq(min(dose),
max(dose),
length.out = 60)))
# Extrai a especificação do tempo base splines.
form <- attr(terms(formula(m1)), "term.labels")[3]
B <- eval(parse(text = form), envir = bio)
str(B)## 'bs' num [1:90, 1:3] 0 0 0 0.373 0.373 ...
## - attr(*, "dimnames")=List of 2
## ..$ : NULL
## ..$ : chr [1:3] "1" "2" "3"
## - attr(*, "degree")= int 3
## - attr(*, "knots")= num(0)
## - attr(*, "Boundary.knots")= num [1:2] 0 5.96
## - attr(*, "intercept")= logi FALSE# Gera a matriz para os valores de predição.
X <- predict(B, pred$dos)
# Cria a matriz do modelo para fazer a predição.
L <- model.matrix(~bloc + biocar * X, data = pred)
# Dá pesos para os efeitos de blocos correspondente à LSmeans.
i <- attr(L, "assign") == 1
L[, i] <- 1/(sum(i) + 1)
# Obtém os valores preditos com IC.
ci <- confint(glht(m1, linfct = L),
calpha = univariate_calpha())$confint
# Junta as colunas estimadas com os valores do grid.
pred <- cbind(pred, as.data.frame(ci))
xyplot(arg ~ dose,
groups = biocar,
data = bio,
auto.key = list(points = FALSE,
lines = TRUE,
divide = 1,
type = "o")) +
as.layer(xyplot(Estimate ~ dose,
groups = biocar,
data = pred,
type = "l",
ly = pred$lwr,
uy = pred$upr,
cty = "bands",
alpha = 0.3,
prepanel = prepanel.cbH,
panel.groups = panel.cbH,
panel = panel.superpose))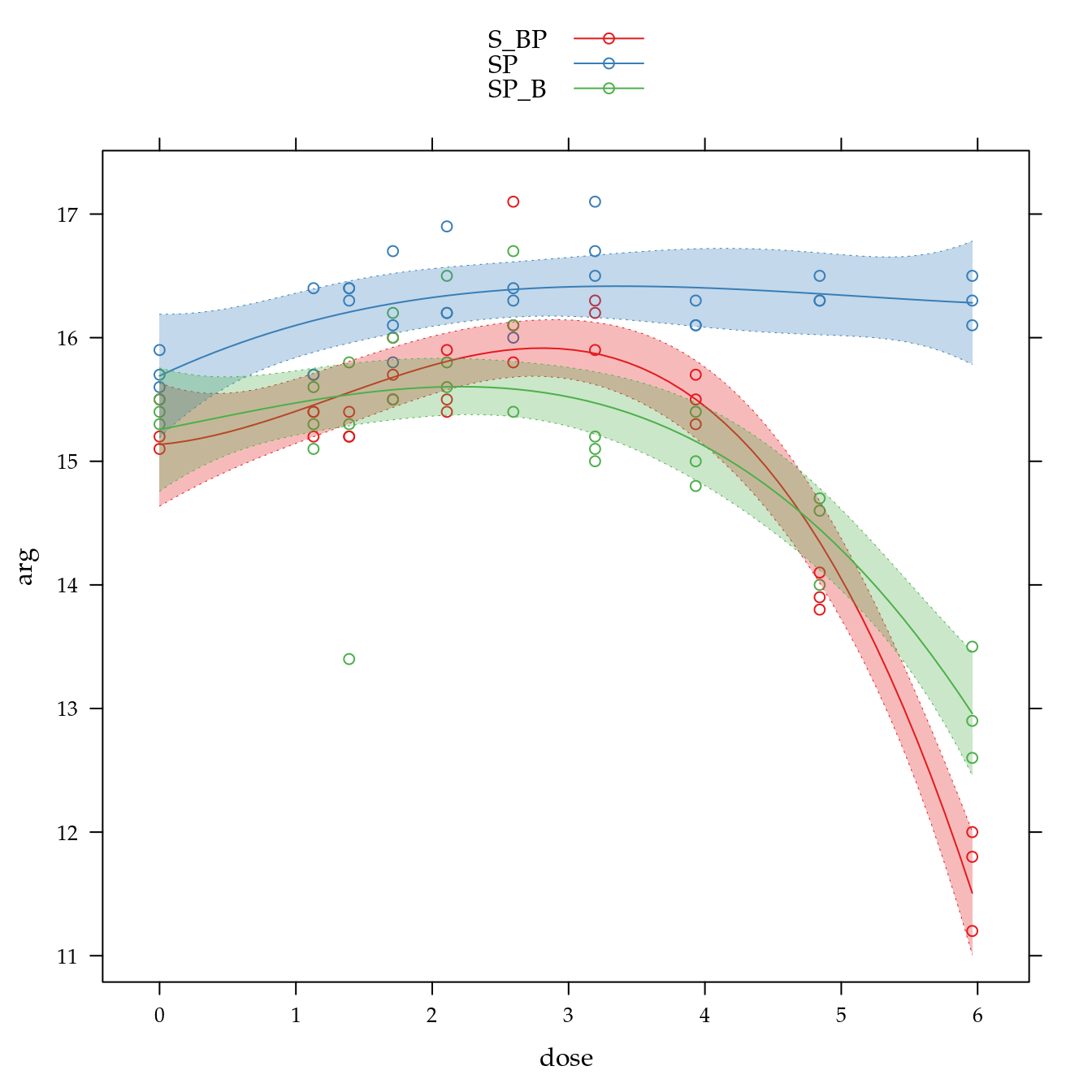
Informações da Sessão
## Atualizado em 11 de julho de 2019.
##
## R version 3.6.1 (2019-07-05)
## Platform: x86_64-pc-linux-gnu (64-bit)
## Running under: Ubuntu 18.04.2 LTS
##
## Matrix products: default
## BLAS: /usr/lib/x86_64-linux-gnu/blas/libblas.so.3.7.1
## LAPACK: /usr/lib/x86_64-linux-gnu/lapack/liblapack.so.3.7.1
##
## locale:
## [1] LC_CTYPE=pt_BR.UTF-8 LC_NUMERIC=C
## [3] LC_TIME=pt_BR.UTF-8 LC_COLLATE=en_US.UTF-8
## [5] LC_MONETARY=pt_BR.UTF-8 LC_MESSAGES=en_US.UTF-8
## [7] LC_PAPER=pt_BR.UTF-8 LC_NAME=C
## [9] LC_ADDRESS=C LC_TELEPHONE=C
## [11] LC_MEASUREMENT=pt_BR.UTF-8 LC_IDENTIFICATION=C
##
## attached base packages:
## [1] splines stats graphics grDevices utils datasets
## [7] methods base
##
## other attached packages:
## [1] multcomp_1.4-10 TH.data_1.0-10 MASS_7.3-51.4
## [4] survival_2.44-1.1 mvtnorm_1.0-11 doBy_4.6-2
## [7] reshape_0.8.8 EACS_0.0-7 wzRfun_0.91
## [10] captioner_2.2.3 latticeExtra_0.6-28 RColorBrewer_1.1-2
## [13] lattice_0.20-38 knitr_1.23
##
## loaded via a namespace (and not attached):
## [1] zoo_1.8-6 tidyselect_0.2.5 xfun_0.8
## [4] remotes_2.1.0 purrr_0.3.2 testthat_2.1.1
## [7] htmltools_0.3.6 usethis_1.5.1 yaml_2.2.0
## [10] rlang_0.4.0 pkgbuild_1.0.3 pkgdown_1.3.0
## [13] pillar_1.4.2 glue_1.3.1 withr_2.1.2
## [16] sessioninfo_1.1.1 plyr_1.8.4 stringr_1.4.0
## [19] commonmark_1.7 devtools_2.1.0 codetools_0.2-16
## [22] memoise_1.1.0 evaluate_0.14 callr_3.3.0
## [25] ps_1.3.0 Rcpp_1.0.1 backports_1.1.4
## [28] desc_1.2.0 pkgload_1.0.2 fs_1.3.1
## [31] digest_0.6.20 stringi_1.4.3 processx_3.4.0
## [34] dplyr_0.8.3 grid_3.6.1 rprojroot_1.3-2
## [37] cli_1.1.0 tools_3.6.1 sandwich_2.5-1
## [40] magrittr_1.5 tibble_2.1.3 crayon_1.3.4
## [43] pkgconfig_2.0.2 Matrix_1.2-17 xml2_1.2.0
## [46] prettyunits_1.0.2 assertthat_0.2.1 rmarkdown_1.13
## [49] roxygen2_6.1.1 rstudioapi_0.10 R6_2.4.0
## [52] compiler_3.6.1