Capítulo 16 Delineamento em látice
https://pbgworks.org/sites/pbgworks.org/files/user/4/Lattice_seminar.pdf
RamalhoTb11.1: látice quadrado balanceado.RamalhoEg11.4: látice quadrado parcialmente balanceado.
library(car) # Anova().
library(emmeans) # emmeans().
library(multcomp) # glht().
library(lme4) # lmer().
library(lmerTest) # anova() e summary() para classe lmerMod.
library(tidyverse) # Manipulação e visualização.
# Funções.
source("mpaer_functions.R")TODO: látice quadrado e látice retangular. Qual a diferença entre os BIB tipo 1 e 2?
16.1 RamalhoTb11.1
O dados em RamalhoTb11.1 são de um experimento para a avaliação de 16 cultivares de sorgo consudizo no delineamento de látice quadrado balanceado. A produção (kg/ha) das parcelas é a variável resposta.
Nesse ensaio foram avaliados \(t = 16\) cultivares usando \(b = 20\) blocos de tamanho \(k = 4\) agrupados em \(r = 5\) repetições, portanto são \(tr = bk = 80\) unidades experimentais. Este experimento é um látice quadrado pois \(t = k^2\) e \(r = k + 1\).
# Importação dos dados.
da <- as_tibble(labestData::RamalhoTb11.1)
da$rept <- gl(5, 16)
str(da)## Classes 'tbl_df', 'tbl' and 'data.frame': 80 obs. of 4 variables:
## $ bloc: Factor w/ 4 levels "1","2","3","4": 1 1 1 1 2 2 2 2 3 3 ...
## $ cult: Factor w/ 16 levels "1","2","3","4",..: 1 2 3 4 5 6 7 8 9 10 ...
## $ prod: num 10.2 10.7 10.8 12.7 9.3 6.4 10.5 10.6 9.2 5.2 ...
## $ rept: Factor w/ 5 levels "1","2","3","4",..: 1 1 1 1 1 1 1 1 1 1 ...# Os blocos são de tamanho 4, dispostos em 5 repetições de 4 blocos cada
# para acomodar 16 cultivares.
# with(da, table(bloc, cult, rept)) %>%
# addmargins()
with(da, table(rept, bloc, cult)) %>%
ftable()## cult 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16
## rept bloc
## 1 1 1 1 1 1 0 0 0 0 0 0 0 0 0 0 0 0
## 2 0 0 0 0 1 1 1 1 0 0 0 0 0 0 0 0
## 3 0 0 0 0 0 0 0 0 1 1 1 1 0 0 0 0
## 4 0 0 0 0 0 0 0 0 0 0 0 0 1 1 1 1
## 2 1 1 0 0 0 1 0 0 0 1 0 0 0 1 0 0 0
## 2 0 1 0 0 0 1 0 0 0 1 0 0 0 1 0 0
## 3 0 0 1 0 0 0 1 0 0 0 1 0 0 0 1 0
## 4 0 0 0 1 0 0 0 1 0 0 0 1 0 0 0 1
## 3 1 1 0 0 0 0 1 0 0 0 0 1 0 0 0 0 1
## 2 0 1 0 0 1 0 0 0 0 0 0 1 0 0 1 0
## 3 0 0 1 0 0 0 0 1 1 0 0 0 0 1 0 0
## 4 0 0 0 1 0 0 1 0 0 1 0 0 1 0 0 0
## 4 1 1 0 0 0 0 0 1 0 0 0 0 1 0 1 0 0
## 2 0 1 0 0 0 0 0 1 0 0 1 0 1 0 0 0
## 3 0 0 1 0 1 0 0 0 0 1 0 0 0 0 0 1
## 4 0 0 0 1 0 1 0 0 1 0 0 0 0 0 1 0
## 5 1 1 0 0 0 0 0 0 1 0 1 0 0 0 0 1 0
## 2 0 1 0 0 0 0 1 0 1 0 0 0 0 0 0 1
## 3 0 0 1 0 0 1 0 0 0 0 0 1 1 0 0 0
## 4 0 0 0 1 1 0 0 0 0 0 1 0 0 1 0 0library(igraph)
# Cria bloco dentro de repetição (identificador único).
da$cond <- with(da, interaction(rept, bloc))
# Determina as conexões entre tratamentos.
edg <- by(data = as.integer(da$cult),
INDICES = da$cond,
FUN = combn,
m = 2) %>%
flatten_int()
# Exibe o grafo.
ghp <- graph(edg, directed = FALSE)
plot(ghp,
layout = layout_in_circle,
edge.curved = FALSE)O modelo de efeitos fixos é \[ \begin{aligned} Y | i, j, k &\sim \text{Normal}(\mu_{ijk}, \sigma^2)\\ \mu_{ijk} &= R_i + B_{j(i)} + T_k\\ \sigma^2 &\propto 1 \end{aligned} \] em que:
- \(Y\) é a variável aleatória observada na repetição \(i\), bloco \(j\) e tratamento \(k\) que assume-se ter distribuição normal com média condicionada a especificação dos efeitos das fontes de variação e com variância constante (\(\sigma^2\)).
- \(\mu_{ijk}\) é a média composta pelos efeito da repetição (\(R_i\)), do bloco dentro da repetição (\(B_{j(i)}\)) e do tratamento (\(T_k\)) em um preditor linear de efeitos aditivos.
No modelo de efeitos fixos, todos os termos do preditor linear são parâmetros. Logo, serão estimados \(1 + (r - 1) + r(k - 1) + (t - 1)\) parâmetros para o preditor da média. O modelo é ajustado aos dados por mínimos quadrados (ordinários).
# Modelo de efeitos fixos.
m0 <- lm(terms(prod ~ rept/bloc + cult, keep.order = TRUE),
data = da)
# Quado de anova com hipóteses marginais.
Anova(m0)## Anova Table (Type II tests)
##
## Response: prod
## Sum Sq Df F value Pr(>F)
## rept 20.136 4 1.7708 0.151399
## rept:bloc 123.942 15 2.9066 0.002877 **
## cult 105.224 15 2.4676 0.009903 **
## Residuals 127.926 45
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Médias marginais ajustadas.
emm <- emmeans(m0, specs = ~cult)## NOTE: A nesting structure was detected in the fitted model:
## bloc %in% reptemm## cult emmean SE df lower.CL upper.CL
## 1 8.12 0.838 45 6.43 9.81
## 2 7.69 0.838 45 6.00 9.38
## 3 8.44 0.838 45 6.75 10.13
## 4 9.06 0.838 45 7.37 10.75
## 5 9.77 0.838 45 8.08 11.45
## 6 6.88 0.838 45 5.19 8.56
## 7 10.59 0.838 45 8.90 12.28
## 8 10.70 0.838 45 9.02 12.39
## 9 10.43 0.838 45 8.75 12.12
## 10 7.87 0.838 45 6.18 9.55
## 11 7.32 0.838 45 5.63 9.01
## 12 8.04 0.838 45 6.35 9.73
## 13 7.51 0.838 45 5.83 9.20
## 14 9.85 0.838 45 8.17 11.54
## 15 7.66 0.838 45 5.97 9.35
## 16 6.91 0.838 45 5.22 8.60
##
## Results are averaged over the levels of: bloc, rept
## Confidence level used: 0.95# Extração da matriz de funções lineares.
L <- attr(emm, "linfct")
grid <- attr(emm, "grid")
rownames(L) <- grid[[1]]
# Entenda como são obtidas as médias marginais.
MASS::fractions(t(L[1:4, ])) %>%
unique()## 1 2 3 4
## (Intercept) 1 1 1 1
## rept2 1/5 1/5 1/5 1/5
## rept1:bloc2 1/20 1/20 1/20 1/20
## cult2 0 1 0 0
## cult3 0 0 1 0
## cult4 0 0 0 1
## cult5 0 0 0 0# Comparações múltiplas a 10%.
results_m0 <- wzRfun::apmc(X = L,
model = m0,
focus = "cult",
test = "fdr",
level = 0.1) %>%
mutate(cld = wzRfun::ordered_cld(let = cld, means = fit))
results_m0## cult fit lwr upr cld
## 1 1 8.12125 6.714319 9.528181 ab
## 2 2 7.69000 6.283069 9.096931 ab
## 3 3 8.44000 7.033069 9.846931 ab
## 4 4 9.05875 7.651819 10.465681 ab
## 5 5 9.76500 8.358069 11.171931 ab
## 6 6 6.87750 5.470569 8.284431 b
## 7 7 10.59000 9.183069 11.996931 a
## 8 8 10.70250 9.295569 12.109431 a
## 9 9 10.43375 9.026819 11.840681 a
## 10 10 7.86500 6.458069 9.271931 ab
## 11 11 7.32125 5.914319 8.728181 ab
## 12 12 8.04000 6.633069 9.446931 ab
## 13 13 7.51500 6.108069 8.921931 ab
## 14 14 9.85250 8.445569 11.259431 ab
## 15 15 7.65875 6.251819 9.065681 ab
## 16 16 6.90875 5.501819 8.315681 b# Gráfico de segmentos para as estimativas intervalares.
ggplot(data = results_m0,
mapping = aes(x = fit, y = reorder(cult, fit))) +
geom_point() +
geom_errorbarh(mapping = aes(xmin = lwr, xmax = upr),
height = 0) +
geom_text(mapping = aes(label = sprintf("%0.2f %s", fit, cld)),
vjust = 0,
nudge_y = 0.1) +
labs(x = "Produção",
y = "Cultivares")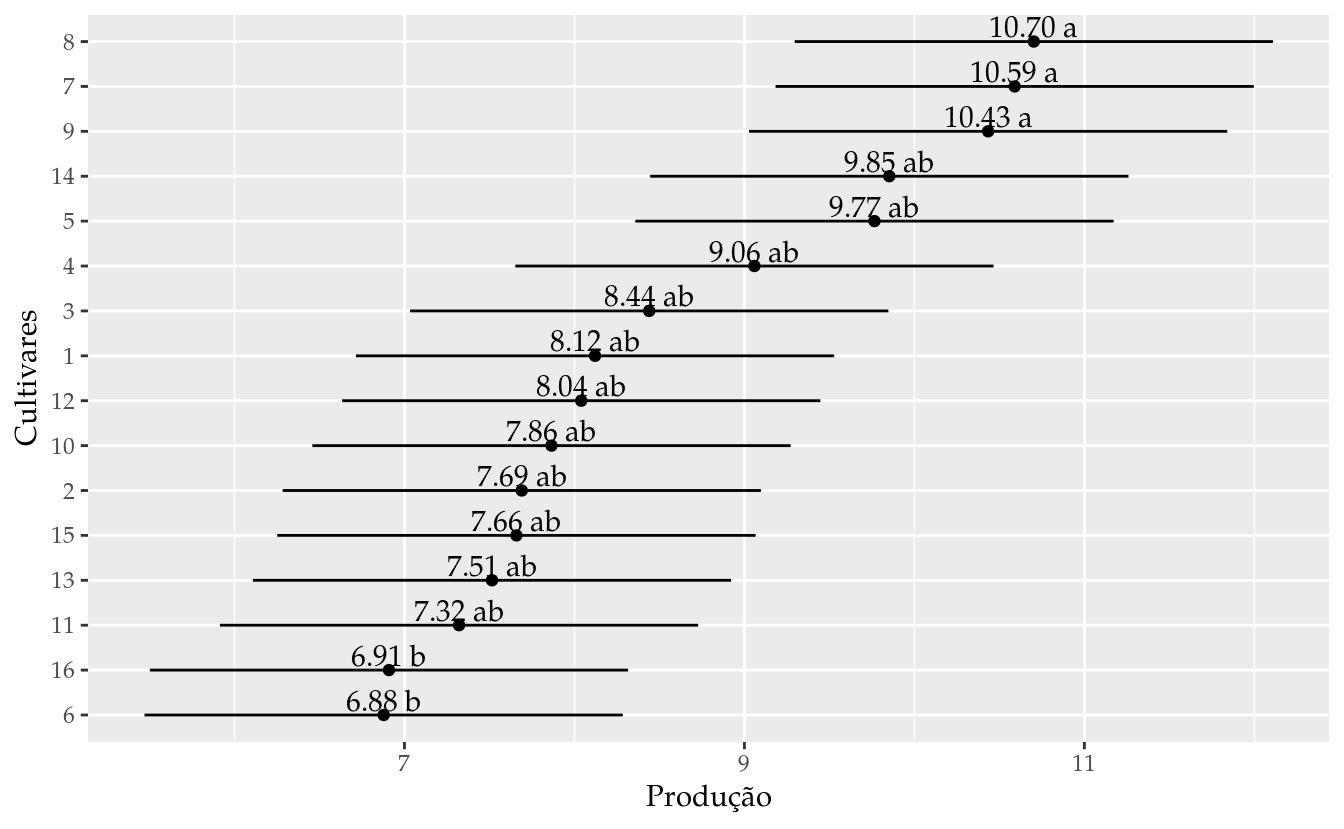
Atenção: as médias ajustadas da Tabela 3 na página 168 em RAMALHO et al. (2005) foram obtidas com o modelo de efeito aleatório para bloco dentro de repetição que será ajustado a seguir. Essas médias podem ser obtidas por meio da análise com recuperação da informação interbloco. Se houver diferença em relação ao modelo de efeitos aleatórios será por causa da diferença na estimativa do componente de variância de bloco. Considerar o modelo de efeitos aleatório é mais interessante por permitir, com facilidade, determinar o erro padrão das médias e realizar comparações múltiplas.
# Cria bloco dentro de repetição (identificador único).
da$cond <- with(da, interaction(rept, bloc))
# Médias ajustadas com a análise da recuperação interbloco.
u <- emmeans_interbloco(trt = "cult", blc = "cond", resp = "prod", data = da)
u## cult emmeans
## 1 1 8.368436
## 2 2 8.042155
## 3 3 8.796025
## 4 4 8.927659
## 5 5 9.855941
## 6 6 7.002302
## 7 7 10.307502
## 8 8 10.647355
## 9 9 10.095623
## 10 10 7.816627
## 11 11 7.274328
## 12 12 8.016781
## 13 13 7.238307
## 14 14 10.180469
## 15 15 7.241291
## 16 16 7.029199# Componentes de variância.
attr(u, "information") %>%
as.data.frame()## n_trat n_block block_variance residual_variance df_residual
## 1 16 20 1.407264 2.842799 45No modelo a seguir, o efeito de bloco dentro de repetições é aleatório. O efeito de repetições foi mantido como fixo, bem como o efeito de cultivares.
# Ajuste do modelo de efeito aleatório de bloc.
mm0 <- lmer(prod ~ rept + (1 | rept:bloc) + cult, data = da)
# Quadro de teste de Wald.
anova(mm0)## Type III Analysis of Variance Table with Satterthwaite's method
## Sum Sq Mean Sq NumDF DenDF F value Pr(>F)
## rept 5.952 1.4879 4 12.986 0.5234 0.720445
## cult 108.862 7.2574 15 51.484 2.5529 0.006463 **
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Estimativas dos parâmetros dos termos de efeito.
summary(mm0)## Linear mixed model fit by REML. t-tests use Satterthwaite's method
## [lmerModLmerTest]
## Formula: prod ~ rept + (1 | rept:bloc) + cult
## Data: da
##
## REML criterion at convergence: 284.2
##
## Scaled residuals:
## Min 1Q Median 3Q Max
## -1.71014 -0.46545 -0.04598 0.48518 2.00915
##
## Random effects:
## Groups Name Variance Std.Dev.
## rept:bloc (Intercept) 1.694 1.301
## Residual 2.843 1.686
## Number of obs: 80, groups: rept:bloc, 20
##
## Fixed effects:
## Estimate Std. Error df t value Pr(>|t|)
## (Intercept) 8.4510 1.1052 39.1512 7.646 2.75e-09 ***
## rept2 -0.2500 1.0965 12.9862 -0.228 0.8232
## rept3 0.1938 1.0965 12.9862 0.177 0.8625
## rept4 0.4938 1.0965 12.9862 0.450 0.6599
## rept5 -0.9875 1.0965 12.9862 -0.901 0.3842
## cult2 -0.3379 1.1505 51.4843 -0.294 0.7701
## cult3 0.4155 1.1505 51.4843 0.361 0.7195
## cult4 0.6012 1.1505 51.4843 0.523 0.6035
## cult5 1.5048 1.1505 51.4843 1.308 0.1967
## cult6 -1.3526 1.1505 51.4843 -1.176 0.2451
## cult7 1.9978 1.1505 51.4843 1.737 0.0885 .
## cult8 2.3125 1.1505 51.4843 2.010 0.0497 *
## cult9 1.7921 1.1505 51.4843 1.558 0.1254
## cult10 -0.5190 1.1505 51.4843 -0.451 0.6538
## cult11 -1.0615 1.1505 51.4843 -0.923 0.3605
## cult12 -0.3217 1.1505 51.4843 -0.280 0.7809
## cult13 -1.0720 1.1505 51.4843 -0.932 0.3558
## cult14 1.8031 1.1505 51.4843 1.567 0.1232
## cult15 -1.0534 1.1505 51.4843 -0.916 0.3641
## cult16 -1.3252 1.1505 51.4843 -1.152 0.2547
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1##
## Correlation matrix not shown by default, as p = 20 > 12.
## Use print(x, correlation=TRUE) or
## vcov(x) if you need it# Médias marginais ajustadas.
emm <- emmeans(mm0, specs = ~cult)
emm## cult emmean SE df lower.CL upper.CL
## 1 8.34 0.87 59.9 6.60 10.08
## 2 8.00 0.87 59.9 6.26 9.74
## 3 8.76 0.87 59.9 7.02 10.50
## 4 8.94 0.87 59.9 7.20 10.68
## 5 9.85 0.87 59.9 8.11 11.59
## 6 6.99 0.87 59.9 5.25 8.73
## 7 10.34 0.87 59.9 8.60 12.08
## 8 10.65 0.87 59.9 8.91 12.39
## 9 10.13 0.87 59.9 8.39 11.87
## 10 7.82 0.87 59.9 6.08 9.56
## 11 7.28 0.87 59.9 5.54 9.02
## 12 8.02 0.87 59.9 6.28 9.76
## 13 7.27 0.87 59.9 5.53 9.01
## 14 10.14 0.87 59.9 8.40 11.88
## 15 7.29 0.87 59.9 5.55 9.03
## 16 7.02 0.87 59.9 5.28 8.76
##
## Results are averaged over the levels of: rept
## Degrees-of-freedom method: kenward-roger
## Confidence level used: 0.95# Extração da matriz de funções lineares.
grid <- attr(emm, "grid")
L <- attr(emm, "linfct")
rownames(L) <- grid[[1]]
# As mesmas comparações múltiplas.
results_mm0 <- wzRfun::apmc(L,
model = mm0,
focus = "cult",
test = "fdr",
level = 0.1)
results_mm0## cult fit lwr upr cld
## 1 1 8.341010 6.925425 9.756595 ad
## 2 2 8.003083 6.587498 9.418667 ad
## 3 3 8.756523 7.340938 10.172108 ad
## 4 4 8.942204 7.526619 10.357789 ad
## 5 5 9.845851 8.430266 11.261436 ac
## 6 6 6.988455 5.572870 8.404040 d
## 7 7 10.338846 8.923261 11.754431 ab
## 8 8 10.653473 9.237888 12.069058 a
## 9 9 10.133139 8.717554 11.548724 ab
## 10 10 7.821994 6.406409 9.237579 bcd
## 11 11 7.279534 5.863949 8.695119 cd
## 12 12 8.019357 6.603772 9.434942 ad
## 13 13 7.269007 5.853422 8.684592 cd
## 14 14 10.144080 8.728495 11.559665 ab
## 15 15 7.287610 5.872025 8.703195 cd
## 16 16 7.015835 5.600250 8.431419 d# Gráfico de segmentos para as estimativas intervalares.
ggplot(data = results_mm0,
mapping = aes(x = fit, y = reorder(cult, fit))) +
geom_point() +
geom_errorbarh(mapping = aes(xmin = lwr, xmax = upr),
height = 0) +
geom_text(mapping = aes(label = sprintf("%0.2f %s", fit, cld)),
vjust = 0,
nudge_y = 0.1) +
labs(x = "Produção",
y = "Cultivares")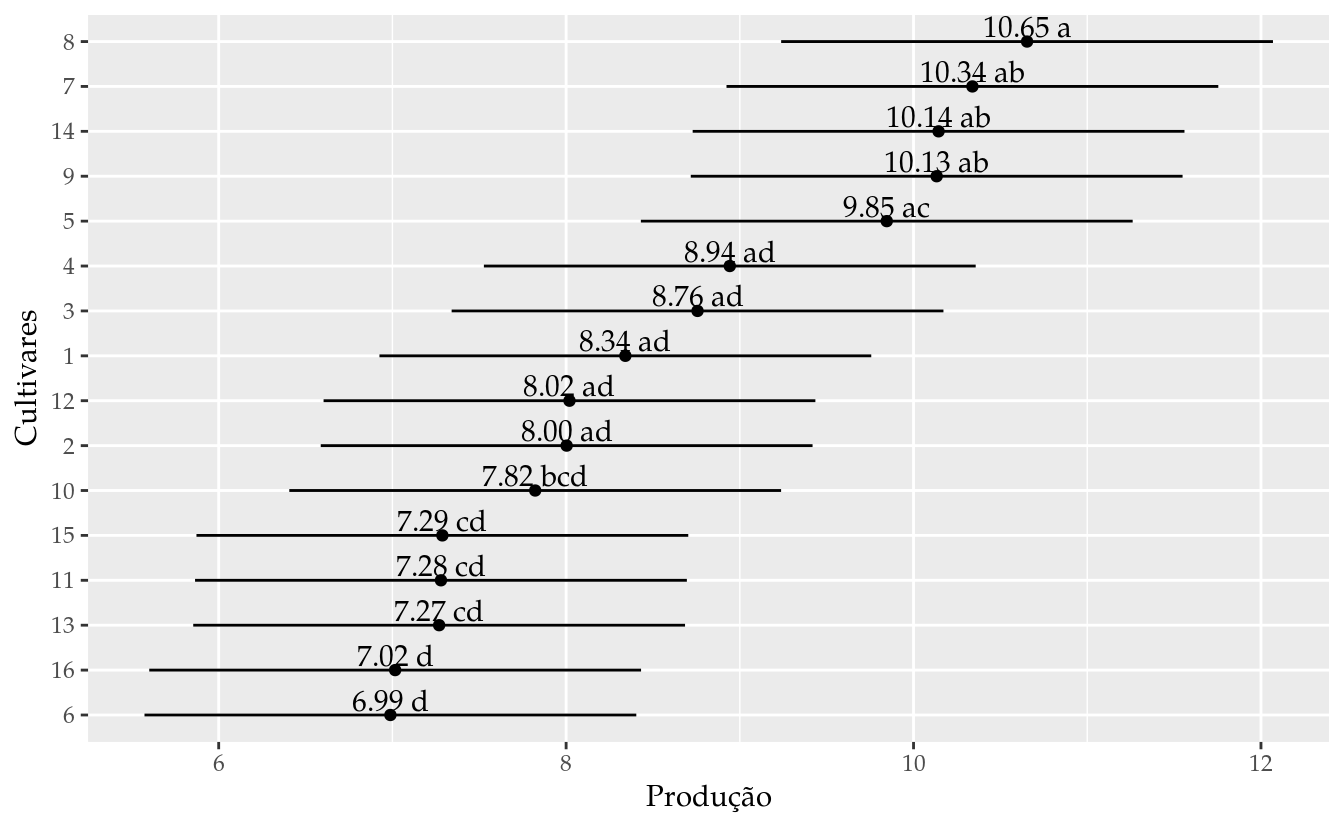
16.2 RamalhoEg11.4
O objeto RamalhoEg11.4 armazena dados de um experimento feito em delineamento de látice quadrado parcialmente balanceado para avaliar a produção (kg/parcela) de 36 cultivares de milho.
No delineamento de látice parcialmente balanceado apenas alguns pares de tratamentos ocorrem juntos nos blocos. Fazer com que todos os pares sejam observados simultaneamente exige um número grande de blocos, o que torna inivável a realização do experimento.
Com isso, vão existir 3 tipos de associação. O primeiro tipo corresponde ao conjunto de tratamentos que ocorrem juntos \(\lambda_1 > 0\) vezes. O segundo tipo corresponde ao conjunto de tratamentos que ocorre \(0 \leq \lambda_2 < \lambda_1\) vezes. O terceiro, caso \(\lambda_2 > 0\), é o grupo de tratamentos que não ocorre junto.
Dessa forma, os contrastes entre tratamentos terão erro padrão de tamanho correspondente a estrutura de associação. O erro padrão do contraste entre primeiros associados será o menor, seguido pelos segundos associados e, por fim, o maior erro padrão será para o contraste entre tratamentos que não ocorreram juntos.
# Dados do experimento.
da <- as_tibble(labestData::RamalhoEg11.4)
da$rept <- gl(2, 36)
str(da)## Classes 'tbl_df', 'tbl' and 'data.frame': 72 obs. of 4 variables:
## $ bloc: Factor w/ 6 levels "1","2","3","4",..: 1 1 1 1 1 1 2 2 2 2 ...
## $ cult: Factor w/ 36 levels "1","2","3","4",..: 1 2 3 4 5 6 7 8 9 10 ...
## $ prod: num 4.39 3.66 2 1.92 4.69 ...
## $ rept: Factor w/ 2 levels "1","2": 1 1 1 1 1 1 1 1 1 1 ...# São 6 blocos de tamanho 6 em cada uma das 2 repetições para acomodar
# 36 cultivares de milho.
with(da, table(cult, bloc, rept)) %>%
addmargins(1:2) %>%
print.table(zero.print = ".")## , , rept = 1
##
## bloc
## cult 1 2 3 4 5 6 Sum
## 1 1 . . . . . 1
## 2 1 . . . . . 1
## 3 1 . . . . . 1
## 4 1 . . . . . 1
## 5 1 . . . . . 1
## 6 1 . . . . . 1
## 7 . 1 . . . . 1
## 8 . 1 . . . . 1
## 9 . 1 . . . . 1
## 10 . 1 . . . . 1
## 11 . 1 . . . . 1
## 12 . 1 . . . . 1
## 13 . . 1 . . . 1
## 14 . . 1 . . . 1
## 15 . . 1 . . . 1
## 16 . . 1 . . . 1
## 17 . . 1 . . . 1
## 18 . . 1 . . . 1
## 19 . . . 1 . . 1
## 20 . . . 1 . . 1
## 21 . . . 1 . . 1
## 22 . . . 1 . . 1
## 23 . . . 1 . . 1
## 24 . . . 1 . . 1
## 25 . . . . 1 . 1
## 26 . . . . 1 . 1
## 27 . . . . 1 . 1
## 28 . . . . 1 . 1
## 29 . . . . 1 . 1
## 30 . . . . 1 . 1
## 31 . . . . . 1 1
## 32 . . . . . 1 1
## 33 . . . . . 1 1
## 34 . . . . . 1 1
## 35 . . . . . 1 1
## 36 . . . . . 1 1
## Sum 6 6 6 6 6 6 36
##
## , , rept = 2
##
## bloc
## cult 1 2 3 4 5 6 Sum
## 1 1 . . . . . 1
## 2 . 1 . . . . 1
## 3 . . 1 . . . 1
## 4 . . . 1 . . 1
## 5 . . . . 1 . 1
## 6 . . . . . 1 1
## 7 1 . . . . . 1
## 8 . 1 . . . . 1
## 9 . . 1 . . . 1
## 10 . . . 1 . . 1
## 11 . . . . 1 . 1
## 12 . . . . . 1 1
## 13 1 . . . . . 1
## 14 . 1 . . . . 1
## 15 . . 1 . . . 1
## 16 . . . 1 . . 1
## 17 . . . . 1 . 1
## 18 . . . . . 1 1
## 19 1 . . . . . 1
## 20 . 1 . . . . 1
## 21 . . 1 . . . 1
## 22 . . . 1 . . 1
## 23 . . . . 1 . 1
## 24 . . . . . 1 1
## 25 1 . . . . . 1
## 26 . 1 . . . . 1
## 27 . . 1 . . . 1
## 28 . . . 1 . . 1
## 29 . . . . 1 . 1
## 30 . . . . . 1 1
## 31 1 . . . . . 1
## 32 . 1 . . . . 1
## 33 . . 1 . . . 1
## 34 . . . 1 . . 1
## 35 . . . . 1 . 1
## 36 . . . . . 1 1
## Sum 6 6 6 6 6 6 36library(igraph)
da$cond <- with(da, interaction(rept, bloc))
edg <- by(data = as.integer(da$cult),
INDICES = da$cond,
FUN = combn,
m = 2) %>%
flatten_int()
ghp <- graph(edg, directed = FALSE)
plot(ghp,
vertex.size = 5,
vertex.label.dist = 1,
# layout = layout_components,
layout = layout_in_circle,
edge.curved = FALSE)
# Quantidade de pares possíveis.
choose(nlevels(da$cult), k = 2)
# Quantas vezes cada par ocorre junto.
by(data = da$cult,
INDICES = da$cond,
FUN = function(x) {
apply(combn(sort(x), m = 2),
MARGIN = 2,
FUN = paste0,
collapse = "_")
}) %>%
flatten_chr() %>%
table() %>%
enframe(name = "par", value = "freq")# CAUTION: quadro de anova não bate com o livro!
# Modelo de efeitos fixos.
m0 <- lm(terms(prod ~ rept/bloc + cult, keep.order = TRUE),
data = da)
# Quado de anova com hipóteses marginais.
Anova(m0)## Anova Table (Type II tests)
##
## Response: prod
## Sum Sq Df F value Pr(>F)
## rept 32.334 1 49.5050 2.247e-07 ***
## rept:bloc 16.611 10 2.5432 0.02841 *
## cult 41.034 35 1.7950 0.06541 .
## Residuals 16.329 25
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Médias marginais ajustadas.
emm <- emmeans(m0, specs = ~cult)## NOTE: A nesting structure was detected in the fitted model:
## bloc %in% reptemm## cult emmean SE df lower.CL upper.CL
## 1 4.98 0.646 25 3.64 6.31
## 2 3.80 0.646 25 2.47 5.13
## 3 3.34 0.646 25 2.01 4.67
## 4 3.04 0.646 25 1.71 4.37
## 5 4.78 0.646 25 3.45 6.12
## 6 6.14 0.646 25 4.81 7.47
## 7 3.84 0.646 25 2.51 5.17
## 8 3.38 0.646 25 2.05 4.71
## 9 4.76 0.646 25 3.43 6.09
## 10 5.08 0.646 25 3.74 6.41
## 11 4.36 0.646 25 3.03 5.70
## 12 4.03 0.646 25 2.70 5.36
## 13 4.89 0.646 25 3.56 6.22
## 14 3.25 0.646 25 1.92 4.58
## 15 5.29 0.646 25 3.96 6.62
## 16 4.80 0.646 25 3.47 6.13
## 17 4.68 0.646 25 3.35 6.01
## 18 3.18 0.646 25 1.85 4.51
## 19 5.19 0.646 25 3.86 6.52
## 20 3.68 0.646 25 2.35 5.01
## 21 5.40 0.646 25 4.07 6.73
## 22 4.53 0.646 25 3.20 5.86
## 23 5.18 0.646 25 3.85 6.51
## 24 2.63 0.646 25 1.30 3.96
## 25 5.35 0.646 25 4.02 6.68
## 26 4.21 0.646 25 2.88 5.54
## 27 4.77 0.646 25 3.44 6.10
## 28 4.69 0.646 25 3.36 6.02
## 29 5.08 0.646 25 3.74 6.41
## 30 5.61 0.646 25 4.28 6.94
## 31 4.65 0.646 25 3.32 5.98
## 32 3.70 0.646 25 2.36 5.03
## 33 4.63 0.646 25 3.30 5.96
## 34 5.24 0.646 25 3.91 6.57
## 35 5.17 0.646 25 3.84 6.50
## 36 5.58 0.646 25 4.24 6.91
##
## Results are averaged over the levels of: bloc, rept
## Confidence level used: 0.95# Extração da matriz de funções lineares.
L <- attr(emm, "linfct")
grid <- attr(emm, "grid")
rownames(L) <- grid[[1]]
# Entenda como são obtidas as médias marginais.
MASS::fractions(t(L[1:4, ])) %>%
unique()## 1 2 3 4
## (Intercept) 1 1 1 1
## rept2 1/2 1/2 1/2 1/2
## rept1:bloc2 1/12 1/12 1/12 1/12
## cult2 0 1 0 0
## cult3 0 0 1 0
## cult4 0 0 0 1
## cult5 0 0 0 0# Contrastes par a par.
ctr <- summary(glht(m0, linfct = all_pairwise(L)),
test = adjusted(type = "fdr"))
# Erros padrões de dois tamanhos conforme estrutura de associados.
v <- c("coefficients", "sigma", "tstat", "pvalues")
ctr$test[v] %>%
as.data.frame() %>%
split(., round(.$sigma, 4)) %>%
map(head, n = 10)## $`0.8729`
## coefficients sigma tstat pvalues
## 1vs2 1.17916667 0.8729308 1.35081343 0.5726127
## 1vs3 1.63625000 0.8729308 1.87443263 0.4317107
## 1vs4 1.93291667 0.8729308 2.21428393 0.3938518
## 1vs5 0.19083333 0.8729308 0.21861221 0.9668481
## 1vs6 -1.16083333 0.8729308 -1.32981139 0.5812251
## 1vs7 1.13250000 0.8729308 1.29735368 0.5970728
## 1vs13 0.08541667 0.8729308 0.09785044 0.9693983
## 1vs19 -0.21625000 0.8729308 -0.24772868 0.9605105
## 1vs25 -0.37083333 0.8729308 -0.42481412 0.9023365
## 1vs31 0.32583333 0.8729308 0.37326364 0.9241553
##
## $`0.9332`
## coefficients sigma tstat pvalues
## 1vs8 1.59916667 0.9332023 1.7136335 0.4711698
## 1vs9 0.21625000 0.9332023 0.2317290 0.9644038
## 1vs10 -0.09958333 0.9332023 -0.1067114 0.9693983
## 1vs11 0.61083333 0.9332023 0.6545562 0.8688576
## 1vs12 0.94416667 0.9332023 1.0117492 0.7153741
## 1vs14 1.72708333 0.9332023 1.8507063 0.4317107
## 1vs15 -0.31583333 0.9332023 -0.3384404 0.9241553
## 1vs16 0.17333333 0.9332023 0.1857404 0.9693983
## 1vs17 0.29875000 0.9332023 0.3201342 0.9323001
## 1vs18 1.79458333 0.9332023 1.9230379 0.4238813# Comparações múltiplas a 10%.
results_m0 <- wzRfun::apmc(X = L,
model = m0,
focus = "cult",
test = "fdr",
level = 0.1)
results_m0## cult fit lwr upr cld
## 1 1 4.975417 3.871992 6.078841 a
## 2 2 3.796250 2.692825 4.899675 a
## 3 3 3.339167 2.235742 4.442591 a
## 4 4 3.042500 1.939075 4.145925 a
## 5 5 4.784583 3.681159 5.888008 a
## 6 6 6.136250 5.032825 7.239675 a
## 7 7 3.842917 2.739492 4.946341 a
## 8 8 3.376250 2.272825 4.479675 a
## 9 9 4.759167 3.655742 5.862591 a
## 10 10 5.075000 3.971575 6.178425 a
## 11 11 4.364583 3.261159 5.468008 a
## 12 12 4.031250 2.927825 5.134675 a
## 13 13 4.890000 3.786575 5.993425 a
## 14 14 3.248333 2.144909 4.351758 a
## 15 15 5.291250 4.187825 6.394675 a
## 16 16 4.802083 3.698659 5.905508 a
## 17 17 4.676667 3.573242 5.780091 a
## 18 18 3.180833 2.077409 4.284258 a
## 19 19 5.191667 4.088242 6.295091 a
## 20 20 3.680000 2.576575 4.783425 a
## 21 21 5.402917 4.299492 6.506341 a
## 22 22 4.533750 3.430325 5.637175 a
## 23 23 5.180833 4.077409 6.284258 a
## 24 24 2.630000 1.526575 3.733425 a
## 25 25 5.346250 4.242825 6.449675 a
## 26 26 4.209583 3.106159 5.313008 a
## 27 27 4.770000 3.666575 5.873425 a
## 28 28 4.690833 3.587409 5.794258 a
## 29 29 5.075417 3.971992 6.178841 a
## 30 30 5.612083 4.508659 6.715508 a
## 31 31 4.649583 3.546159 5.753008 a
## 32 32 3.695417 2.591992 4.798841 a
## 33 33 4.628333 3.524909 5.731758 a
## 34 34 5.236667 4.133242 6.340091 a
## 35 35 5.173750 4.070325 6.277175 a
## 36 36 5.575417 4.471992 6.678841 a# Gráfico de segmentos para as estimativas intervalares.
ggplot(data = results_m0,
mapping = aes(x = fit, y = reorder(cult, fit))) +
geom_point() +
geom_errorbarh(mapping = aes(xmin = lwr, xmax = upr),
height = 0) +
geom_label(mapping = aes(label = sprintf("%0.2f%s", fit, cld)),
label.padding = unit(0.15, "lines"),
fill = "black",
colour = "white",
size = 3,
nudge_x = 0.25,
vjust = 0.5) +
labs(x = "Produção",
y = "Cultivares")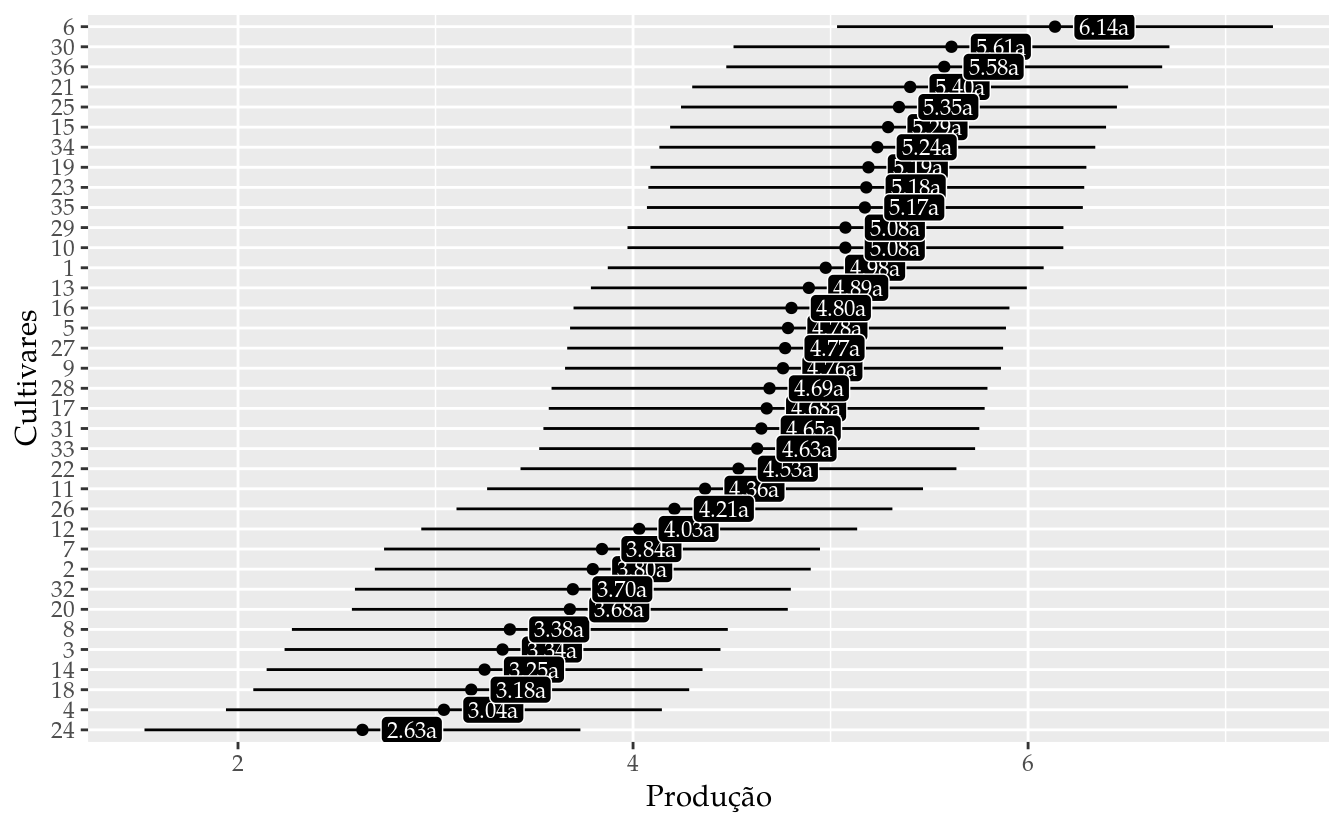
ATTENTION: pode ocorrer de duas médias consecutivas diferirem entre si e elas podem estar dentro do intervalo formado por duas médias que não diferem! As primeiras médias podem ser de tratamentos em primeiro associados e as outras de tratamentos como segundo associados. Acredito que seja um evento bem raro, mas é possível ter algo como exemplificado a seguir.
10.7 abc – 11.2 b – 13.9 c – 14.1 abc
TODO
# Ajuste do modelo de efeito aleatório de bloc.
mm0 <- lmer(prod ~ rept + (1 | rept:bloc) + cult, data = da)
# Quadro de teste de Wald.
anova(mm0)## Type III Analysis of Variance Table with Satterthwaite's method
## Sum Sq Mean Sq NumDF DenDF F value Pr(>F)
## rept 7.913 7.9127 1 6.3562 12.1147 0.01197 *
## cult 39.280 1.1223 35 27.2185 1.7183 0.07414 .
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Estimativas dos parâmetros dos termos de efeito.
summary(mm0)## Linear mixed model fit by REML. t-tests use Satterthwaite's method
## [lmerModLmerTest]
## Formula: prod ~ rept + (1 | rept:bloc) + cult
## Data: da
##
## REML criterion at convergence: 121.6
##
## Scaled residuals:
## Min 1Q Median 3Q Max
## -1.5841 -0.3997 0.0000 0.3997 1.5841
##
## Random effects:
## Groups Name Variance Std.Dev.
## rept:bloc (Intercept) 0.3360 0.5796
## Residual 0.6531 0.8082
## Number of obs: 72, groups: rept:bloc, 12
##
## Fixed effects:
## Estimate Std. Error df t value Pr(>|t|)
## (Intercept) 4.27071 0.66835 34.99991 6.390 2.37e-07 ***
## rept2 1.34028 0.38507 6.35616 3.481 0.0120 *
## cult2 -0.94652 0.84806 27.98407 -1.116 0.2739
## cult3 -1.54630 0.84806 27.98407 -1.823 0.0790 .
## cult4 -1.92391 0.84806 27.98407 -2.269 0.0312 *
## cult5 -0.27210 0.84806 27.98407 -0.321 0.7507
## cult6 1.27355 0.84806 27.98407 1.502 0.1444
## cult7 -0.82776 0.84806 27.98407 -0.976 0.3374
## cult8 -1.06178 0.88615 30.35602 -1.198 0.2401
## cult9 0.17843 0.88615 30.35602 0.201 0.8418
## cult10 0.41333 0.88615 30.35602 0.466 0.6442
## cult11 -0.38736 0.88615 30.35602 -0.437 0.6651
## cult12 -0.52671 0.88615 30.35602 -0.594 0.5567
## cult13 -0.14128 0.84806 27.98407 -0.167 0.8689
## cult14 -1.55030 0.88615 30.35602 -1.749 0.0903 .
## cult15 0.34991 0.88615 30.35602 0.395 0.6957
## cult16 -0.22019 0.88615 30.35602 -0.248 0.8054
## cult17 -0.43588 0.88615 30.35602 -0.492 0.6263
## cult18 -1.73773 0.88615 30.35602 -1.961 0.0591 .
## cult19 0.03980 0.84806 27.98407 0.047 0.9629
## cult20 -1.23922 0.88615 30.35602 -1.398 0.1721
## cult21 0.34099 0.88615 30.35602 0.385 0.7031
## cult22 -0.60911 0.88615 30.35602 -0.687 0.4971
## cult23 -0.05230 0.88615 30.35602 -0.059 0.9533
## cult24 -2.40915 0.88615 30.35602 -2.719 0.0107 *
## cult25 0.20536 0.84806 27.98407 0.242 0.8104
## cult26 -0.69866 0.88615 30.35602 -0.788 0.4366
## cult27 -0.28095 0.88615 30.35602 -0.317 0.7534
## cult28 -0.44105 0.88615 30.35602 -0.498 0.6223
## cult29 -0.14674 0.88615 30.35602 -0.166 0.8696
## cult30 0.58391 0.88615 30.35602 0.659 0.5149
## cult31 -0.38842 0.84806 27.98407 -0.458 0.6505
## cult32 -1.10994 0.88615 30.35602 -1.253 0.2199
## cult33 -0.31972 0.88615 30.35602 -0.361 0.7207
## cult34 0.20768 0.88615 30.35602 0.234 0.8163
## cult35 0.05448 0.88615 30.35602 0.061 0.9514
## cult36 0.65013 0.88615 30.35602 0.734 0.4688
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1##
## Correlation matrix not shown by default, as p = 37 > 12.
## Use print(x, correlation=TRUE) or
## vcov(x) if you need it# Médias marginais ajustadas.
emm <- emmeans(mm0, specs = ~cult)
emm## cult emmean SE df lower.CL upper.CL
## 1 4.94 0.655 33.8 3.61 6.27
## 2 3.99 0.655 33.8 2.66 5.33
## 3 3.39 0.655 33.8 2.06 4.73
## 4 3.02 0.655 33.8 1.68 4.35
## 5 4.67 0.655 33.8 3.34 6.00
## 6 6.21 0.655 33.8 4.88 7.55
## 7 4.11 0.655 33.8 2.78 5.45
## 8 3.88 0.655 33.8 2.55 5.21
## 9 5.12 0.655 33.8 3.79 6.45
## 10 5.35 0.655 33.8 4.02 6.69
## 11 4.55 0.655 33.8 3.22 5.89
## 12 4.41 0.655 33.8 3.08 5.75
## 13 4.80 0.655 33.8 3.47 6.13
## 14 3.39 0.655 33.8 2.06 4.72
## 15 5.29 0.655 33.8 3.96 6.62
## 16 4.72 0.655 33.8 3.39 6.05
## 17 4.50 0.655 33.8 3.17 5.84
## 18 3.20 0.655 33.8 1.87 4.54
## 19 4.98 0.655 33.8 3.65 6.31
## 20 3.70 0.655 33.8 2.37 5.03
## 21 5.28 0.655 33.8 3.95 6.61
## 22 4.33 0.655 33.8 3.00 5.66
## 23 4.89 0.655 33.8 3.56 6.22
## 24 2.53 0.655 33.8 1.20 3.86
## 25 5.15 0.655 33.8 3.81 6.48
## 26 4.24 0.655 33.8 2.91 5.57
## 27 4.66 0.655 33.8 3.33 5.99
## 28 4.50 0.655 33.8 3.17 5.83
## 29 4.79 0.655 33.8 3.46 6.13
## 30 5.52 0.655 33.8 4.19 6.86
## 31 4.55 0.655 33.8 3.22 5.88
## 32 3.83 0.655 33.8 2.50 5.16
## 33 4.62 0.655 33.8 3.29 5.95
## 34 5.15 0.655 33.8 3.82 6.48
## 35 5.00 0.655 33.8 3.66 6.33
## 36 5.59 0.655 33.8 4.26 6.92
##
## Results are averaged over the levels of: rept
## Degrees-of-freedom method: kenward-roger
## Confidence level used: 0.95# Extração da matriz de funções lineares.
grid <- attr(emm, "grid")
L <- attr(emm, "linfct")
rownames(L) <- grid[[1]]
# Contrastes par a par.
ctr <- summary(glht(mm0, linfct = all_pairwise(L)),
test = adjusted(type = "fdr"))
# Erros padrões de dois tamanhos conforme estrutura de associados.
v <- c("coefficients", "sigma", "tstat", "pvalues")
ctr$test[v] %>%
as.data.frame() %>%
split(., round(.$sigma, 4)) %>%
map(head, n = 10)## $`0.8481`
## coefficients sigma tstat pvalues
## 1vs2 0.94651874 0.8480587 1.11610050 0.7148448
## 1vs3 1.54630373 0.8480587 1.82334516 0.4290162
## 1vs4 1.92390566 0.8480587 2.26859963 0.2883864
## 1vs5 0.27209627 0.8480587 0.32084603 0.9353039
## 1vs6 -1.27355289 0.8480587 -1.50172729 0.5435869
## 1vs7 0.82776399 0.8480587 0.97606921 0.7705449
## 1vs13 0.14128494 0.8480587 0.16659806 0.9554397
## 1vs19 -0.03979802 0.8480587 -0.04692838 0.9790704
## 1vs25 -0.20535840 0.8480587 -0.24215116 0.9481991
## 1vs31 0.38841890 0.8480587 0.45800945 0.9102064
##
## $`0.8861`
## coefficients sigma tstat pvalues
## 1vs8 1.0617827 0.8861478 1.1982005 0.6732803
## 1vs9 -0.1784323 0.8861478 -0.2013573 0.9481991
## 1vs10 -0.4133304 0.8861478 -0.4664350 0.9100631
## 1vs11 0.3873603 0.8861478 0.4371283 0.9108261
## 1vs12 0.5267111 0.8861478 0.5943829 0.8850169
## 1vs14 1.5503037 0.8861478 1.7494866 0.4520915
## 1vs15 -0.3499113 0.8861478 -0.3948679 0.9239990
## 1vs16 0.2201906 0.8861478 0.2484807 0.9481991
## 1vs17 0.4358812 0.8861478 0.4918832 0.8978609
## 1vs18 1.7377320 0.8861478 1.9609958 0.3879518# As mesmas comparações múltiplas.
results_mm0 <- wzRfun::apmc(L,
model = mm0,
focus = "cult",
test = "fdr",
level = 0.1)
results_mm0## cult fit lwr upr cld
## 1 1 4.940847 3.888118 5.993577 ac
## 2 2 3.994328 2.941599 5.047058 ac
## 3 3 3.394543 2.341814 4.447273 bc
## 4 4 3.016941 1.964212 4.069671 bc
## 5 5 4.668751 3.616022 5.721480 ac
## 6 6 6.214400 5.161671 7.267129 a
## 7 7 4.113083 3.060354 5.165813 ac
## 8 8 3.879064 2.826335 4.931794 ac
## 9 9 5.119279 4.066550 6.172009 ac
## 10 10 5.354178 4.301448 6.406907 ac
## 11 11 4.553487 3.500758 5.606216 ac
## 12 12 4.414136 3.361407 5.466865 ac
## 13 13 4.799562 3.746833 5.852292 ac
## 14 14 3.390543 2.337814 4.443273 ac
## 15 15 5.290758 4.238029 6.343488 ac
## 16 16 4.720657 3.667927 5.773386 ac
## 17 17 4.504966 3.452237 5.557695 ac
## 18 18 3.203115 2.150386 4.255844 bc
## 19 19 4.980645 3.927916 6.033375 ac
## 20 20 3.701626 2.648897 4.754356 ac
## 21 21 5.281841 4.229112 6.334571 ac
## 22 22 4.331740 3.279010 5.384469 ac
## 23 23 4.888549 3.835820 5.941278 ac
## 24 24 2.531698 1.478969 3.584427 c
## 25 25 5.146206 4.093476 6.198935 ac
## 26 26 4.242187 3.189457 5.294916 ac
## 27 27 4.659902 3.607172 5.712631 ac
## 28 28 4.499800 3.447071 5.552529 ac
## 29 29 4.794109 3.741380 5.846839 ac
## 30 30 5.524758 4.472029 6.577488 ab
## 31 31 4.552428 3.499699 5.605158 ac
## 32 32 3.830910 2.778180 4.883639 ac
## 33 33 4.621125 3.568395 5.673854 ac
## 34 34 5.148523 4.095793 6.201252 ac
## 35 35 4.995332 3.942603 6.048061 ac
## 36 36 5.590981 4.538252 6.643711 ab# Gráfico de segmentos para as estimativas intervalares.
ggplot(data = results_mm0,
mapping = aes(x = fit, y = reorder(cult, fit))) +
geom_point() +
geom_errorbarh(mapping = aes(xmin = lwr, xmax = upr),
height = 0) +
geom_label(mapping = aes(label = sprintf("%0.2f%s", fit, cld)),
label.padding = unit(0.15, "lines"),
fill = "black",
colour = "white",
size = 3,
nudge_x = 0.25,
vjust = 0.5) +
labs(x = "Produção",
y = "Cultivares")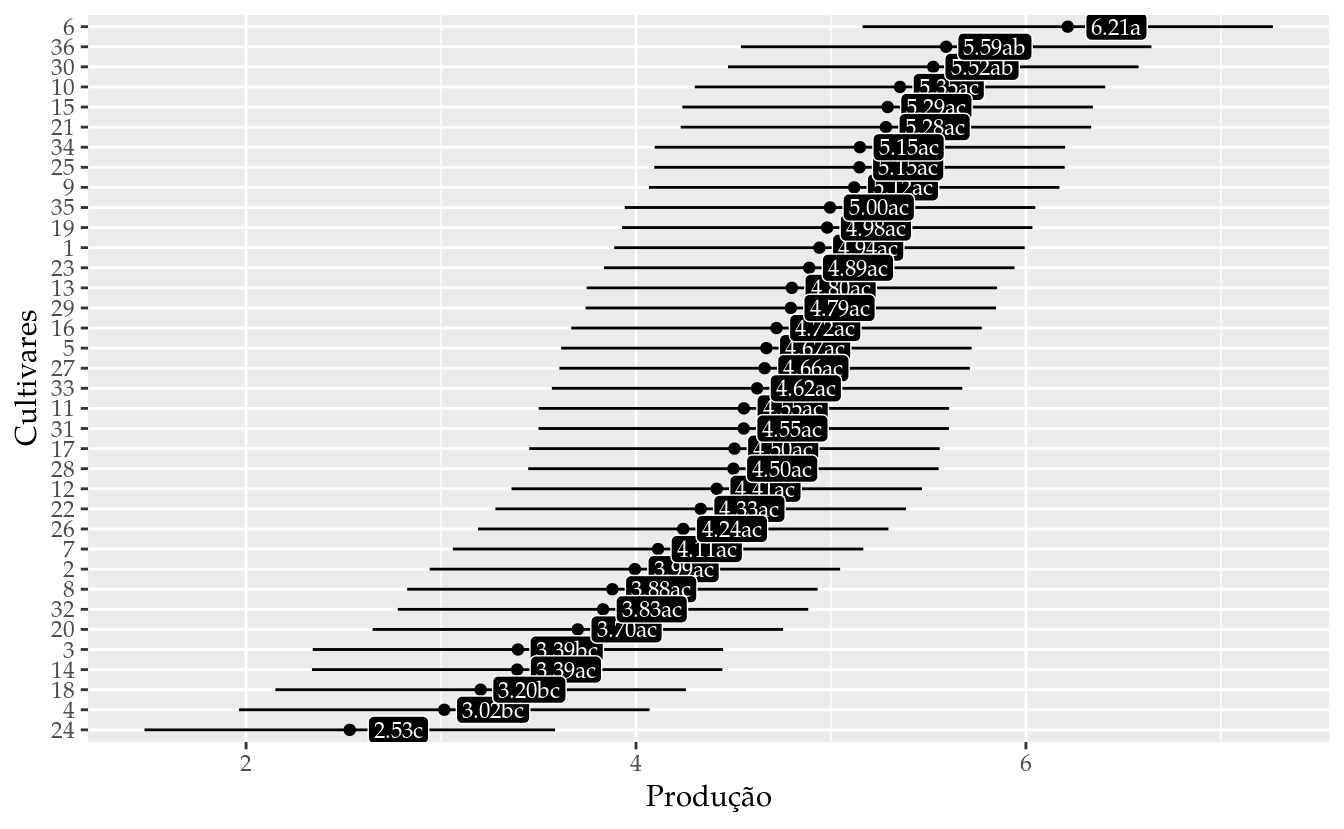
ATTENTION: A tabela 7 da página 175 não foi incluída na obra.
16.3 ZimmermannTb7.1
Os dados em ZimmermannTb7.1 correspondem a produção de arroz em terras altas (kg/ha) de um experimento em delineamento látice reticulado quadrado \(5^2\) com 3 repetições. No experimento foram estudados \(t = 5^2 = 25\) tratamentos em \(b = 15\) blocos de tamanho \(k = 5\) agrupados em \(r = 3\) repetições, perfazendo assim o total de \(tr = bk = 75\) unidades experimentais.
da <- labestData::ZimmermannTb7.1
str(da)## 'data.frame': 75 obs. of 4 variables:
## $ rept: Factor w/ 3 levels "1","2","3": 1 1 1 1 1 1 1 1 1 1 ...
## $ bloc: Factor w/ 5 levels "1","2","3","4",..: 1 1 1 1 1 2 2 2 2 2 ...
## $ cult: Factor w/ 25 levels "1","2","3","4",..: 17 20 16 19 18 10 9 7 8 6 ...
## $ prod: num 1875 2125 2250 1562 844 ...# TODO FIXME: arrumar dado no labestData.
aggregate(prod ~ bloc + rept, data = da, FUN = sum)## bloc rept prod
## 1 1 1 8656.25
## 2 2 1 11500.00
## 3 3 1 8500.00
## 4 4 1 11375.00
## 5 5 1 9250.00
## 6 1 2 9906.25
## 7 2 2 11843.75
## 8 3 2 13375.00
## 9 4 2 15843.75
## 10 5 2 11937.50
## 11 1 3 10718.75
## 12 2 3 14375.00
## 13 3 3 13311.25
## 14 4 3 14031.25
## 15 5 3 8562.50i <- which(with(da, rept == "3" & bloc == "3" & cult == "4"))
da$prod[i] <- 1906.25
# São 6 blocos de tamanho 6 em cada uma das 2 repetições para acomodar
# 36 cultivares de milho.
with(da, table(cult, bloc, rept)) %>%
addmargins(1:2) %>%
print.table(zero.print = ".")## , , rept = 1
##
## bloc
## cult 1 2 3 4 5 Sum
## 1 . . 1 . . 1
## 2 . . 1 . . 1
## 3 . . 1 . . 1
## 4 . . 1 . . 1
## 5 . . 1 . . 1
## 6 . 1 . . . 1
## 7 . 1 . . . 1
## 8 . 1 . . . 1
## 9 . 1 . . . 1
## 10 . 1 . . . 1
## 11 . . . 1 . 1
## 12 . . . 1 . 1
## 13 . . . 1 . 1
## 14 . . . 1 . 1
## 15 . . . 1 . 1
## 16 1 . . . . 1
## 17 1 . . . . 1
## 18 1 . . . . 1
## 19 1 . . . . 1
## 20 1 . . . . 1
## 21 . . . . 1 1
## 22 . . . . 1 1
## 23 . . . . 1 1
## 24 . . . . 1 1
## 25 . . . . 1 1
## Sum 5 5 5 5 5 25
##
## , , rept = 2
##
## bloc
## cult 1 2 3 4 5 Sum
## 1 . . 1 . . 1
## 2 . . . 1 . 1
## 3 . . . . 1 1
## 4 1 . . . . 1
## 5 . 1 . . . 1
## 6 . . 1 . . 1
## 7 . . . 1 . 1
## 8 . . . . 1 1
## 9 1 . . . . 1
## 10 . 1 . . . 1
## 11 . . 1 . . 1
## 12 . . . 1 . 1
## 13 . . . . 1 1
## 14 1 . . . . 1
## 15 . 1 . . . 1
## 16 . . 1 . . 1
## 17 . . . 1 . 1
## 18 . . . . 1 1
## 19 1 . . . . 1
## 20 . 1 . . . 1
## 21 . . 1 . . 1
## 22 . . . 1 . 1
## 23 . . . . 1 1
## 24 1 . . . . 1
## 25 . 1 . . . 1
## Sum 5 5 5 5 5 25
##
## , , rept = 3
##
## bloc
## cult 1 2 3 4 5 Sum
## 1 . . . 1 . 1
## 2 . . . . 1 1
## 3 1 . . . . 1
## 4 . . 1 . . 1
## 5 . 1 . . . 1
## 6 . 1 . . . 1
## 7 . . . 1 . 1
## 8 . . . . 1 1
## 9 1 . . . . 1
## 10 . . 1 . . 1
## 11 . . 1 . . 1
## 12 . 1 . . . 1
## 13 . . . 1 . 1
## 14 . . . . 1 1
## 15 1 . . . . 1
## 16 1 . . . . 1
## 17 . . 1 . . 1
## 18 . 1 . . . 1
## 19 . . . 1 . 1
## 20 . . . . 1 1
## 21 . . . . 1 1
## 22 1 . . . . 1
## 23 . . 1 . . 1
## 24 . 1 . . . 1
## 25 . . . 1 . 1
## Sum 5 5 5 5 5 25library(igraph)
da$cond <- with(da, interaction(rept, bloc))
edg <- by(data = as.integer(da$cult),
INDICES = da$cond,
FUN = combn,
m = 2) %>%
flatten_int()
ghp <- graph(edg, directed = FALSE)
plot(ghp,
vertex.size = 5,
vertex.label.dist = 1,
# layout = layout_components,
layout = layout_in_circle,
edge.curved = FALSE)
# Quantidade de pares possíveis.
choose(nlevels(da$cult), k = 2)
# Todos os possíveis pares.
pares <- apply(combn(sort(as.character(levels(da$cult))), m = 2),
MARGIN = 2,
FUN = paste0,
collapse = "_") %>%
tibble(name = ., value = 0)
# Quantas vezes cada par ocorre junto.
by(data = da$cult,
INDICES = da$cond,
FUN = function(x) {
apply(combn(sort(x), m = 2),
MARGIN = 2,
FUN = paste0,
collapse = "_")
}) %>%
flatten_chr() %>%
table() %>%
enframe() %>%
bind_rows(pares) %>%
distinct(name, .keep_all = TRUE) %>%
split(x = .$name, f = .$value)# Modelo de efeitos fixos.
m0 <- lm(terms(prod ~ rept/bloc + cult, keep.order = TRUE),
data = da)
# Quado de anova com hipóteses marginais.
Anova(m0)## Anova Table (Type II tests)
##
## Response: prod
## Sum Sq Df F value Pr(>F)
## rept 4346563 2 14.8998 1.929e-05 ***
## rept:bloc 6003398 12 3.4299 0.0020356 **
## cult 12026003 24 3.4354 0.0004157 ***
## Residuals 5250951 36
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Médias marginais ajustadas.
emm <- emmeans(m0, specs = ~cult)## NOTE: A nesting structure was detected in the fitted model:
## bloc %in% reptemm## cult emmean SE df lower.CL upper.CL
## 1 2539 246 36 2041 3037
## 2 2075 246 36 1577 2573
## 3 1849 246 36 1351 2347
## 4 1794 246 36 1296 2292
## 5 2462 246 36 1965 2960
## 6 2415 246 36 1917 2913
## 7 3307 246 36 2809 3805
## 8 2786 246 36 2288 3284
## 9 2178 246 36 1680 2676
## 10 1767 246 36 1269 2265
## 11 2225 246 36 1727 2723
## 12 2138 246 36 1640 2635
## 13 2410 246 36 1912 2908
## 14 2434 246 36 1936 2932
## 15 2886 246 36 2388 3384
## 16 3042 246 36 2544 3540
## 17 2551 246 36 2053 3049
## 18 1562 246 36 1065 2060
## 19 2130 246 36 1632 2628
## 20 2496 246 36 1998 2994
## 21 1596 246 36 1098 2094
## 22 2167 246 36 1669 2665
## 23 2129 246 36 1631 2627
## 24 2623 246 36 2125 3121
## 25 2157 246 36 1659 2655
##
## Results are averaged over the levels of: bloc, rept
## Confidence level used: 0.95# Extração da matriz de funções lineares.
L <- attr(emm, "linfct")
grid <- attr(emm, "grid")
rownames(L) <- grid[[1]]
# Entenda como são obtidas as médias marginais.
MASS::fractions(t(L[1:4, ])) %>%
unique()## 1 2 3 4
## (Intercept) 1 1 1 1
## rept2 1/3 1/3 1/3 1/3
## rept1:bloc2 1/15 1/15 1/15 1/15
## cult2 0 1 0 0
## cult3 0 0 1 0
## cult4 0 0 0 1
## cult5 0 0 0 0# Contrastes par a par.
ctr <- summary(glht(m0, linfct = all_pairwise(L)),
test = adjusted(type = "fdr"))
# Erros padrões de dois tamanhos conforme estrutura de associados.
v <- c("coefficients", "sigma", "tstat", "pvalues")
ctr$test[v] %>%
as.data.frame() %>%
split(., round(.$sigma, 4)) %>%
map(head, n = 10)## $`341.5959`
## coefficients sigma tstat pvalues
## 1vs2 463.54167 341.5959 1.3569882 0.4164385
## 1vs3 689.58333 341.5959 2.0187105 0.1987545
## 1vs4 744.79167 341.5959 2.1803294 0.1614960
## 1vs5 76.04167 341.5959 0.2226071 0.9376131
## 1vs6 123.95833 341.5959 0.3628800 0.8695361
## 1vs7 -768.75000 341.5959 -2.2504658 0.1481772
## 1vs11 313.54167 341.5959 0.9178729 0.5759909
## 1vs13 128.12500 341.5959 0.3750776 0.8621095
## 1vs16 -503.12500 341.5959 -1.4728658 0.3675671
## 1vs19 408.33333 341.5959 1.1953694 0.4960509
##
## $`355.5443`
## coefficients sigma tstat pvalues
## 1vs8 -247.91667 355.5443 -0.69728764 0.65638622
## 1vs9 360.41667 355.5443 1.01370387 0.55055898
## 1vs10 771.87500 355.5443 2.17096697 0.16149598
## 1vs12 401.04167 355.5443 1.12796529 0.51874673
## 1vs14 104.16667 355.5443 0.29297800 0.92076611
## 1vs15 -347.91667 355.5443 -0.97854651 0.56348526
## 1vs17 -12.50000 355.5443 -0.03515736 0.98992520
## 1vs18 976.04167 355.5443 2.74520385 0.08539424
## 1vs20 42.70833 355.5443 0.12012098 0.97306706
## 1vs22 371.87500 355.5443 1.04593145 0.54268083# Comparações múltiplas a 10%.
results_m0 <- wzRfun::apmc(X = L,
model = m0,
focus = "cult",
test = "bonferroni",
level = 0.05)
results_m0## cult fit lwr upr cld
## 1 1 2538.542 2040.568 3036.515 bc
## 2 2 2075.000 1577.027 2572.973 bc
## 3 3 1848.958 1350.985 2346.932 bc
## 4 4 1793.750 1295.777 2291.723 ac
## 5 5 2462.500 1964.527 2960.473 bc
## 6 6 2414.583 1916.610 2912.557 bc
## 7 7 3307.292 2809.318 3805.265 b
## 8 8 2786.458 2288.485 3284.432 bc
## 9 9 2178.125 1680.152 2676.098 bc
## 10 10 1766.667 1268.693 2264.640 ac
## 11 11 2225.000 1727.027 2722.973 bc
## 12 12 2137.500 1639.527 2635.473 bc
## 13 13 2410.417 1912.443 2908.390 bc
## 14 14 2434.375 1936.402 2932.348 bc
## 15 15 2886.458 2388.485 3384.432 bc
## 16 16 3041.667 2543.693 3539.640 ab
## 17 17 2551.042 2053.068 3049.015 bc
## 18 18 1562.500 1064.527 2060.473 c
## 19 19 2130.208 1632.235 2628.182 bc
## 20 20 2495.833 1997.860 2993.807 bc
## 21 21 1595.833 1097.860 2093.807 c
## 22 22 2166.667 1668.693 2664.640 bc
## 23 23 2129.167 1631.193 2627.140 bc
## 24 24 2622.917 2124.943 3120.890 bc
## 25 25 2157.292 1659.318 2655.265 bc# Gráfico de segmentos para as estimativas intervalares.
ggplot(data = results_m0,
mapping = aes(x = fit, y = reorder(cult, fit))) +
geom_point() +
geom_errorbarh(mapping = aes(xmin = lwr, xmax = upr),
height = 0) +
geom_label(mapping = aes(label = sprintf("%0.0f%s", fit, cld)),
label.padding = unit(0.15, "lines"),
fill = "black",
colour = "white",
size = 3,
nudge_x = 75,
vjust = 0.5) +
labs(x = "Produção",
y = "Cultivares")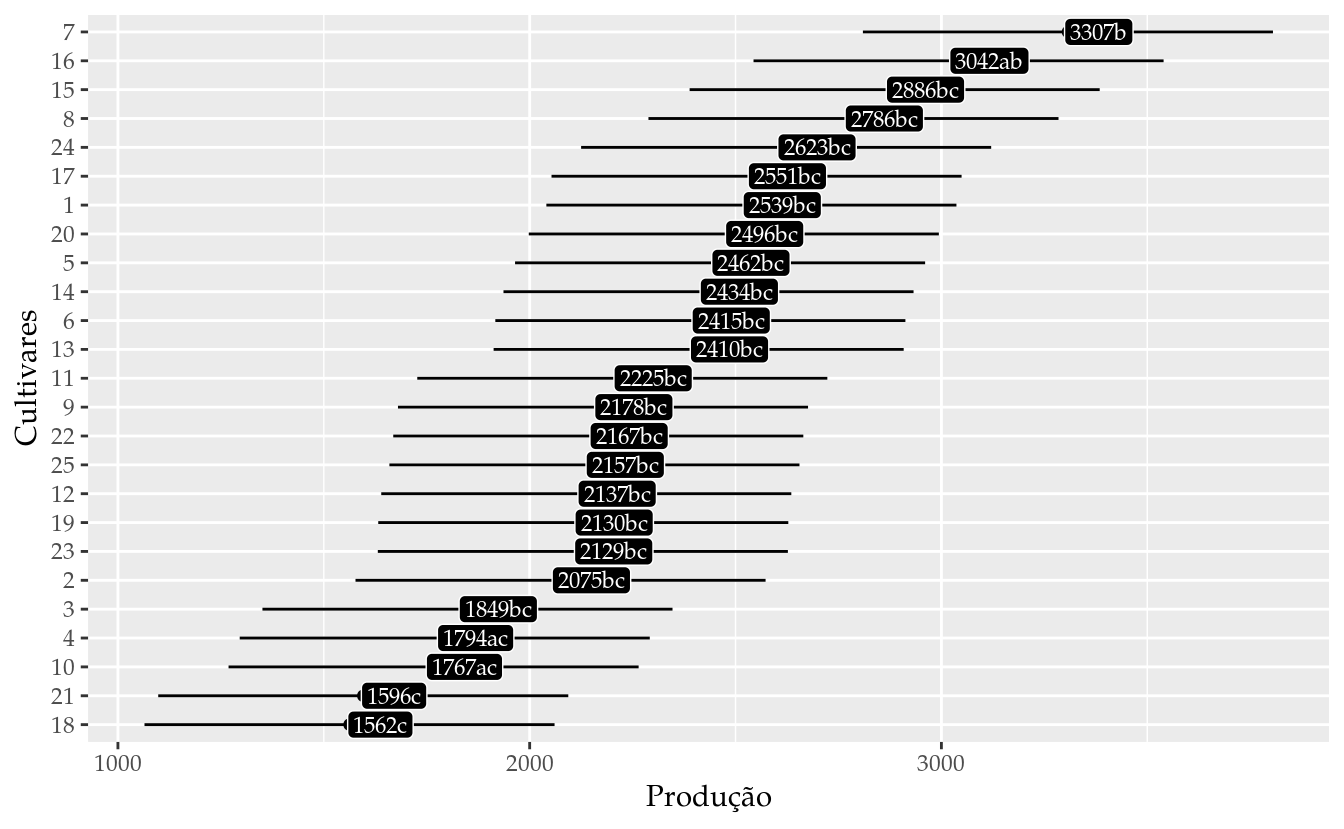
# Ajuste do modelo de efeito aleatório de bloc.
mm0 <- lmer(prod ~ rept + (1 | rept:bloc) + cult, data = da)
# Quadro de teste de Wald.
anova(mm0)## Type III Analysis of Variance Table with Satterthwaite's method
## Sum Sq Mean Sq NumDF DenDF F value Pr(>F)
## rept 935780 467890 2 9.750 3.2078 0.0850170 .
## cult 12545810 522742 24 39.345 3.5839 0.0001915 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Estimativas dos parâmetros dos termos de efeito.
summary(mm0)## Linear mixed model fit by REML. t-tests use Satterthwaite's method
## [lmerModLmerTest]
## Formula: prod ~ rept + (1 | rept:bloc) + cult
## Data: da
##
## REML criterion at convergence: 754.6
##
## Scaled residuals:
## Min 1Q Median 3Q Max
## -2.08640 -0.51285 0.04899 0.45165 1.78751
##
## Random effects:
## Groups Name Variance Std.Dev.
## rept:bloc (Intercept) 106327 326.1
## Residual 145860 381.9
## Number of obs: 75, groups: rept:bloc, 15
##
## Fixed effects:
## Estimate Std. Error df t value Pr(>|t|)
## (Intercept) 2217.14 286.43 42.40 7.741 1.21e-09 ***
## rept2 545.00 232.81 9.75 2.341 0.04189 *
## rept3 467.50 232.81 9.75 2.008 0.07313 .
## cult2 -507.58 333.19 39.63 -1.523 0.13560
## cult3 -752.75 333.19 39.63 -2.259 0.02944 *
## cult4 -773.64 333.19 39.63 -2.322 0.02546 *
## cult5 -72.09 333.19 39.63 -0.216 0.82981
## cult6 -66.56 333.19 39.63 -0.200 0.84269
## cult7 833.13 333.19 39.63 2.500 0.01664 *
## cult8 181.71 343.38 41.07 0.529 0.59952
## cult9 -446.67 343.38 41.07 -1.301 0.20057
## cult10 -750.31 343.38 41.07 -2.185 0.03464 *
## cult11 -258.57 333.19 39.63 -0.776 0.44233
## cult12 -299.30 343.38 41.07 -0.872 0.38847
## cult13 -105.95 333.19 39.63 -0.318 0.75216
## cult14 -213.50 343.38 41.07 -0.622 0.53753
## cult15 292.04 343.38 41.07 0.850 0.39999
## cult16 429.33 333.19 39.63 1.289 0.20504
## cult17 60.48 343.38 41.07 0.176 0.86104
## cult18 -967.84 343.38 41.07 -2.819 0.00739 **
## cult19 -480.61 333.19 39.63 -1.442 0.15703
## cult20 -173.00 343.38 41.07 -0.504 0.61709
## cult21 -1011.04 333.19 39.63 -3.034 0.00424 **
## cult22 -372.79 343.38 41.07 -1.086 0.28396
## cult23 -375.06 343.38 41.07 -1.092 0.28108
## cult24 78.00 343.38 41.07 0.227 0.82143
## cult25 -394.61 333.19 39.63 -1.184 0.24333
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1##
## Correlation matrix not shown by default, as p = 27 > 12.
## Use print(x, correlation=TRUE) or
## vcov(x) if you need it# Médias marginais ajustadas.
emm <- emmeans(mm0, specs = ~cult)
emm## cult emmean SE df lower.CL upper.CL
## 1 2555 256 47.6 2040 3069
## 2 2047 256 47.6 1532 2562
## 3 1802 256 47.6 1287 2317
## 4 1781 256 47.6 1266 2296
## 5 2483 256 47.6 1968 2997
## 6 2488 256 47.6 1973 3003
## 7 3388 256 47.6 2873 3902
## 8 2736 256 47.6 2222 3251
## 9 2108 256 47.6 1593 2623
## 10 1804 256 47.6 1290 2319
## 11 2296 256 47.6 1781 2811
## 12 2255 256 47.6 1741 2770
## 13 2449 256 47.6 1934 2963
## 14 2341 256 47.6 1826 2856
## 15 2847 256 47.6 2332 3361
## 16 2984 256 47.6 2469 3499
## 17 2615 256 47.6 2100 3130
## 18 1587 256 47.6 1072 2101
## 19 2074 256 47.6 1559 2589
## 20 2382 256 47.6 1867 2896
## 21 1544 256 47.6 1029 2058
## 22 2182 256 47.6 1667 2696
## 23 2180 256 47.6 1665 2694
## 24 2633 256 47.6 2118 3147
## 25 2160 256 47.6 1645 2675
##
## Results are averaged over the levels of: rept
## Degrees-of-freedom method: kenward-roger
## Confidence level used: 0.95# Extração da matriz de funções lineares.
grid <- attr(emm, "grid")
L <- attr(emm, "linfct")
rownames(L) <- grid[[1]]
# Contrastes par a par.
ctr <- summary(glht(mm0, linfct = all_pairwise(L)),
test = adjusted(type = "fdr"))
# Erros padrões de dois tamanhos conforme estrutura de associados.
v <- c("coefficients", "sigma", "tstat", "pvalues")
ctr$test[v] %>%
as.data.frame() %>%
split(., round(.$sigma, 4)) %>%
map(head, n = 10)## $`333.193`
## coefficients sigma tstat pvalues
## 1vs2 507.57836 333.193 1.5233765 0.31259352
## 1vs3 752.75321 333.193 2.2592108 0.10874220
## 1vs4 773.64329 333.193 2.3219075 0.09562056
## 1vs5 72.09355 333.193 0.2163717 0.89427841
## 1vs6 66.55878 333.193 0.1997605 0.90178707
## 1vs7 -833.13468 333.193 -2.5004568 0.07155765
## 1vs11 258.57173 333.193 0.7760419 0.63438296
## 1vs13 105.95480 333.193 0.3179983 0.85050862
## 1vs16 -429.32558 333.193 -1.2885192 0.40221438
## 1vs19 480.61425 333.193 1.4424501 0.34112996
##
## $`343.3751`
## coefficients sigma tstat pvalues
## 1vs8 -181.70978 343.3751 -0.5291873 0.74735016
## 1vs9 446.66784 343.3751 1.3008159 0.39755658
## 1vs10 750.31221 343.3751 2.1851093 0.12203170
## 1vs12 299.30172 343.3751 0.8716464 0.58984813
## 1vs14 213.49914 343.3751 0.6217665 0.69968836
## 1vs15 -292.03562 343.3751 -0.8504856 0.59856841
## 1vs17 -60.48481 343.3751 -0.1761479 0.91184921
## 1vs18 967.84173 343.3751 2.8186133 0.04136174
## 1vs20 172.99620 343.3751 0.5038111 0.76480598
## 1vs22 372.78610 343.3751 1.0856526 0.48707497# As mesmas comparações múltiplas.
results_mm0 <- wzRfun::apmc(L,
model = mm0,
focus = "cult",
test = "bonferroni",
level = 0.05)
results_mm0## cult fit lwr upr cld
## 1 1 2554.638 2058.898 3050.377 ad
## 2 2 2047.059 1551.320 2542.799 bcd
## 3 3 1801.885 1306.145 2297.624 bcd
## 4 4 1780.995 1285.255 2276.734 bcd
## 5 5 2482.544 1986.805 2978.284 ad
## 6 6 2488.079 1992.340 2983.818 ad
## 7 7 3387.773 2892.033 3883.512 a
## 8 8 2736.348 2240.608 3232.087 ad
## 9 9 2107.970 1612.231 2603.709 bcd
## 10 10 1804.326 1308.586 2300.065 bcd
## 11 11 2296.066 1800.327 2791.806 ad
## 12 12 2255.336 1759.597 2751.076 ad
## 13 13 2448.683 1952.944 2944.422 ad
## 14 14 2341.139 1845.399 2836.878 ad
## 15 15 2846.673 2350.934 3342.413 ac
## 16 16 2983.963 2488.224 3479.703 ab
## 17 17 2615.123 2119.383 3110.862 ad
## 18 18 1586.796 1091.057 2082.536 cd
## 19 19 2074.024 1578.284 2569.763 bcd
## 20 20 2381.642 1885.902 2877.381 ad
## 21 21 1543.597 1047.857 2039.336 d
## 22 22 2181.852 1686.112 2677.591 ad
## 23 23 2179.581 1683.842 2675.321 ad
## 24 24 2632.635 2136.896 3128.375 ad
## 25 25 2160.025 1664.286 2655.764 ad# Gráfico de segmentos para as estimativas intervalares.
ggplot(data = results_mm0,
mapping = aes(x = fit, y = reorder(cult, fit))) +
geom_point() +
geom_errorbarh(mapping = aes(xmin = lwr, xmax = upr),
height = 0) +
geom_label(mapping = aes(label = sprintf("%0.0f%s", fit, cld)),
label.padding = unit(0.15, "lines"),
fill = "black",
colour = "white",
size = 3,
nudge_x = 75,
vjust = 0.5) +
labs(x = "Produção",
y = "Cultivares")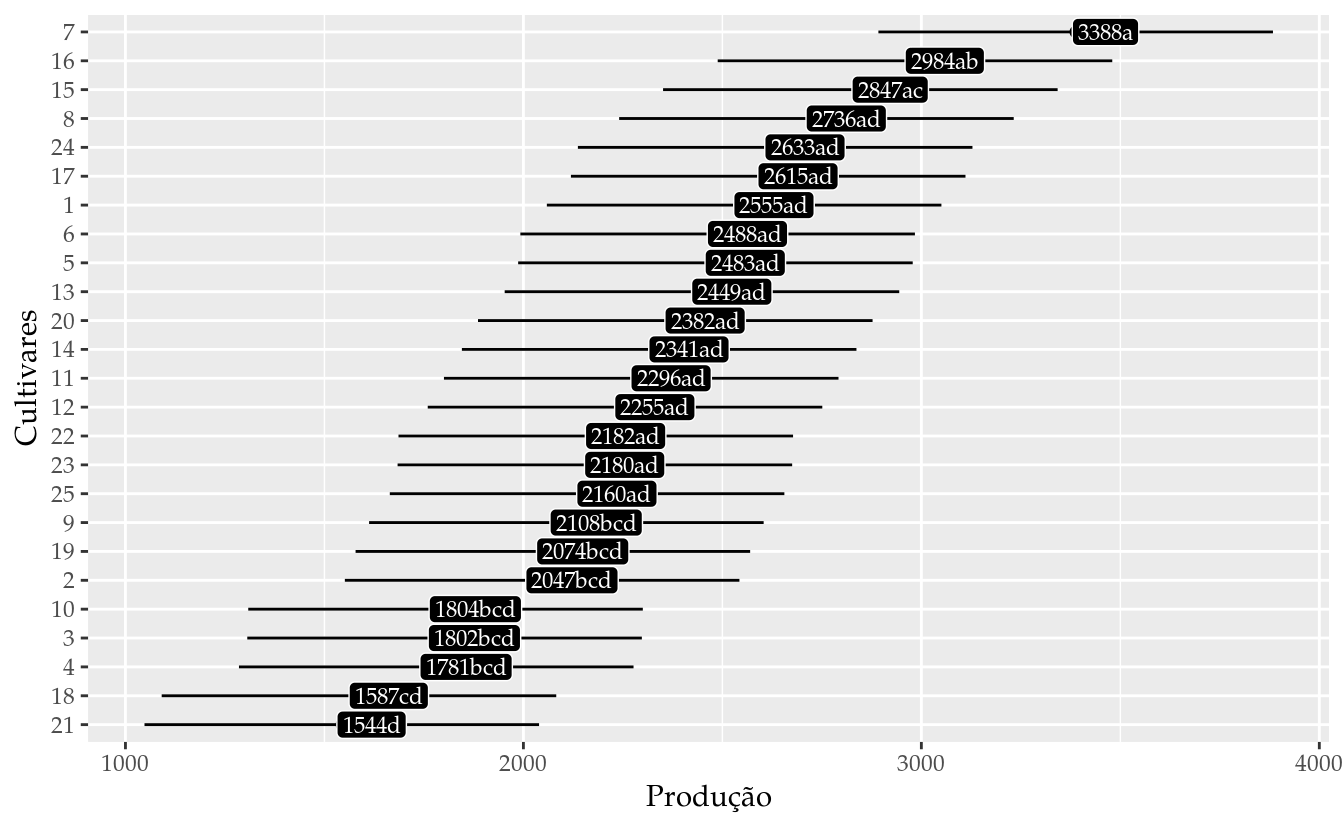
16.4 ZimmermannTb7.4
Os dados em ZimmermannTb7.4 são registros da produção de grãos (kg/ha) de um experimento em látice reticulado retangular de competição de cultivares e linhagens de arroz irrigado. Foram avaliadas \(t = 30\) tratamentos com \(r = 3\) repetições usando-se \(b = 18\) blocos de tamanho \(k = 5\), o que soma \(tr = bk = 90\) unidades experimentais.
da <- labestData::ZimmermannTb7.4
str(da)## 'data.frame': 90 obs. of 4 variables:
## $ rept: Factor w/ 3 levels "X","Y","Z": 1 1 1 1 1 1 1 1 1 1 ...
## $ bloc: Factor w/ 18 levels "X1","X2","X3",..: 1 1 1 1 1 2 2 2 2 2 ...
## $ cult: Factor w/ 30 levels "1","2","3","4",..: 5 1 4 3 2 10 9 6 8 7 ...
## $ prod: num 4256 4628 4890 3713 4602 ...# NOTE: Codificar bloco dentro de repetição.
da <- da %>%
group_by(rept) %>%
mutate(bloc = factor(as.numeric(bloc), labels = 1:6)) %>%
ungroup()
# São 6 blocos de tamanho 5 em cada uma das 3 repetições para acomodar
# 30 tratamentos.
with(da, table(cult, rept:bloc)) %>%
addmargins(1:2) %>%
print.table(zero.print = ".")##
## cult X:1 X:2 X:3 X:4 X:5 X:6 Y:1 Y:2 Y:3 Y:4 Y:5 Y:6 Z:1 Z:2 Z:3
## 1 1 . . . . . . 1 . . . . . . 1
## 2 1 . . . . . . . 1 . . . . . .
## 3 1 . . . . . . . . 1 . . . . .
## 4 1 . . . . . . . . . 1 . . . .
## 5 1 . . . . . . . . . . 1 . 1 .
## 6 . 1 . . . . 1 . . . . . . . .
## 7 . 1 . . . . . . 1 . . . 1 . .
## 8 . 1 . . . . . . . 1 . . . . 1
## 9 . 1 . . . . . . . . 1 . . . .
## 10 . 1 . . . . . . . . . 1 . . .
## 11 . . 1 . . . 1 . . . . . . . .
## 12 . . 1 . . . . 1 . . . . . . .
## 13 . . 1 . . . . . . 1 . . 1 . .
## 14 . . 1 . . . . . . . 1 . . 1 .
## 15 . . 1 . . . . . . . . 1 . . .
## 16 . . . 1 . . 1 . . . . . . 1 .
## 17 . . . 1 . . . 1 . . . . . . .
## 18 . . . 1 . . . . 1 . . . . . .
## 19 . . . 1 . . . . . . 1 . 1 . .
## 20 . . . 1 . . . . . . . 1 . . 1
## 21 . . . . 1 . 1 . . . . . . . 1
## 22 . . . . 1 . . 1 . . . . . . .
## 23 . . . . 1 . . . 1 . . . . 1 .
## 24 . . . . 1 . . . . 1 . . . . .
## 25 . . . . 1 . . . . . . 1 1 . .
## 26 . . . . . 1 1 . . . . . . . .
## 27 . . . . . 1 . 1 . . . . 1 . .
## 28 . . . . . 1 . . 1 . . . . . .
## 29 . . . . . 1 . . . 1 . . . 1 .
## 30 . . . . . 1 . . . . 1 . . . 1
## Sum 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5
##
## cult Z:4 Z:5 Z:6 Sum
## 1 . . . 3
## 2 1 . . 3
## 3 . 1 . 3
## 4 . . 1 3
## 5 . . . 3
## 6 . . 1 3
## 7 . . . 3
## 8 . . . 3
## 9 1 . . 3
## 10 . 1 . 3
## 11 . 1 . 3
## 12 . . 1 3
## 13 . . . 3
## 14 . . . 3
## 15 1 . . 3
## 16 . . . 3
## 17 . 1 . 3
## 18 . . 1 3
## 19 . . . 3
## 20 . . . 3
## 21 . . . 3
## 22 1 . . 3
## 23 . . . 3
## 24 . . 1 3
## 25 . . . 3
## 26 1 . . 3
## 27 . . . 3
## 28 . 1 . 3
## 29 . . . 3
## 30 . . . 3
## Sum 5 5 5 90library(igraph)
da$cond <- with(da, interaction(rept, bloc))
edg <- by(data = as.integer(da$cult),
INDICES = da$cond,
FUN = combn,
m = 2) %>%
flatten_int()
ghp <- graph(edg, directed = FALSE)
plot(ghp,
vertex.size = 5,
vertex.label.dist = 1,
# layout = layout_components,
layout = layout_in_circle,
edge.curved = FALSE)
# Quantidade de pares possíveis.
choose(nlevels(da$cult), k = 2)
# Todos os possíveis pares.
pares <- apply(combn(sort(as.character(levels(da$cult))), m = 2),
MARGIN = 2,
FUN = paste0,
collapse = "_") %>%
tibble(name = ., value = 0)
# Quantas vezes cada par ocorre junto.
by(data = da$cult,
INDICES = da$cond,
FUN = function(x) {
apply(combn(sort(x), m = 2),
MARGIN = 2,
FUN = paste0,
collapse = "_")
}) %>%
flatten_chr() %>%
table() %>%
enframe() %>%
bind_rows(pares) %>%
distinct(name, .keep_all = TRUE) %>%
split(x = .$name, f = .$value)# Modelo de efeitos fixos.
m0 <- lm(terms(prod ~ rept/bloc + cult, keep.order = TRUE),
data = da)
# Quado de anova com hipóteses marginais.
Anova(m0)## Anova Table (Type II tests)
##
## Response: prod
## Sum Sq Df F value Pr(>F)
## rept 1439299 2 2.4512 0.09815 .
## rept:bloc 8493032 15 1.9285 0.04698 *
## cult 51348862 29 6.0309 7.942e-08 ***
## Residuals 12624615 43
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Médias marginais ajustadas.
emm <- emmeans(m0, specs = ~cult)## NOTE: A nesting structure was detected in the fitted model:
## bloc %in% reptemm## cult emmean SE df lower.CL upper.CL
## 1 5011 352 43 4302 5721
## 2 5150 352 43 4441 5860
## 3 4203 352 43 3494 4913
## 4 4389 352 43 3680 5099
## 5 4249 352 43 3540 4958
## 6 4997 352 43 4287 5706
## 7 5561 352 43 4851 6270
## 8 5339 352 43 4629 6048
## 9 5741 352 43 5032 6451
## 10 5555 352 43 4845 6264
## 11 5253 352 43 4544 5962
## 12 5232 352 43 4522 5941
## 13 4603 352 43 3894 5312
## 14 5325 352 43 4616 6035
## 15 6742 352 43 6033 7452
## 16 6064 352 43 5355 6773
## 17 6428 352 43 5719 7137
## 18 5657 352 43 4947 6366
## 19 6134 352 43 5425 6843
## 20 5635 352 43 4926 6345
## 21 5251 352 43 4542 5961
## 22 4902 352 43 4193 5611
## 23 4498 352 43 3789 5207
## 24 5360 352 43 4651 6070
## 25 5502 352 43 4793 6212
## 26 3939 352 43 3230 4649
## 27 4837 352 43 4127 5546
## 28 7800 352 43 7091 8509
## 29 5135 352 43 4426 5845
## 30 6961 352 43 6251 7670
##
## Results are averaged over the levels of: bloc, rept
## Confidence level used: 0.95# Extração da matriz de funções lineares.
L <- attr(emm, "linfct")
grid <- attr(emm, "grid")
rownames(L) <- grid[[1]]
# Entenda como são obtidas as médias marginais.
MASS::fractions(t(L[1:4, ])) %>%
unique()## 1 2 3 4
## (Intercept) 1 1 1 1
## reptY 1/3 1/3 1/3 1/3
## reptX:bloc2 1/18 1/18 1/18 1/18
## cult2 0 1 0 0
## cult3 0 0 1 0
## cult4 0 0 0 1
## cult5 0 0 0 0# Contrastes par a par.
ctr <- summary(glht(m0, linfct = all_pairwise(L)),
test = adjusted(type = "fdr"))
# Erros padrões de vários tamanhos conforme estrutura de associados.
v <- c("coefficients", "sigma", "tstat", "pvalues")
ctr$test[v] %>%
as.data.frame() %>%
split(., round(.$sigma, 4)) %>%
map(head, n = 4)## $`485.388`
## coefficients sigma tstat pvalues
## 1vs3 807.9878 485.388 1.664623 0.21911133
## 1vs4 622.0736 485.388 1.281601 0.36136548
## 1vs17 -1416.6785 485.388 -2.918652 0.02754887
## 1vs20 -623.9886 485.388 -1.285546 0.36136548
##
## $`487.2513`
## coefficients sigma tstat pvalues
## 1vs2 -138.9827 487.2513 -0.2852383 0.8620446
## 1vs5 762.5234 487.2513 1.5649490 0.2533732
## 1vs8 -327.2198 487.2513 -0.6715627 0.6703432
## 1vs12 -220.0973 487.2513 -0.4517122 0.7695007
##
## $`489.1075`
## coefficients sigma tstat pvalues
## 9vs22 839.268889 489.1075 1.715919173 0.20727666
## 11vs21 1.598889 489.1075 0.003268993 0.99740684
## 14vs23 827.227037 489.1075 1.691299120 0.21212478
## 15vs20 1106.852963 489.1075 2.263005630 0.08994443
##
## $`505.5067`
## coefficients sigma tstat pvalues
## 1vs24 -348.9206 505.5067 -0.6902394 0.6588416
## 2vs10 -404.2844 505.5067 -0.7997607 0.5994884
## 3vs12 -1028.0852 505.5067 -2.0337718 0.1270049
## 4vs8 -949.2934 505.5067 -1.8779048 0.1623681
##
## $`507.2961`
## coefficients sigma tstat pvalues
## 1vs9 -729.8814 507.2961 -1.438768 0.297796903
## 1vs10 -543.2671 507.2961 -1.070907 0.469258810
## 1vs15 -1730.8416 507.2961 -3.411896 0.009931507
## 1vs18 -645.2338 507.2961 -1.271908 0.364363491
##
## $`509.0792`
## coefficients sigma tstat pvalues
## 1vs6 14.75448 509.0792 0.02898267 0.9837062
## 1vs11 -241.43049 509.0792 -0.47424938 0.7642118
## 1vs13 408.44096 509.0792 0.80231319 0.5994884
## 1vs14 -313.85901 509.0792 -0.61652296 0.6888182
##
## $`510.8561`
## coefficients sigma tstat pvalues
## 1vs7 -549.1352 510.8561 -1.074931 0.469258810
## 2vs13 547.4237 510.8561 1.071581 0.469258810
## 3vs19 -1930.7022 510.8561 -3.779347 0.004256734
## 4vs25 -1112.8578 510.8561 -2.178417 0.101900890# Comparações múltiplas.
results_m0 <- wzRfun::apmc(X = L,
model = m0,
focus = "cult",
test = "bonferroni",
level = 0.05)
results_m0## cult fit lwr upr cld
## 1 1 5011.481 4302.217 5720.746 bdf
## 2 2 5150.464 4441.199 5859.729 bdf
## 3 3 4203.493 3494.229 4912.758 ef
## 4 4 4389.408 3680.143 5098.672 df
## 5 5 4248.958 3539.693 4958.222 ef
## 6 6 4996.727 4287.462 5705.991 bdf
## 7 7 5560.616 4851.352 6269.881 bdf
## 8 8 5338.701 4629.436 6047.966 bdf
## 9 9 5741.363 5032.098 6450.627 adf
## 10 10 5554.748 4845.484 6264.013 bdf
## 11 11 5252.912 4543.647 5962.176 bdf
## 12 12 5231.579 4522.314 5940.843 bdf
## 13 13 4603.040 3893.776 5312.305 df
## 14 14 5325.340 4616.076 6034.605 bdf
## 15 15 6742.323 6033.058 7451.587 abc
## 16 16 6063.854 5354.589 6773.119 ade
## 17 17 6428.160 5718.895 7137.424 ad
## 18 18 5656.715 4947.450 6365.980 bdf
## 19 19 6134.196 5424.931 6843.460 ade
## 20 20 5635.470 4926.205 6344.734 bdf
## 21 21 5251.313 4542.048 5960.577 bdf
## 22 22 4902.094 4192.829 5611.358 bdf
## 23 23 4498.113 3788.849 5207.378 df
## 24 24 5360.402 4651.137 6069.666 bdf
## 25 25 5502.265 4793.001 6211.530 bdf
## 26 26 3939.261 3229.996 4648.526 f
## 27 27 4836.762 4127.497 5546.026 cdf
## 28 28 7799.973 7090.708 8509.237 a
## 29 29 5135.264 4426.000 5844.529 bdf
## 30 30 6960.754 6251.490 7670.019 ab# Gráfico de segmentos para as estimativas intervalares.
ggplot(data = results_m0,
mapping = aes(x = fit, y = reorder(cult, fit))) +
geom_point() +
geom_errorbarh(mapping = aes(xmin = lwr, xmax = upr),
height = 0) +
geom_label(mapping = aes(label = sprintf("%0.0f%s", fit, cld)),
label.padding = unit(0.15, "lines"),
fill = "black",
colour = "white",
size = 3,
nudge_x = 120,
vjust = 0.5) +
labs(x = "Produção",
y = "Cultivares")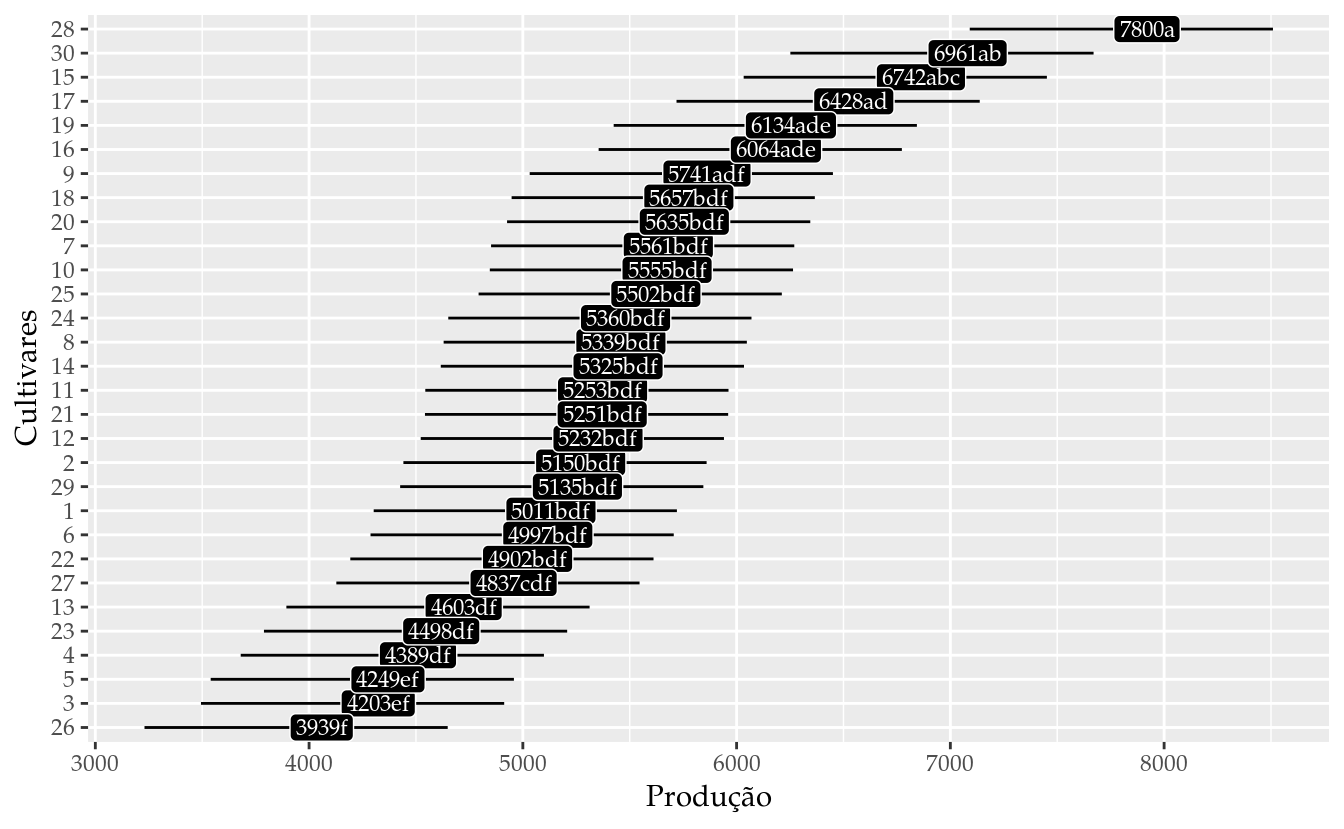
FIXME: por que são 6 tamanhos de erro padrão no reticulado retangular?
# Ajuste do modelo de efeito aleatório de bloc.
mm0 <- lmer(prod ~ rept + (1 | rept:bloc) + cult, data = da)
# Quadro de teste de Wald.
anova(mm0)## Type III Analysis of Variance Table with Satterthwaite's method
## Sum Sq Mean Sq NumDF DenDF F value Pr(>F)
## rept 611521 305761 2 9.274 1.0295 0.3947
## cult 51726584 1783675 29 48.360 6.0056 2.562e-08 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Estimativas dos parâmetros dos termos de efeito.
summary(mm0)## Linear mixed model fit by REML. t-tests use Satterthwaite's method
## [lmerModLmerTest]
## Formula: prod ~ rept + (1 | rept:bloc) + cult
## Data: da
##
## REML criterion at convergence: 943.8
##
## Scaled residuals:
## Min 1Q Median 3Q Max
## -1.94004 -0.52107 -0.04114 0.47160 1.87929
##
## Random effects:
## Groups Name Variance Std.Dev.
## rept:bloc (Intercept) 80407 283.6
## Residual 297003 545.0
## Number of obs: 90, groups: rept:bloc, 18
##
## Fixed effects:
## Estimate Std. Error df t value Pr(>|t|)
## (Intercept) 4906.836 361.949 56.728 13.557 < 2e-16 ***
## reptY 303.314 215.876 9.274 1.405 0.19261
## reptZ 206.111 215.876 9.274 0.955 0.36392
## cult2 -27.959 466.149 50.862 -0.060 0.95241
## cult3 -880.042 465.690 50.532 -1.890 0.06453 .
## cult4 -620.446 465.690 50.532 -1.332 0.18873
## cult5 -891.870 466.149 50.862 -1.913 0.06135 .
## cult6 16.176 476.608 54.222 0.034 0.97305
## cult7 511.132 477.056 54.463 1.071 0.28870
## cult8 519.040 466.149 50.862 1.113 0.27074
## cult9 596.159 476.159 53.973 1.252 0.21596
## cult10 245.437 476.159 53.973 0.515 0.60834
## cult11 28.563 476.608 54.222 0.060 0.95243
## cult12 240.731 466.149 50.862 0.516 0.60779
## cult13 -271.559 476.608 54.222 -0.570 0.57118
## cult14 286.280 476.608 54.222 0.601 0.55057
## cult15 1513.008 476.159 53.973 3.178 0.00246 **
## cult16 960.478 476.608 54.222 2.015 0.04885 *
## cult17 1125.649 465.690 50.532 2.417 0.01929 *
## cult18 586.485 476.159 53.973 1.232 0.22340
## cult19 1021.613 476.159 53.973 2.146 0.03643 *
## cult20 493.714 465.690 50.532 1.060 0.29411
## cult21 316.072 466.149 50.862 0.678 0.50081
## cult22 -198.869 465.690 50.532 -0.427 0.67116
## cult23 -502.825 476.608 54.222 -1.055 0.29610
## cult24 586.609 475.710 53.717 1.233 0.22290
## cult25 427.123 476.159 53.973 0.897 0.37369
## cult26 -1264.213 476.159 53.973 -2.655 0.01040 *
## cult27 -268.424 466.149 50.862 -0.576 0.56727
## cult28 2497.388 476.159 53.973 5.245 2.68e-06 ***
## cult29 201.801 476.159 53.973 0.424 0.67339
## cult30 1909.155 465.690 50.532 4.100 0.00015 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1##
## Correlation matrix not shown by default, as p = 32 > 12.
## Use print(x, correlation=TRUE) or
## vcov(x) if you need it# Médias marginais ajustadas.
emm <- emmeans(mm0, specs = ~cult)
emm## cult emmean SE df lower.CL upper.CL
## 1 5077 347 57 4382 5771
## 2 5049 347 57 4354 5743
## 3 4197 347 57 3502 4891
## 4 4456 347 57 3762 5151
## 5 4185 347 57 3490 4879
## 6 5093 347 57 4398 5787
## 7 5588 347 57 4893 6282
## 8 5596 347 57 4901 6290
## 9 5673 347 57 4978 6367
## 10 5322 347 57 4628 6016
## 11 5105 347 57 4411 5800
## 12 5317 347 57 4623 6012
## 13 4805 347 57 4111 5499
## 14 5363 347 57 4669 6057
## 15 6590 347 57 5895 7284
## 16 6037 347 57 5343 6731
## 17 6202 347 57 5508 6897
## 18 5663 347 57 4969 6357
## 19 6098 347 57 5404 6793
## 20 5570 347 57 4876 6265
## 21 5393 347 57 4698 6087
## 22 4878 347 57 4183 5572
## 23 4574 347 57 3879 5268
## 24 5663 347 57 4969 6358
## 25 5504 347 57 4809 6198
## 26 3812 347 57 3118 4507
## 27 4808 347 57 4114 5503
## 28 7574 347 57 6880 8268
## 29 5278 347 57 4584 5973
## 30 6986 347 57 6291 7680
##
## Results are averaged over the levels of: rept
## Degrees-of-freedom method: kenward-roger
## Confidence level used: 0.95# Extração da matriz de funções lineares.
grid <- attr(emm, "grid")
L <- attr(emm, "linfct")
rownames(L) <- grid[[1]]
# Contrastes par a par.
ctr <- summary(glht(mm0, linfct = all_pairwise(L)),
test = adjusted(type = "fdr"))
# Erros padrões de dois tamanhos conforme estrutura de associados.
v <- c("coefficients", "sigma", "tstat", "pvalues")
ctr$test[v] %>%
as.data.frame() %>%
split(., round(.$sigma, 4)) %>%
map(head, n = 4)## $`465.6904`
## coefficients sigma tstat pvalues
## 1vs3 880.0416 465.6904 1.889757 0.14177285
## 1vs4 620.4462 465.6904 1.332315 0.33686104
## 1vs17 -1125.6486 465.6904 -2.417161 0.05121174
## 1vs20 -493.7140 465.6904 -1.060176 0.44589718
##
## $`466.1488`
## coefficients sigma tstat pvalues
## 1vs2 27.95906 466.1488 0.05997882 0.9769152
## 1vs5 891.87026 466.1488 1.91327355 0.1376998
## 1vs8 -519.03956 466.1488 -1.11346314 0.4293556
## 1vs12 -240.73145 466.1488 -0.51642613 0.7264838
##
## $`466.6068`
## coefficients sigma tstat pvalues
## 9vs22 795.0282 466.6068 1.703850 0.19521798
## 11vs21 -287.5087 466.6068 -0.616169 0.68804584
## 14vs23 789.1052 466.6068 1.691157 0.19750502
## 15vs20 1019.2939 466.6068 2.184481 0.08388801
##
## $`475.7104`
## coefficients sigma tstat pvalues
## 1vs24 -586.6085 475.7104 -1.2331210 0.37641531
## 2vs10 -273.3956 475.7104 -0.5747102 0.69714866
## 3vs12 -1120.7730 475.7104 -2.3559985 0.05825712
## 4vs8 -1139.4857 475.7104 -2.3953348 0.05390492
##
## $`476.1592`
## coefficients sigma tstat pvalues
## 1vs9 -596.1593 476.1592 -1.2520168 0.369335609
## 1vs10 -245.4366 476.1592 -0.5154506 0.726483752
## 1vs15 -1513.0079 476.1592 -3.1775252 0.008501835
## 1vs18 -586.4852 476.1592 -1.2316997 0.376415309
##
## $`476.6076`
## coefficients sigma tstat pvalues
## 1vs6 -16.17564 476.6076 -0.03393911 0.9911538
## 1vs11 -28.56294 476.6076 -0.05992968 0.9769152
## 1vs13 271.55915 476.6076 0.56977514 0.6971487
## 1vs14 -286.27995 476.6076 -0.60066178 0.6950682
##
## $`477.0555`
## coefficients sigma tstat pvalues
## 1vs7 -511.1321 477.0555 -1.0714310 0.4411764685
## 2vs13 243.6001 477.0555 0.5106326 0.7285154857
## 3vs19 -1901.6550 477.0555 -3.9862341 0.0006213121
## 4vs25 -1047.5694 477.0555 -2.1959067 0.0820328981# As mesmas comparações múltiplas.
results_mm0 <- wzRfun::apmc(L,
model = mm0,
focus = "cult",
test = "bonferroni",
level = 0.05)
results_mm0## cult fit lwr upr cld
## 1 1 5076.645 4410.624 5742.665 cdf
## 2 2 5048.686 4382.665 5714.706 cdf
## 3 3 4196.603 3530.583 4862.624 ef
## 4 4 4456.199 3790.178 5122.219 df
## 5 5 4184.775 3518.754 4850.795 ef
## 6 6 5092.820 4426.800 5758.841 cdf
## 7 7 5587.777 4921.756 6253.797 bcdf
## 8 8 5595.684 4929.664 6261.705 bcdf
## 9 9 5672.804 5006.784 6338.825 bcde
## 10 10 5322.081 4656.061 5988.102 bcdf
## 11 11 5105.208 4439.187 5771.228 cdf
## 12 12 5317.376 4651.356 5983.397 bcdf
## 13 13 4805.086 4139.065 5471.106 cdf
## 14 14 5362.925 4696.904 6028.945 bcdf
## 15 15 6589.653 5923.632 7255.673 ac
## 16 16 6037.122 5371.102 6703.143 ad
## 17 17 6202.293 5536.273 6868.314 ad
## 18 18 5663.130 4997.109 6329.150 bcde
## 19 19 6098.258 5432.238 6764.279 ad
## 20 20 5570.359 4904.338 6236.379 bcdf
## 21 21 5392.716 4726.696 6058.737 bcdf
## 22 22 4877.776 4211.755 5543.796 cdf
## 23 23 4573.820 3907.799 5239.840 df
## 24 24 5663.253 4997.233 6329.274 bcde
## 25 25 5503.768 4837.747 6169.789 bcdf
## 26 26 3812.432 3146.411 4478.452 f
## 27 27 4808.220 4142.200 5474.241 cdf
## 28 28 7574.033 6908.012 8240.053 a
## 29 29 5278.446 4612.425 5944.466 bcdf
## 30 30 6985.800 6319.780 7651.821 ab# Gráfico de segmentos para as estimativas intervalares.
ggplot(data = results_mm0,
mapping = aes(x = fit, y = reorder(cult, fit))) +
geom_point() +
geom_errorbarh(mapping = aes(xmin = lwr, xmax = upr),
height = 0) +
geom_label(mapping = aes(label = sprintf("%0.0f%s", fit, cld)),
label.padding = unit(0.15, "lines"),
fill = "black",
colour = "white",
size = 3,
nudge_x = 120,
vjust = 0.5) +
labs(x = "Produção",
y = "Cultivares")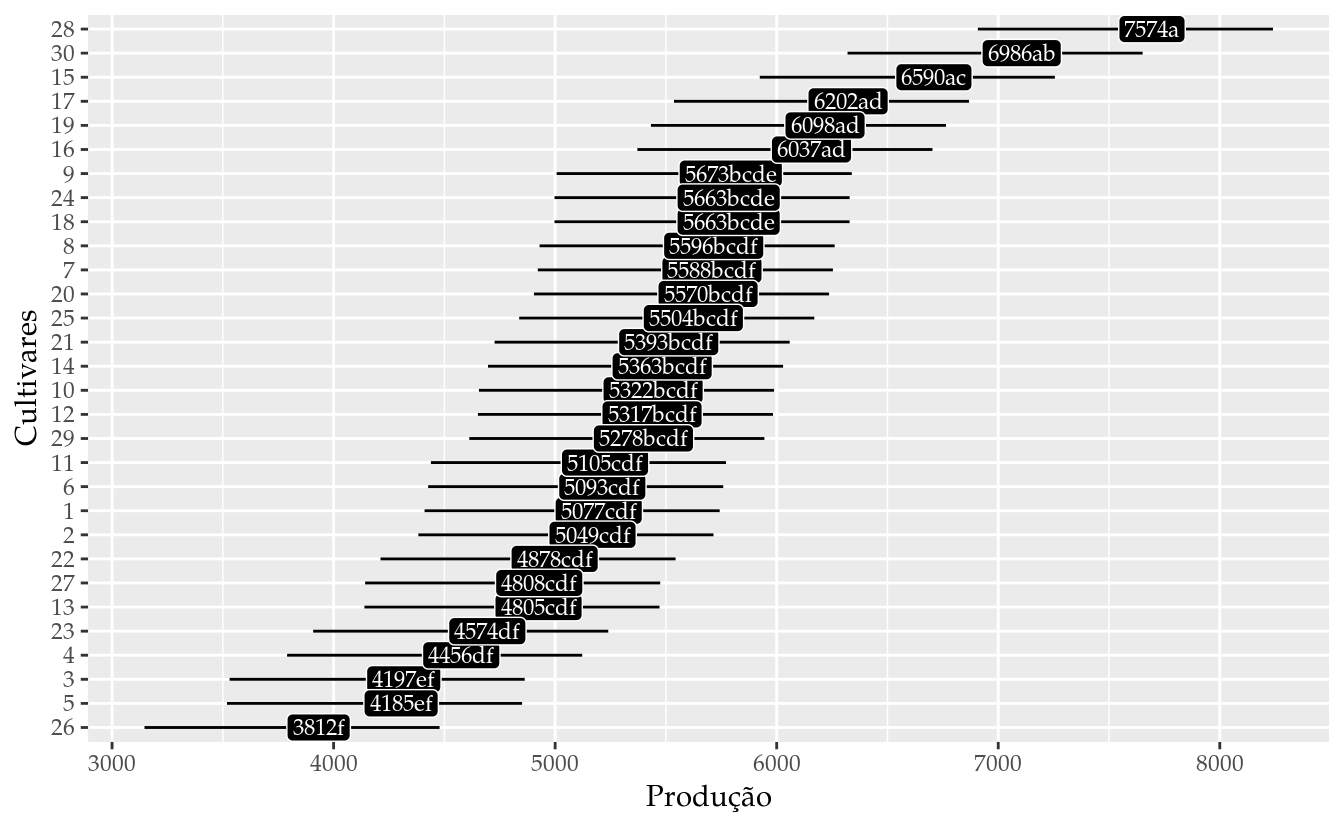
Referências Bibliográficas
RAMALHO, M. A. P.; FERREIRA, D. F.; OLIVEIRA, A. C. Experimentação em genética e melhoramento de plantas. 2nd ed. Lavras, MG: Editora UFLA, 2005.
Manual de Planejamento e Análise de Experimentos com R
Walmes Marques Zeviani