Capítulo 6 Revisão de modelos lineares
6.1 Estimação
#-----------------------------------------------------------------------
library(lattice)
# tá faltando a cov.r e o VIF.
#-----------------------------------------------------------------------
# Importação dos dados.
url <- "http://www.leg.ufpr.br/~walmes/data/business_economics_dataset/EXAMPLES/REALESTA.DAT"
da <- read.table(url, header = FALSE)
names(da) <- c("number", "saleprice", "landvalue",
"improvvalue", "area")
str(da)## 'data.frame': 20 obs. of 5 variables:
## $ number : int 1 2 3 4 5 6 7 8 9 10 ...
## $ saleprice : int 68900 48500 55500 62000 116500 45000 38000 83000 59000 47500 ...
## $ landvalue : int 5960 9000 9500 10000 18000 8500 8000 23000 8100 9000 ...
## $ improvvalue: int 44967 27860 31439 39592 72827 27317 29856 47752 39117 29349 ...
## $ area : int 1873 928 1126 1265 2214 912 899 1803 1204 1725 ...summary(da)## number saleprice landvalue improvvalue
## Min. : 1.00 Min. : 22400 Min. : 1500 Min. : 5779
## 1st Qu.: 5.75 1st Qu.: 40800 1st Qu.: 6965 1st Qu.:26351
## Median :10.50 Median : 49250 Median : 8050 Median :31559
## Mean :10.50 Mean : 56660 Mean : 9213 Mean :35311
## 3rd Qu.:15.25 3rd Qu.: 63725 3rd Qu.: 9625 3rd Qu.:41366
## Max. :20.00 Max. :116500 Max. :23000 Max. :72827
## area
## Min. : 899
## 1st Qu.:1054
## Median :1188
## Mean :1383
## 3rd Qu.:1744
## Max. :2455# Divide valores por 1000
i <- 2:5
da[,i] <- sapply(i,
function(i){
da[,i]/1000
})
str(da)## 'data.frame': 20 obs. of 5 variables:
## $ number : int 1 2 3 4 5 6 7 8 9 10 ...
## $ saleprice : num 68.9 48.5 55.5 62 116.5 ...
## $ landvalue : num 5.96 9 9.5 10 18 8.5 8 23 8.1 9 ...
## $ improvvalue: num 45 27.9 31.4 39.6 72.8 ...
## $ area : num 1.873 0.928 1.126 1.265 2.214 ...#-----------------------------------------------------------------------
# Análise exploratória.
xyplot(saleprice ~ landvalue, data = da, type = c("p", "smooth"))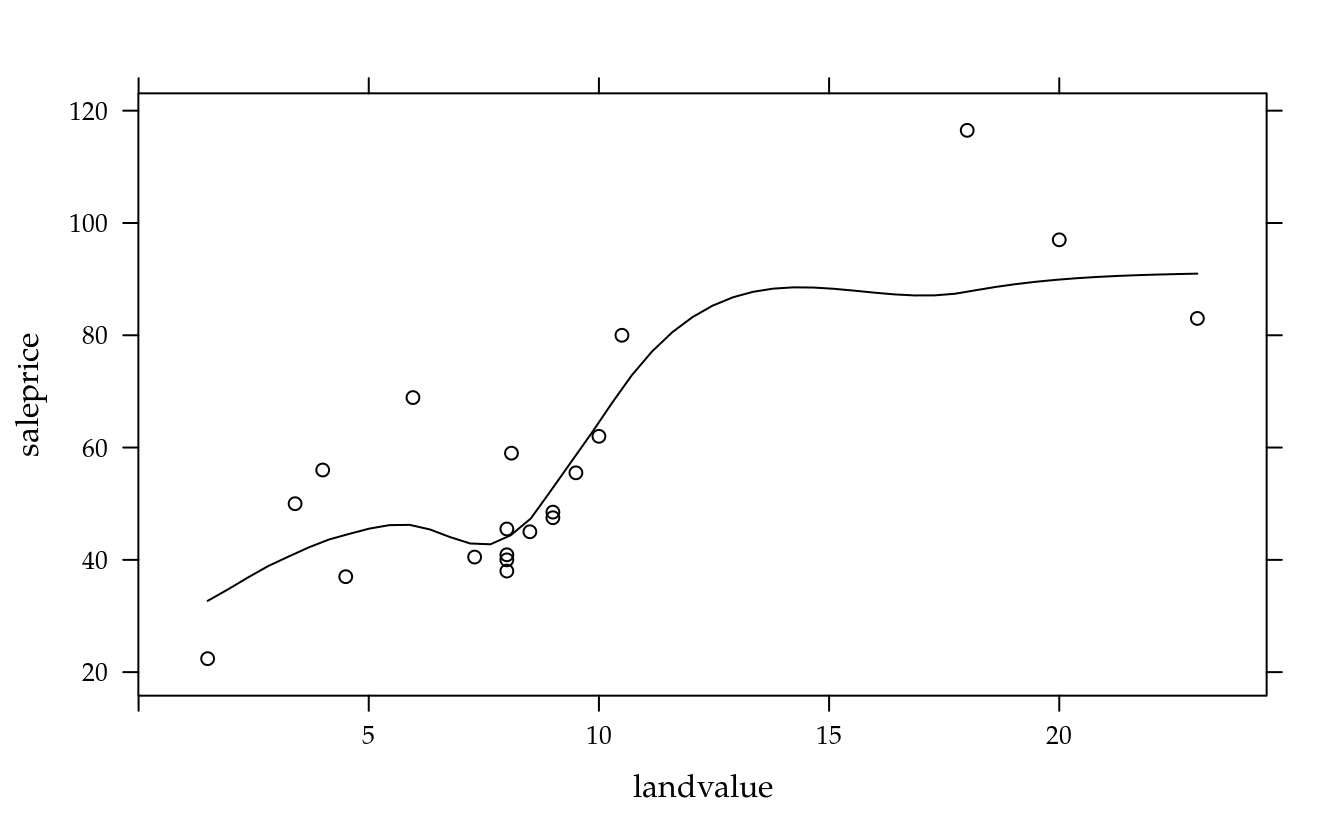
xyplot(saleprice ~ improvvalue, data = da, type = c("p", "smooth"))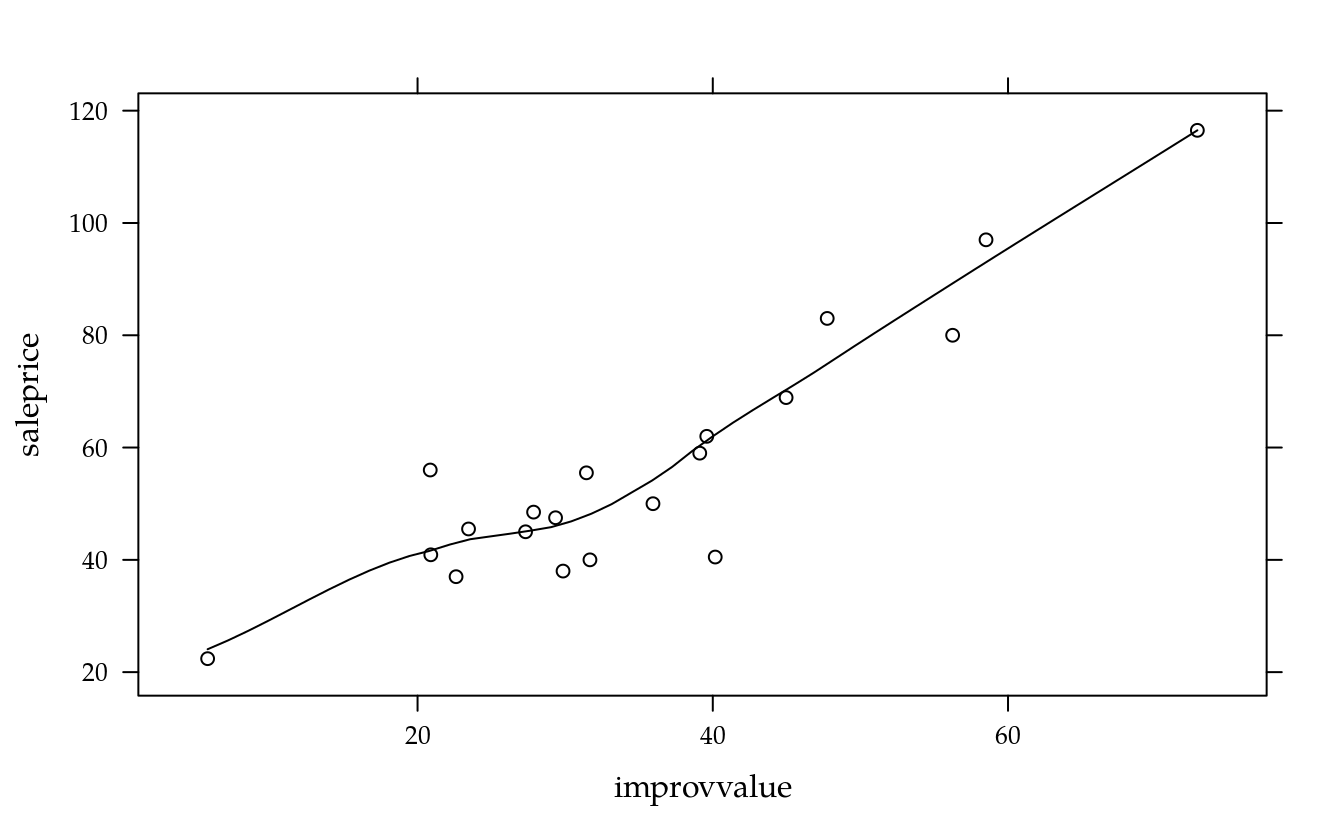
xyplot(saleprice ~ area, data = da, type = c("p", "smooth"))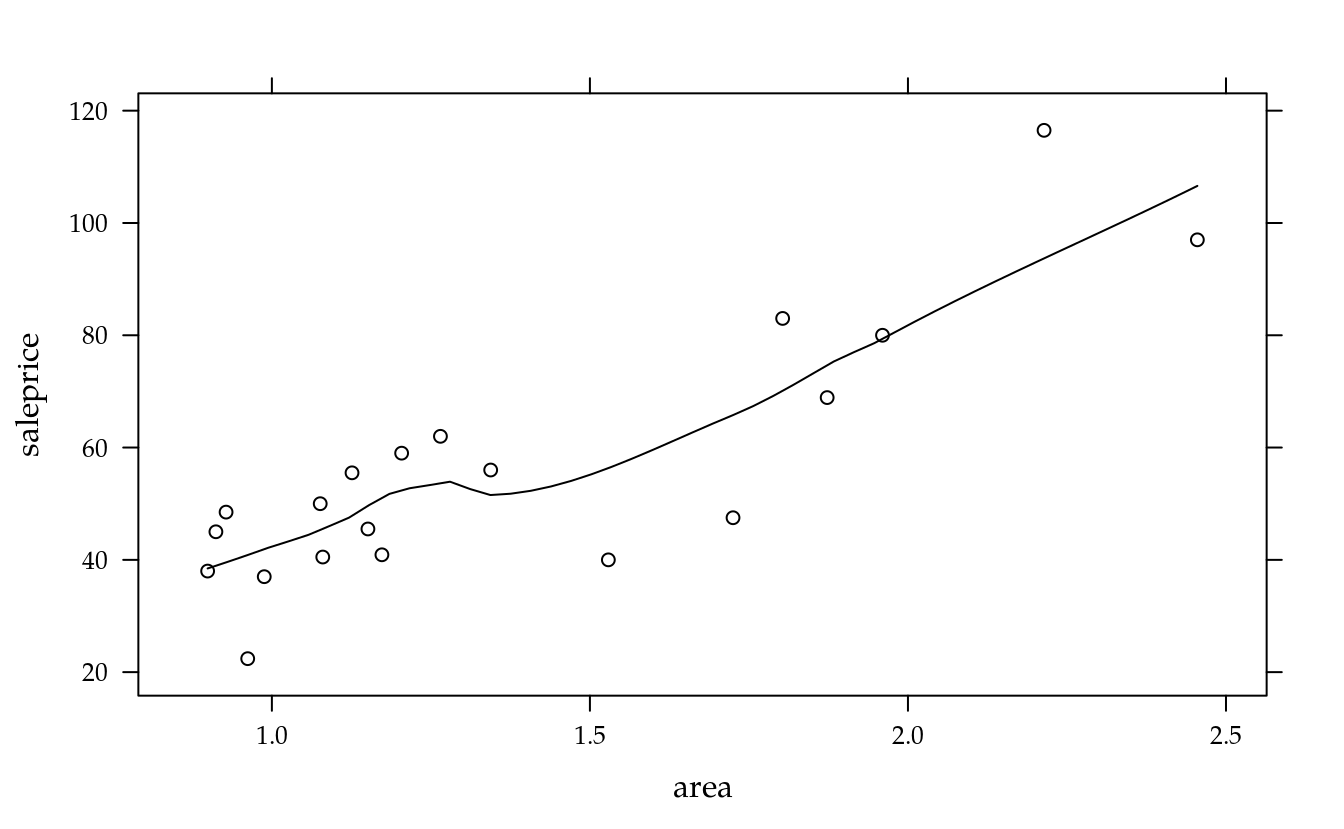
#-----------------------------------------------------------------------
# Ajuste com funções do R.
m0 <- lm(saleprice ~ landvalue + improvvalue + area,
data = da)
par(mfrow = c(2, 2))
plot(m0)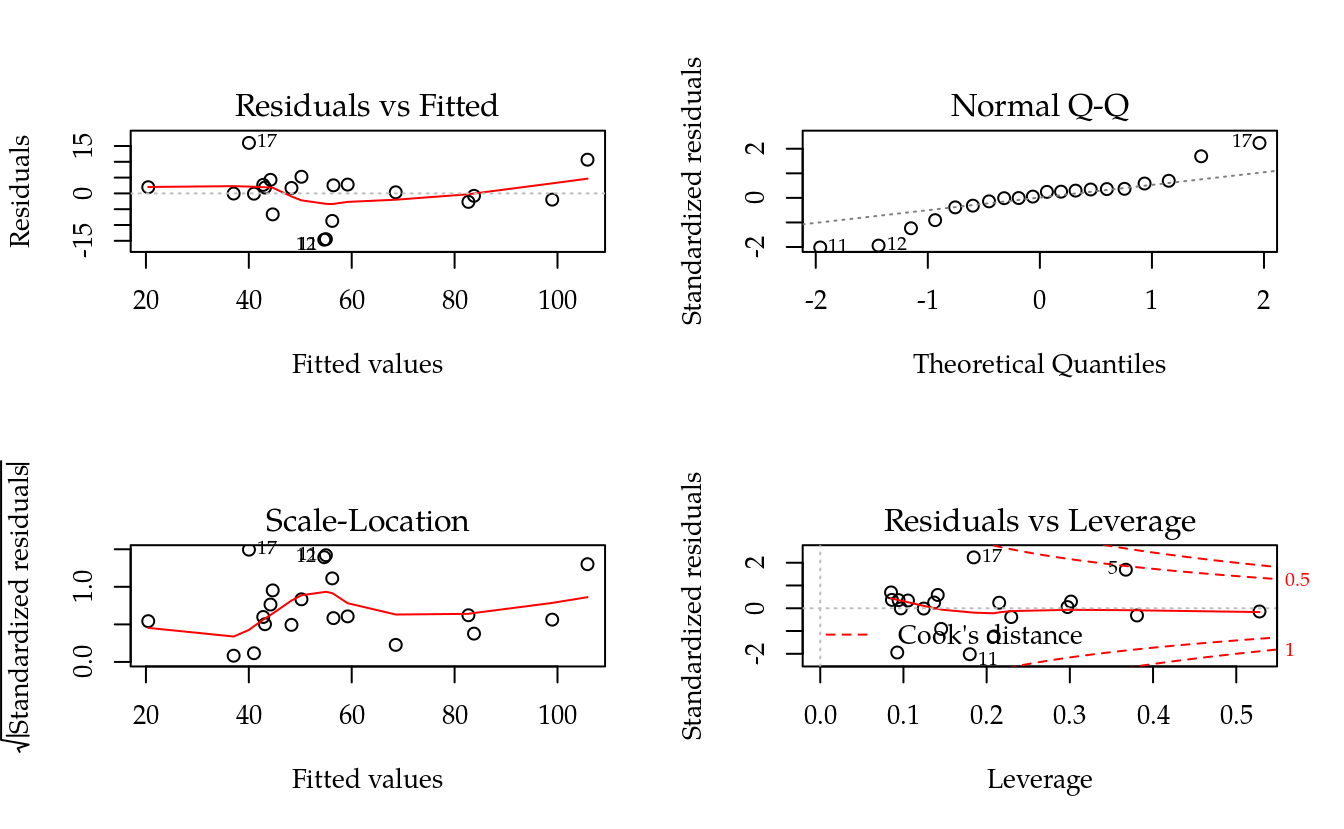
layout(1)
par(mfrow = c(2, 3))
plot(m0, which = 1:6)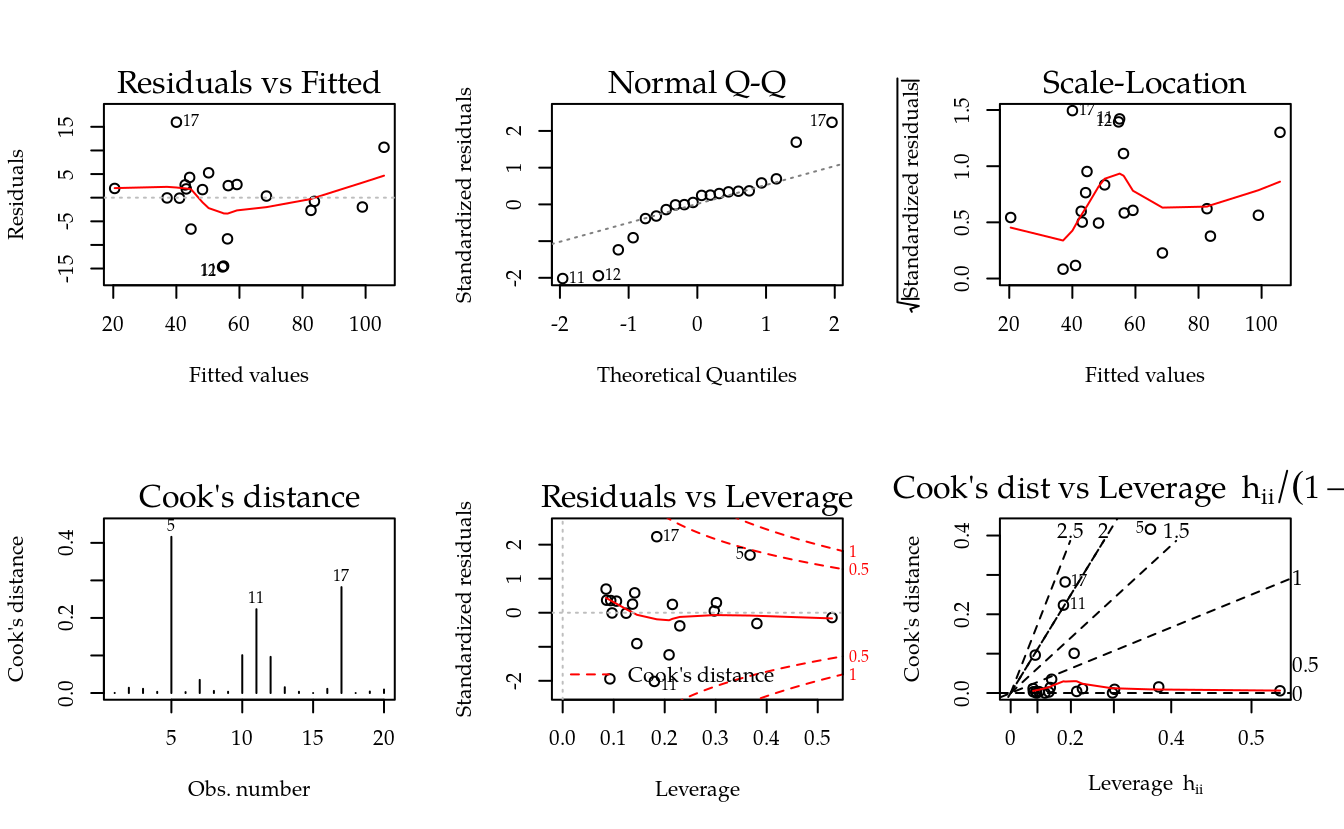
layout(1)
anova(m0)## Analysis of Variance Table
##
## Response: saleprice
## Df Sum Sq Mean Sq F value Pr(>F)
## landvalue 1 6102.2 6102.2 97.296 3.323e-08 ***
## improvvalue 1 2412.8 2412.8 38.470 1.267e-05 ***
## area 1 264.7 264.7 4.220 0.05666 .
## Residuals 16 1003.5 62.7
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1#-----------------------------------------------------------------------
# Ajuste do modelo de forma matricial.
# Matriz do modelo e vetor de observações.
X <- with(da, cbind(b0 = 1,
b1 = landvalue,
b2 = improvvalue,
b3 = area))
y <- with(da, cbind(saleprice))
# Produto de matrizes e inversa.
XlX <- crossprod(X) # X'X
Xly <- crossprod(X, y) # X'y
iXlX <- solve(XlX) # (X'X)^{-1}
# Estimativas dos parâmetros.
b <- solve(XlX, Xly)
b## saleprice
## b0 1.4702759
## b1 0.8144901
## b2 0.8204447
## b3 13.5286499# Matrizes de projeção.
I <- diag(nrow(y)) # I (n x n)
H <- X %*% solve(XlX, t(X)) # X (X'X)^{-1} X' (n x n)
# Função para criar matrizes de projeção.
proj <- function(X) {
X %*% solve(crossprod(X), t(X))
}
# Função para calcular o traço de matrizes.
trace <- function(H){
sum(diag(H))
}
# Traço das matrizes de projeção que são os graus de liberdade que é a
# dimensão do espaço em que é feita a projeção por ela.
n <- trace(I)
p <- trace(H)
# Matriz de projeção da média (modelo nulo).
H_b0 <- proj(X[, "b0"])
# Soma de quadrados da regressão (corrigida) pra média e dos resíduos.
SQReg <- t(y) %*% (H - H_b0) %*% y
SQRes <- t(y) %*% (I - H) %*% y
# Os graus de liberdade.
trace((H - H_b0))## [1] 3trace((I - H))## [1] 16# Quadrados médios
QMReg <- SQReg/(p - 1)
QMR <- SQRes/(n - p)
rQMR <- sqrt(QMR)
# Valor de F para ajuste do modelo.
F <- QMReg/QMR
pf(F, p - 1, n - p, lower = FALSE)## saleprice
## saleprice 3.899898e-08# Valores preditos pelo modelo.
hy <- crossprod(H, y)
#-----------------------------------------------------------------------
# Pontos de alavancagem.
hii <- diag(H)
plot(hii)
abline(h = 2 * p/n, lty = 2, col = 2)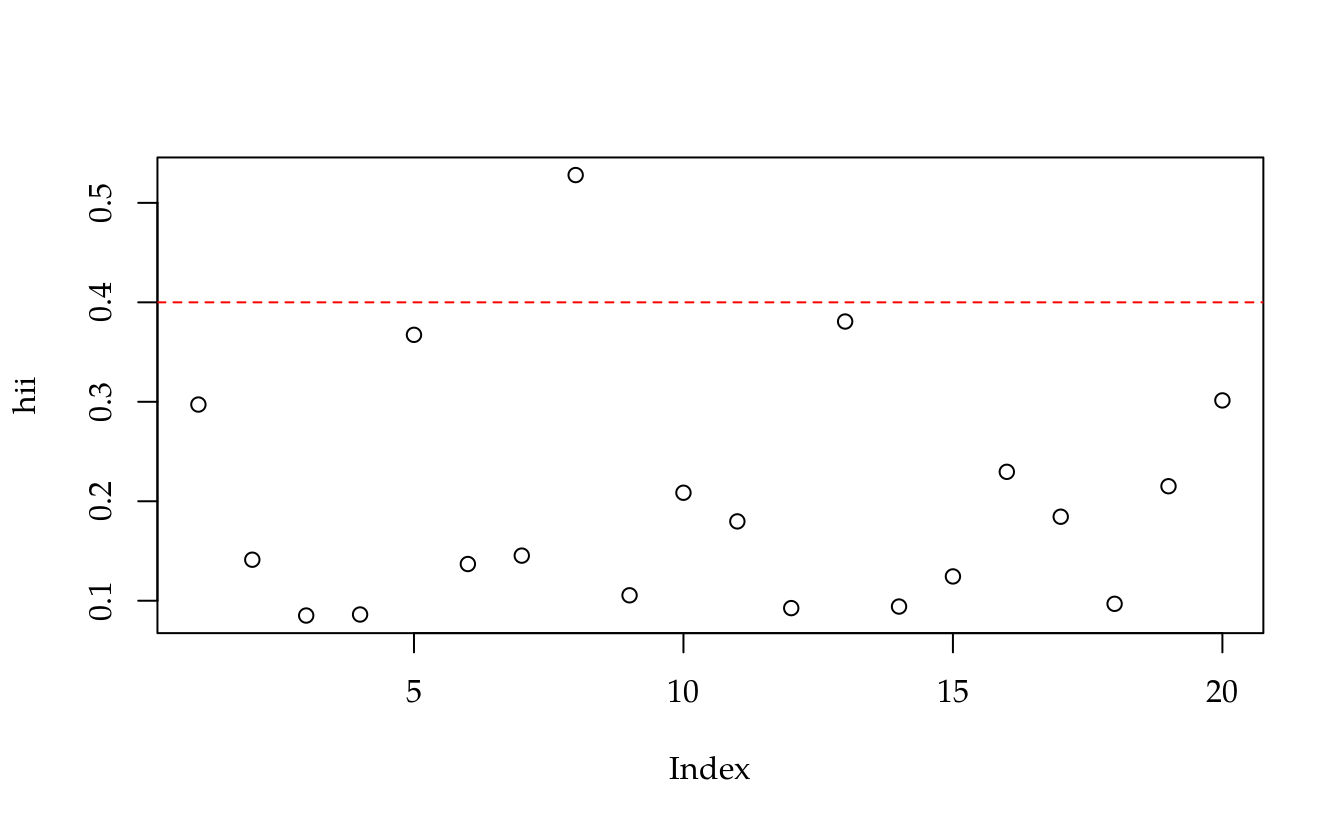
cbind(hatvalues(m0), hii)## hii
## 1 0.29721478 0.29721478
## 2 0.14121680 0.14121680
## 3 0.08509103 0.08509103
## 4 0.08613511 0.08613511
## 5 0.36730340 0.36730340
## 6 0.13688377 0.13688377
## 7 0.14533094 0.14533094
## 8 0.52794088 0.52794088
## 9 0.10542731 0.10542731
## 10 0.20855291 0.20855291
## 11 0.17973851 0.17973851
## 12 0.09255776 0.09255776
## 13 0.38069000 0.38069000
## 14 0.09413692 0.09413692
## 15 0.12441213 0.12441213
## 16 0.22958162 0.22958162
## 17 0.18444624 0.18444624
## 18 0.09688042 0.09688042
## 19 0.21508782 0.21508782
## 20 0.30137165 0.30137165#-----------------------------------------------------------------------
# Resíduos crus.
# e <- y - hy
# e <- y - crossprod(X, b)
e <- c(crossprod(I - H, y))
plot(e)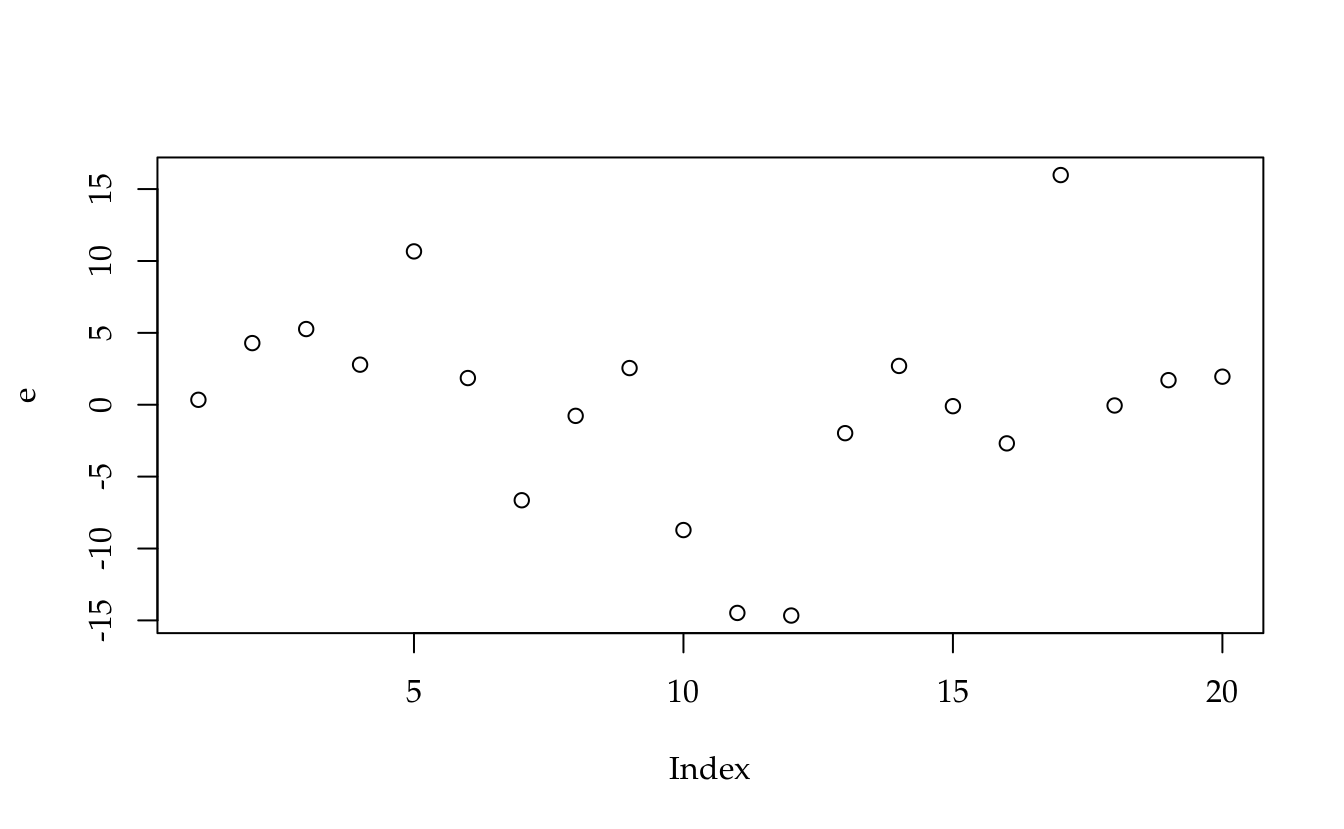
cbind(residuals(m0), e)## e
## 1 0.34326507 0.34326507
## 2 4.28713670 4.28713670
## 3 5.26484739 5.26484739
## 4 2.78803439 2.78803439
## 5 10.66594524 10.66594524
## 6 1.85634162 1.85634162
## 7 -6.64364995 -6.64364995
## 8 -0.77357950 -0.77357950
## 9 2.55052449 2.55052449
## 10 -8.71683945 -8.71683945
## 11 -14.48097731 -14.48097731
## 12 -14.66237009 -14.66237009
## 13 -1.97713293 -1.97713293
## 14 2.69961720 2.69961720
## 15 -0.10013601 -0.10013601
## 16 -2.68694921 -2.68694921
## 17 15.97560222 15.97560222
## 18 -0.05204213 -0.05204213
## 19 1.71028447 1.71028447
## 20 1.95207778 1.95207778#-----------------------------------------------------------------------
# Resíduos padronizados ou internamente studentizados.
r <- e/sqrt(c(QMR) * (1 - hii))
plot(r)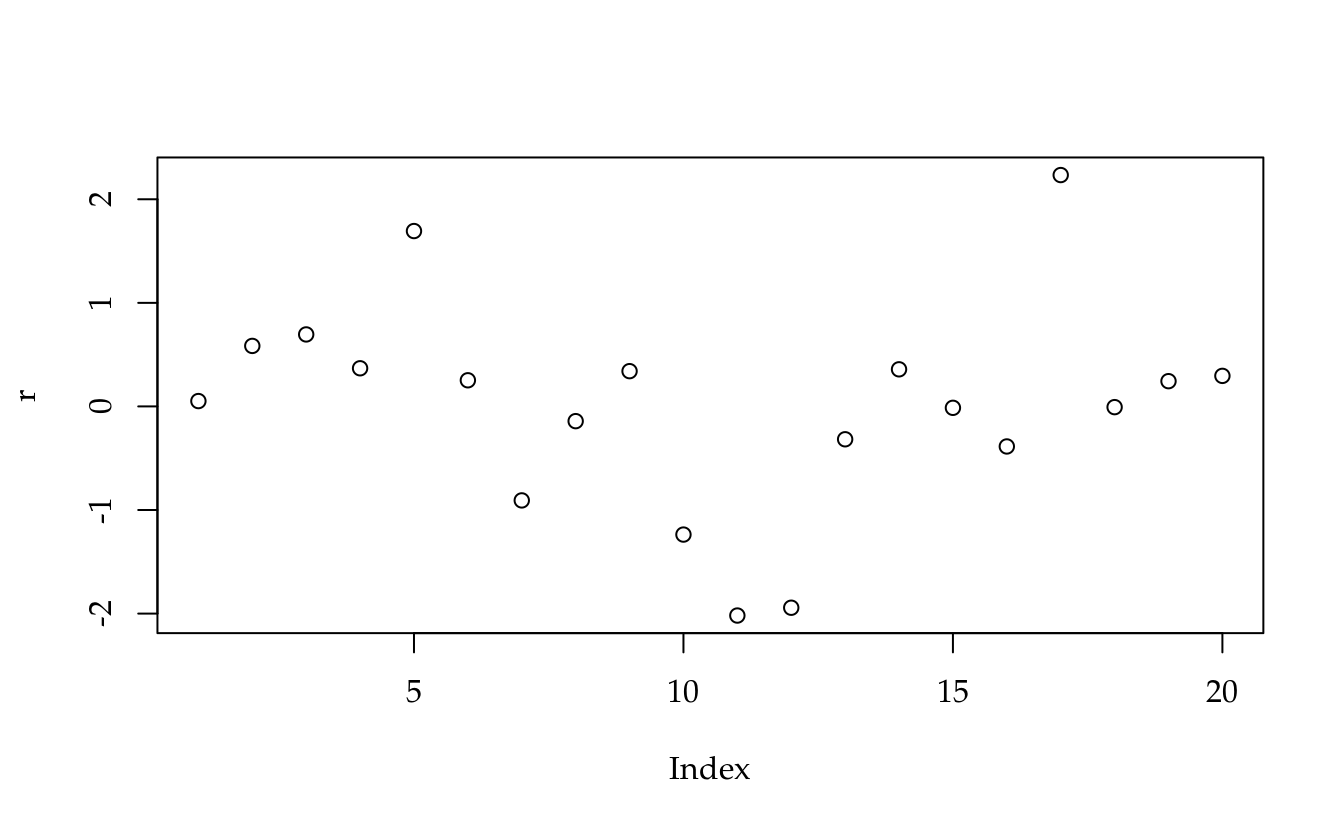
cbind(rstandard(m0), r)## r
## 1 0.051703685 0.051703685
## 2 0.584155884 0.584155884
## 3 0.695024379 0.695024379
## 4 0.368264897 0.368264897
## 5 1.693186488 1.693186488
## 6 0.252305347 0.252305347
## 7 -0.907425428 -0.907425428
## 8 -0.142170649 -0.142170649
## 9 0.340506082 0.340506082
## 10 -1.237232437 -1.237232437
## 11 -2.018946947 -2.018946947
## 12 -1.943559660 -1.943559660
## 13 -0.317237864 -0.317237864
## 14 0.358157543 0.358157543
## 15 -0.013512746 -0.013512746
## 16 -0.386544354 -0.386544354
## 17 2.233747784 2.233747784
## 18 -0.006914895 -0.006914895
## 19 0.243759198 0.243759198
## 20 0.294901656 0.294901656#-----------------------------------------------------------------------
# Estimativas leave one out.
# Estimativas dos parâmetros sem a i-ésima observação.
bi <- 0 * t(X)
for (i in 1:n) {
bi[, i] <- b - e[i] * crossprod(iXlX, X[i, ])/(1 - hii[i])
}
t(bi)## b0 b1 b2 b3
## [1,] 1.5495414 0.8280208 0.8190054 13.400322
## [2,] 0.3553034 0.7589914 0.8170813 14.609679
## [3,] 0.6038282 0.7699289 0.8185515 14.292098
## [4,] 1.1775069 0.8085580 0.8082694 13.980312
## [5,] 5.2862179 0.8539770 0.6561456 14.091765
## [6,] 0.9837925 0.7937361 0.8186134 13.987550
## [7,] 3.2226525 0.8605318 0.8451833 11.604738
## [8,] 1.4400526 0.8843033 0.8099139 13.413586
## [9,] 1.1692898 0.8280933 0.8039131 13.974565
## [10,] 0.6624660 0.7969022 0.7273459 17.004255
## [11,] 4.0310386 0.6898487 0.9741052 9.223385
## [12,] 1.3149387 0.7403023 0.7627215 16.192478
## [13,] 0.5443722 0.8553346 0.8046348 14.444900
## [14,] 1.0614468 0.7940970 0.8341251 13.503081
## [15,] 1.4852686 0.8154461 0.8196796 13.535108
## [16,] 0.8715999 0.7514001 0.8434409 13.920655
## [17,] 0.6960190 1.0269219 0.9385098 8.951832
## [18,] 1.4802808 0.8142129 0.8204106 13.526217
## [19,] 1.2034635 0.8586896 0.8017954 13.824444
## [20,] 0.9792723 0.8281335 0.8485870 12.973374# Valores preditos sem a i-ésima observação.
hyi <- diag(X %*% bi)
hyi## [1] 68.41156 43.50789 49.74550 58.94918 99.64209 42.84926
## [7] 45.77336 84.63873 56.14889 58.51380 58.15410 56.15791
## [13] 100.19248 42.51984 41.01436 83.48765 36.41134 37.05762
## [19] 47.82105 19.60584plot(hyi ~ hy)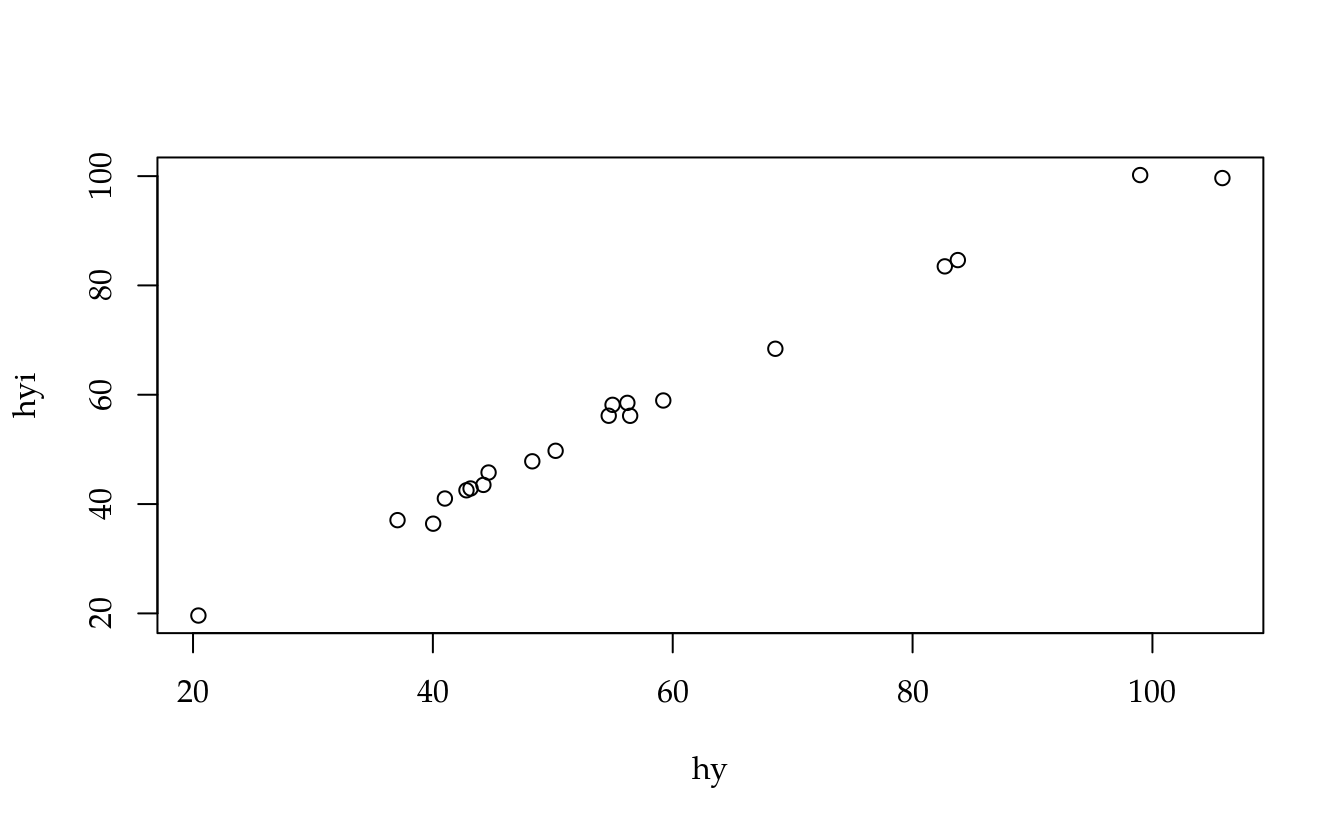
# Quadrado médio sem a i-ésima observação.
QMRi <- ((n - p) * QMR - (e^2/(1 - hii)))/(n - 1 - p)
QMRi## [1] 66.88824 65.47263 64.87964 66.33237 54.91238 66.63325 63.45652
## [8] 66.81490 66.41463 60.49905 49.85618 51.10520 66.47862 66.36306
## [15] 66.89865 66.27467 46.03671 66.89922 66.65098 66.53579#-----------------------------------------------------------------------
# Resíduos studentizados ou externamente studentizados.
s <- e/sqrt(QMRi * (1 - hii))
plot(s)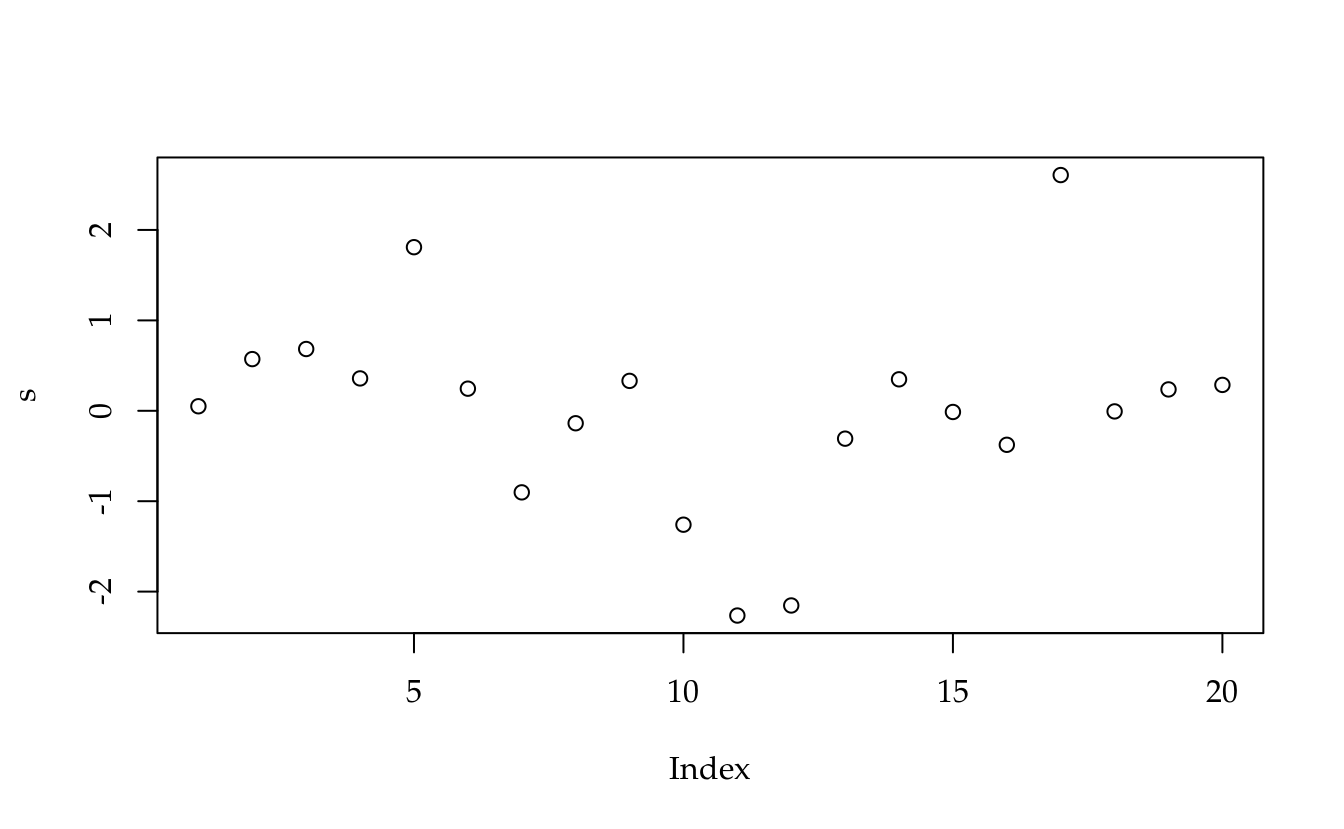
cbind(rstudent(m0), s)## s
## 1 0.050066061 0.050066061
## 2 0.571736179 0.571736179
## 3 0.683349077 0.683349077
## 4 0.358091810 0.358091810
## 5 1.809532865 1.809532865
## 6 0.244781033 0.244781033
## 7 -0.902131048 -0.902131048
## 8 -0.137743171 -0.137743171
## 9 0.330894694 0.330894694
## 10 -1.259719431 -1.259719431
## 11 -2.264447340 -2.264447340
## 12 -2.153089760 -2.153089760
## 13 -0.308134853 -0.308134853
## 14 0.348183103 0.348183103
## 15 -0.013083735 -0.013083735
## 16 -0.376029863 -0.376029863
## 17 2.607226676 2.607226676
## 18 -0.006695329 -0.006695329
## 19 0.236458300 0.236458300
## 20 0.286316488 0.286316488pvals <- pt(abs(s), df = n - 1 - p, lower.tail = FALSE)
p.adjust(pvals, method = "bonferroni")## [1] 1.0000000 1.0000000 1.0000000 1.0000000 0.9044904 1.0000000
## [7] 1.0000000 1.0000000 1.0000000 1.0000000 0.3879525 0.4799291
## [13] 1.0000000 1.0000000 1.0000000 1.0000000 0.1981185 1.0000000
## [19] 1.0000000 1.0000000#-----------------------------------------------------------------------
# Distância de Cook.
D <- hii * (e^2)/(p * c(QMR) * (1 - hii)^2)
plot(D)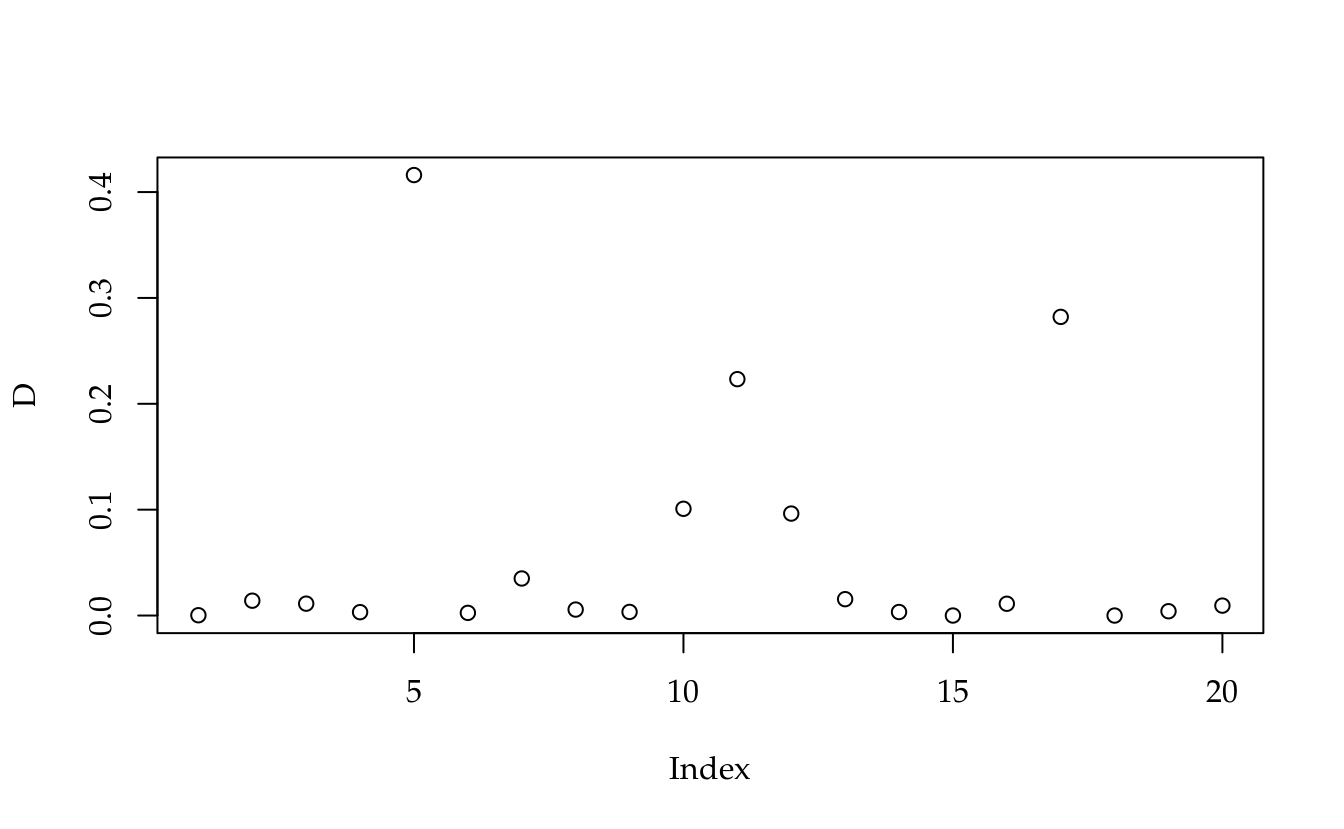
cbind(cooks.distance(m0), D)## D
## 1 2.826382e-04 2.826382e-04
## 2 1.402815e-02 1.402815e-02
## 3 1.123171e-02 1.123171e-02
## 4 3.195647e-03 3.195647e-03
## 5 4.160821e-01 4.160821e-01
## 6 2.523920e-03 2.523920e-03
## 7 3.500435e-02 3.500435e-02
## 8 5.651306e-03 5.651306e-03
## 9 3.416074e-03 3.416074e-03
## 10 1.008410e-01 1.008410e-01
## 11 2.232948e-01 2.232948e-01
## 12 9.632292e-02 9.632292e-02
## 13 1.546584e-02 1.546584e-02
## 14 3.332619e-03 3.332619e-03
## 15 6.486198e-06 6.486198e-06
## 16 1.113138e-02 1.113138e-02
## 17 2.821145e-01 2.821145e-01
## 18 1.282336e-06 1.282336e-06
## 19 4.070585e-03 4.070585e-03
## 20 9.378872e-03 9.378872e-03#-----------------------------------------------------------------------
# DFfits.
# dffits <- (c(hy) - c(hyi))/sqrt(QMRi * hii)
dffits <- s * sqrt((hii/(1 - hii)))
plot(dffits)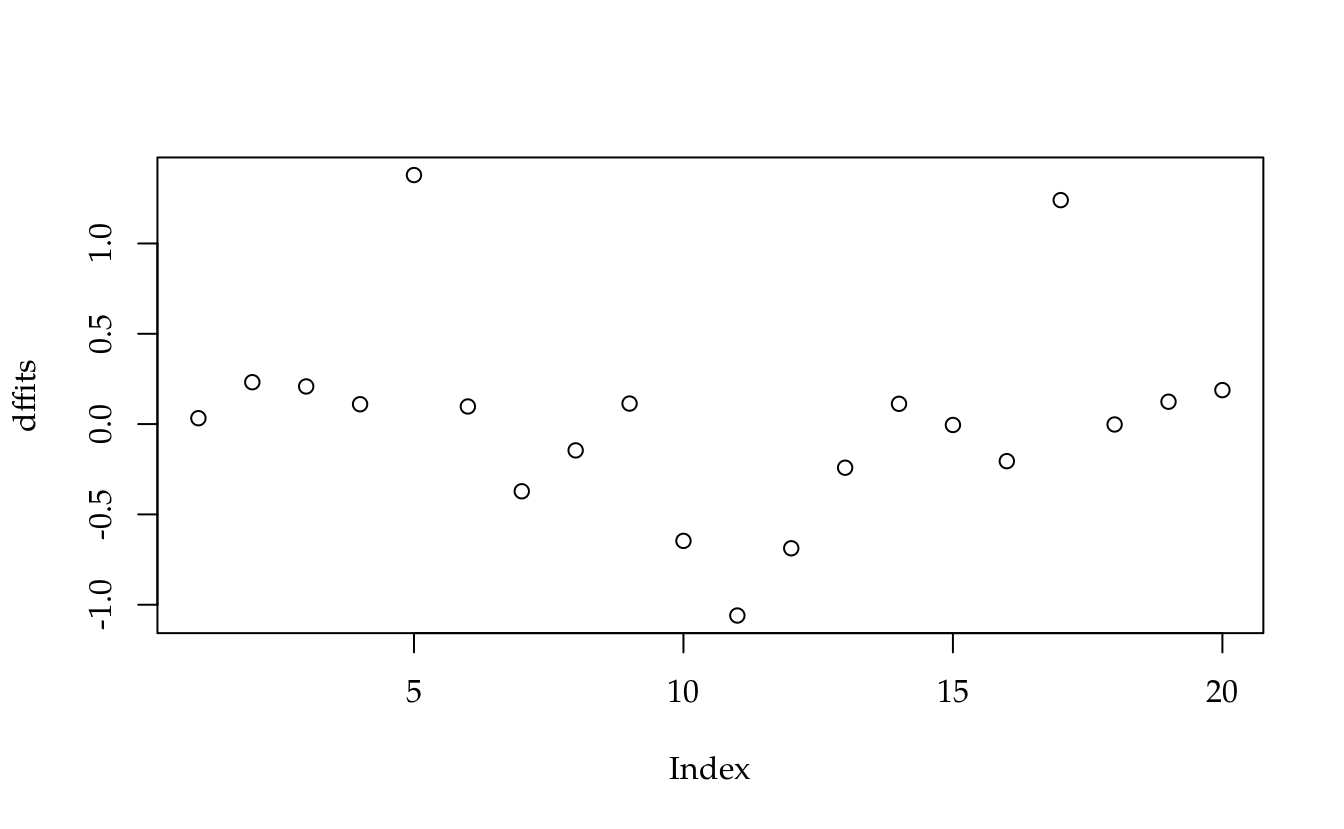
cbind(dffits(m0), dffits)## dffits
## 1 0.032558720 0.032558720
## 2 0.231844656 0.231844656
## 3 0.208398965 0.208398965
## 4 0.109936900 0.109936900
## 5 1.378736269 1.378736269
## 6 0.097480803 0.097480803
## 7 -0.372005775 -0.372005775
## 8 -0.145668122 -0.145668122
## 9 0.113594823 0.113594823
## 10 -0.646652572 -0.646652572
## 11 -1.060001884 -1.060001884
## 12 -0.687636725 -0.687636725
## 13 -0.241586417 -0.241586417
## 14 0.112242260 0.112242260
## 15 -0.004931888 -0.004931888
## 16 -0.205270991 -0.205270991
## 17 1.239902094 1.239902094
## 18 -0.002192892 -0.002192892
## 19 0.123780417 0.123780417
## 20 0.188050485 0.188050485#-----------------------------------------------------------------------
# DFbetas.
# dfbetas <- 0 * t(X)
# for (i in 1:n) {
# num <- e[i] * c(crossprod(iXlX, X[i, ]))/(1 - hii[i])
# den <- sqrt(QMRi[i] * diag(iXlX))
# dfbetas[, i] <- num/den
# }
num <- t(sweep(iXlX %*% t(X), 2, e/(1 - hii), "*"))
den <- outer(sqrt(QMRi), sqrt(diag(iXlX)), "*")
dfbetas <- num/den; dfbetas## b0 b1 b2 b3
## [1,] -0.013357206 -0.0255791757 0.0065994417 0.0188687060
## [2,] 0.189906987 0.1060459514 0.0155876421 -0.1606585314
## [3,] 0.148250023 0.0855351239 0.0088139472 -0.1139780947
## [4,] 0.049541486 0.0112613722 0.0560596793 -0.0666878921
## [5,] -0.709697529 -0.0823869963 0.8314457891 -0.0913814452
## [6,] 0.082135170 0.0393095493 0.0084129640 -0.0676033687
## [7,] -0.303176808 -0.0893623801 -0.1164585556 0.2904312440
## [8,] 0.005095804 -0.1320512114 0.0483122339 0.0169277796
## [9,] 0.050900401 -0.0258076893 0.0760704753 -0.0657985087
## [10,] 0.143133594 0.0349608993 0.4488526692 -0.5373439259
## [11,] -0.499823291 0.2729255496 -0.8160871901 0.7332237670
## [12,] 0.029946754 0.1604506692 0.3027971856 -0.4480947782
## [13,] 0.156506181 -0.0774521758 0.0727144700 -0.1351353794
## [14,] 0.069164813 0.0387045021 -0.0629752868 0.0037743780
## [15,] -0.002526252 -0.0018071770 0.0035078271 -0.0009495220
## [16,] 0.101350226 0.1198196523 -0.1059293160 -0.0579048016
## [17,] 0.157267714 -0.4840706407 -0.6525340542 0.8111621803
## [18,] -0.001685814 0.0005240004 0.0001563683 0.0003576845
## [19,] 0.045041147 -0.0837058087 0.0856631047 -0.0435694922
## [20,] 0.082959029 -0.0258604588 -0.1293794865 0.0818610922dfbetas(m0)## (Intercept) landvalue improvvalue area
## 1 -0.013357206 -0.0255791757 0.0065994417 0.0188687060
## 2 0.189906987 0.1060459514 0.0155876421 -0.1606585314
## 3 0.148250023 0.0855351239 0.0088139472 -0.1139780947
## 4 0.049541486 0.0112613722 0.0560596793 -0.0666878921
## 5 -0.709697529 -0.0823869963 0.8314457891 -0.0913814452
## 6 0.082135170 0.0393095493 0.0084129640 -0.0676033687
## 7 -0.303176808 -0.0893623801 -0.1164585556 0.2904312440
## 8 0.005095804 -0.1320512114 0.0483122339 0.0169277796
## 9 0.050900401 -0.0258076893 0.0760704753 -0.0657985087
## 10 0.143133594 0.0349608993 0.4488526692 -0.5373439259
## 11 -0.499823291 0.2729255496 -0.8160871901 0.7332237670
## 12 0.029946754 0.1604506692 0.3027971856 -0.4480947782
## 13 0.156506181 -0.0774521758 0.0727144700 -0.1351353794
## 14 0.069164813 0.0387045021 -0.0629752868 0.0037743780
## 15 -0.002526252 -0.0018071770 0.0035078271 -0.0009495220
## 16 0.101350226 0.1198196523 -0.1059293160 -0.0579048016
## 17 0.157267714 -0.4840706407 -0.6525340542 0.8111621803
## 18 -0.001685814 0.0005240004 0.0001563683 0.0003576845
## 19 0.045041147 -0.0837058087 0.0856631047 -0.0435694922
## 20 0.082959029 -0.0258604588 -0.1293794865 0.0818610922#-----------------------------------------------------------------------
# Medidas de influência.
class(m0)## [1] "lm"methods(class = "lm")## [1] add1 addterm alias
## [4] anova Anova attrassign
## [7] avPlot bootCase Boot
## [10] boxcox boxCox brief
## [13] case.names ceresPlot coerce
## [16] concordance confidenceEllipse confint
## [19] Confint cooks.distance crPlot
## [22] deltaMethod deviance dfbeta
## [25] dfbetaPlots dfbetas dfbetasPlots
## [28] drop1 dropterm dummy.coef
## [31] durbinWatsonTest effects emm_basis
## [34] extractAIC family formula
## [37] fortify hatvalues hccm
## [40] infIndexPlot influence influencePlot
## [43] initialize inverseResponsePlot kappa
## [46] labels leveneTest leveragePlot
## [49] linearHypothesis logLik logtrans
## [52] mcPlot mmp model.frame
## [55] model.matrix ncvTest nextBoot
## [58] nobs outlierTest plot
## [61] powerTransform predict Predict
## [64] print proj qqnorm
## [67] qqPlot qr recover_data
## [70] residualPlot residualPlots residuals
## [73] rstandard rstudent show
## [76] sigmaHat simulate S
## [79] slotsFromS3 spreadLevelPlot summary
## [82] variable.names vcov
## see '?methods' for accessing help and source codeim <- influence.measures(m0)
summary(im)## Potentially influential observations of
## lm(formula = saleprice ~ landvalue + improvvalue + area, data = da) :
##
## dfb.1_ dfb.lndv dfb.impr dfb.area dffit cov.r cook.d hat
## 1 -0.01 -0.03 0.01 0.02 0.03 1.84_* 0.00 0.30
## 8 0.01 -0.13 0.05 0.02 -0.15 2.73_* 0.01 0.53
## 13 0.16 -0.08 0.07 -0.14 -0.24 2.04_* 0.02 0.38
## 20 0.08 -0.03 -0.13 0.08 0.19 1.81_* 0.01 0.30#-----------------------------------------------------------------------6.2 Decomposições matriciais
#-----------------------------------------------------------------------
# Ajuste pela expressão matricial (X'X)^{-1} (X'y).
plot(demand ~ Time, BOD)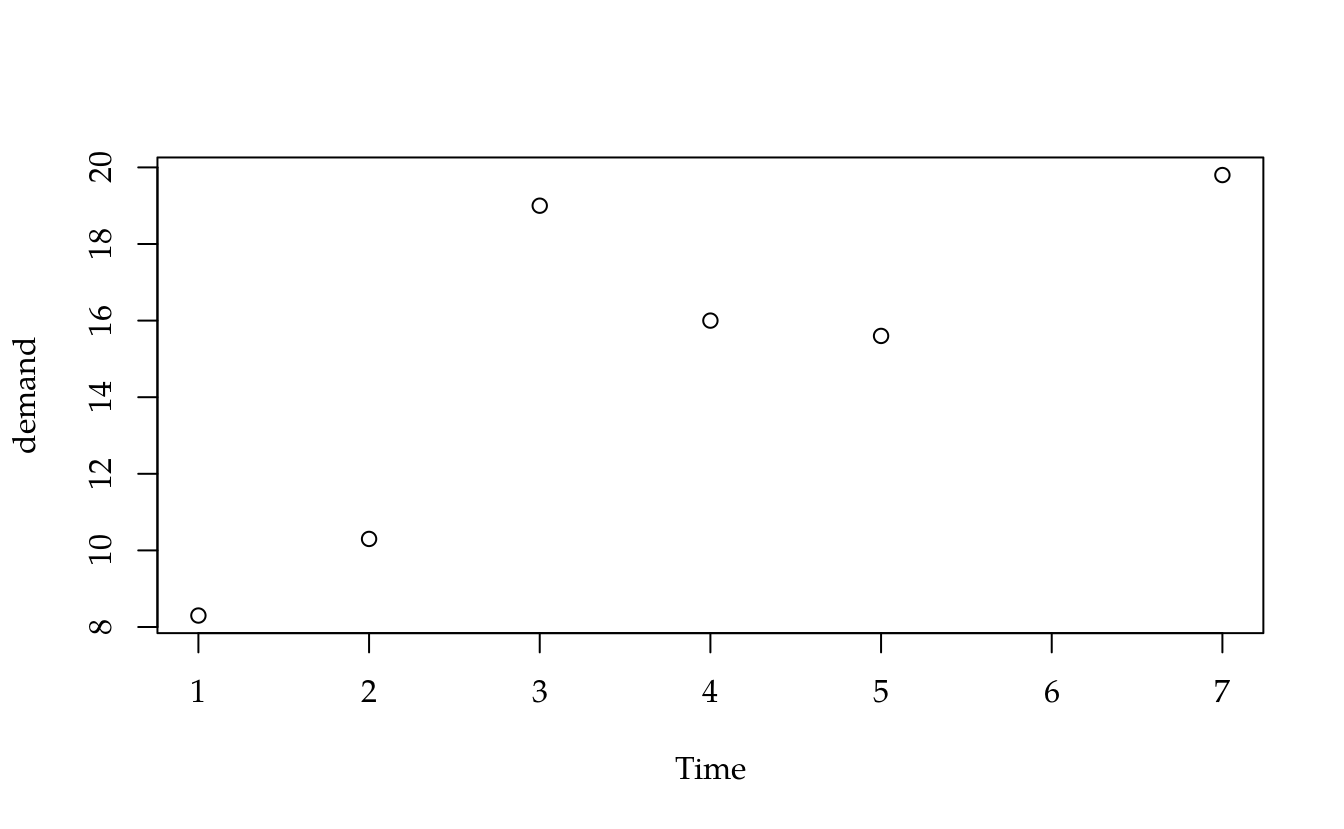
y <- cbind(BOD$demand)
X <- cbind(b0 = 1, b1 = BOD$Time)
# Expressão literal.
betas <- solve(t(X) %*% X) %*% (t(X) %*% y)
betas## [,1]
## b0 8.521429
## b1 1.721429# Código mais apropriado.
betas <- solve(crossprod(X), crossprod(X, y))
betas## [,1]
## b0 8.521429
## b1 1.721429#-----------------------------------------------------------------------
# Usando decomposição QR de X.
QR <- qr(X, LAPACK = TRUE)
str(QR)## List of 4
## $ qr : num [1:6, 1:2] -10.198 0.179 0.268 0.357 0.447 ...
## ..- attr(*, "dimnames")=List of 2
## .. ..$ : NULL
## .. ..$ : chr [1:2] "b1" "b0"
## $ rank : int 2
## $ qraux: num [1:2] 1.1 1.38
## $ pivot: int [1:2] 2 1
## - attr(*, "useLAPACK")= logi TRUE
## - attr(*, "class")= chr "qr"# Matriz Q (n x n).
Q <- qr.Q(QR, complete = TRUE)
Q## [,1] [,2] [,3] [,4] [,5]
## [1,] -0.09805807 -0.67956838 -0.3408594 -0.35352904 -0.36619871
## [2,] -0.19611614 -0.49724516 -0.1937412 0.04700555 0.28775231
## [3,] -0.29417420 -0.31492193 0.8958523 -0.08407540 -0.06400314
## [4,] -0.39223227 -0.13259871 -0.1111203 0.86489961 -0.15908043
## [5,] -0.49029034 0.04972452 -0.1180930 -0.18612537 0.74584228
## [6,] -0.68640647 0.41437097 -0.1320384 -0.28817535 -0.44431231
## [,6]
## [1,] -0.39153805
## [2,] 0.76924583
## [3,] -0.02385861
## [4,] -0.20704051
## [5,] -0.39022242
## [6,] 0.24341377# Q'Q = I.
round(t(Q) %*% Q, 3)## [,1] [,2] [,3] [,4] [,5] [,6]
## [1,] 1 0 0 0 0 0
## [2,] 0 1 0 0 0 0
## [3,] 0 0 1 0 0 0
## [4,] 0 0 0 1 0 0
## [5,] 0 0 0 0 1 0
## [6,] 0 0 0 0 0 1# Matriz R (n x p).
R <- qr.R(QR, complete = TRUE)
R## b1 b0
## [1,] -10.19804 -2.157277
## [2,] 0.00000 -1.160239
## [3,] 0.00000 0.000000
## [4,] 0.00000 0.000000
## [5,] 0.00000 0.000000
## [6,] 0.00000 0.000000z <- t(Q) %*% y
betas <- backsolve(R, z)
betas## [,1]
## [1,] 1.721429
## [2,] 8.521429#-----------------------------------------------------------------------
# Usando decomposição de Cholesky de X'X.
# XlX <- t(X) %*% X
XlX <- crossprod(X)
XlX## b0 b1
## b0 6 22
## b1 22 104L <- chol(XlX) # Triangular superior p x p.
Lt <- t(L) # Triangular inferior p x p.
L## b0 b1
## b0 2.44949 8.981462
## b1 0.00000 4.830459# L'L = X'X.
Lt %*% L## b0 b1
## b0 6 22
## b1 22 104d <- t(X) %*% y
z <- forwardsolve(Lt, d)
z## [,1]
## [1,] 36.33410
## [2,] 8.31529betas <- backsolve(L, z)
betas## [,1]
## [1,] 8.521429
## [2,] 1.721429#-----------------------------------------------------------------------6.3 Quadro de análise de variância
6.4 Teste de Wald
TODO ver a nlme::anova().
6.5 Efeito do promalin
# https://github.com/pet-estatistica/labestData/blob/devel/data-raw/BanzattoQd4.5.2.txt
da <- labestData::BanzattoQd4.5.2
summary(da)## promalin bloco peso
## 12.5 :4 I :5 Min. :130.6
## 25.0 :4 II :5 1st Qu.:136.8
## 50.0 :4 III:5 Median :141.6
## 12.5+12.5 :4 IV :5 Mean :143.0
## Testemunha:4 3rd Qu.:146.4
## Max. :165.0# Passa testemunha para primeiro nível para ordem mais lógica.
da <- da %>%
mutate(promalin = fct_relevel(promalin, "Testemunha", after = 0))
# Tabela de frequência.
xtabs(~promalin + bloco, data = da)## bloco
## promalin I II III IV
## Testemunha 1 1 1 1
## 12.5 1 1 1 1
## 25.0 1 1 1 1
## 50.0 1 1 1 1
## 12.5+12.5 1 1 1 1# reshape2::dcast(da, promalin ~ bloco)
da$peso[2] <- 139.3 # Corrige valor registrado errado no pacote.
ggplot(data = da,
mapping = aes(x = promalin,
y = peso,
color = bloco,
group = bloco)) +
geom_point() +
geom_line() +
stat_summary(mapping = aes(group = 1),
geom = "line",
fun.y = "mean")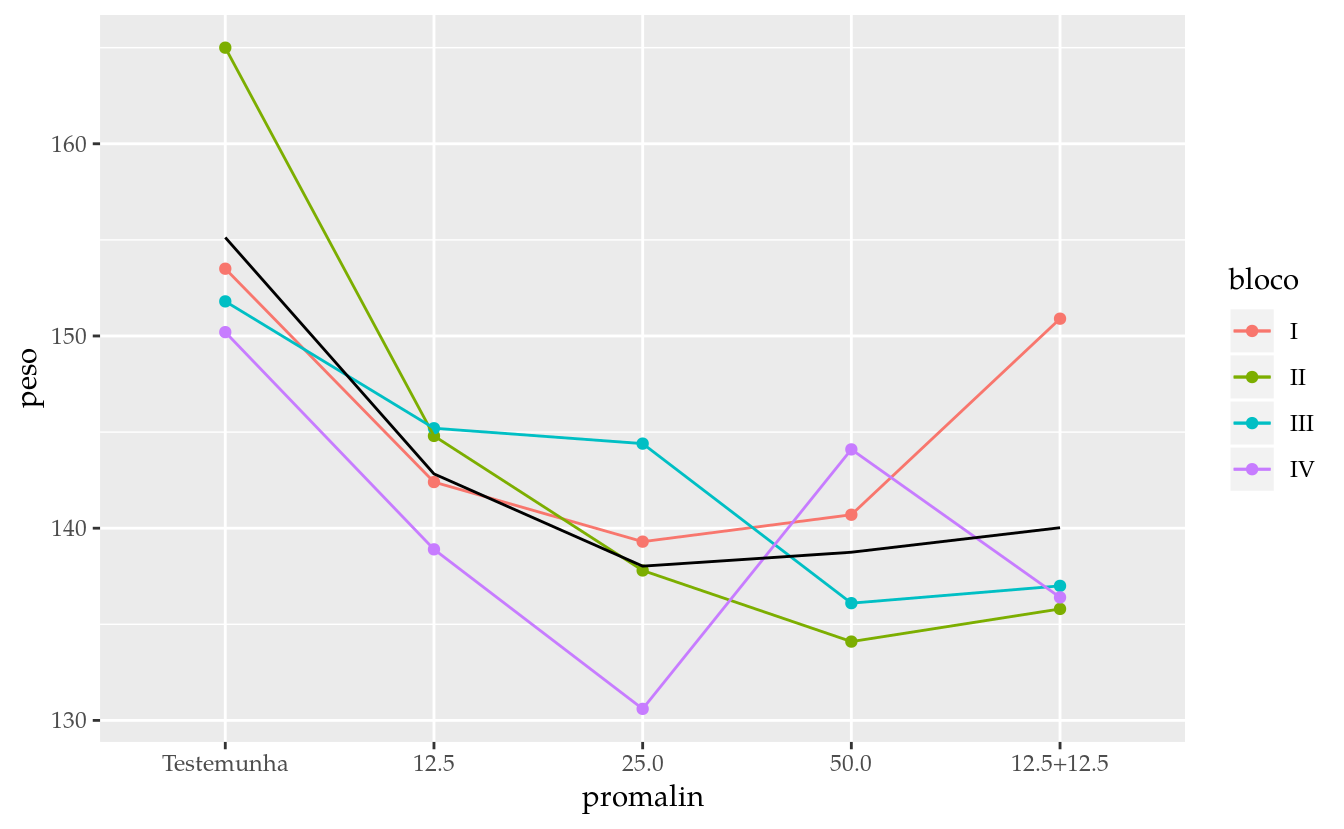
6.6 Castração
PimentelEx6.6.3## leitegada coluna castracao peso
## 1 1 1 56 93.0
## 2 2 1 Test 115.4
## 3 3 1 7 102.1
## 4 4 1 21 117.6
## 5 1 2 Test 108.6
## 6 2 2 21 96.5
## 7 3 2 56 94.9
## 8 4 2 7 114.1
## 9 1 3 7 108.9
## 10 2 3 56 77.9
## 11 3 3 21 116.9
## 12 4 3 Test 118.7
## 13 1 4 21 102.0
## 14 2 4 7 100.2
## 15 3 4 Test 96.0
## 16 4 4 56 97.6
Manual de Planejamento e Análise de Experimentos com R
Walmes Marques Zeviani